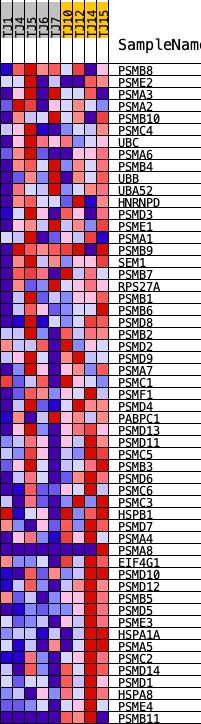
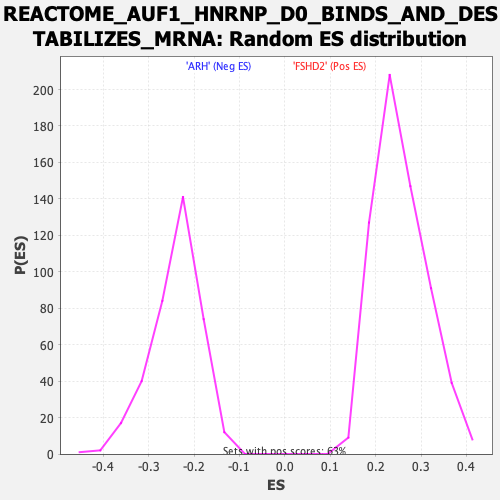
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_AUF1_HNRNP_D0_BINDS_AND_DESTABILIZES_MRNA |
| Enrichment Score (ES) | -0.53477347 |
| Normalized Enrichment Score (NES) | -2.2245557 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.5542145E-4 |
| FWER p-Value | 0.012 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMB8 | ENSG00000204264.11 | 14058 | 0.204 | -0.3232 | No | ||
| 2 | PSME2 | ENSG00000100911.16 | 19274 | 0.057 | -0.4449 | No | ||
| 3 | PSMA3 | ENSG00000100567.13 | 19401 | 0.054 | -0.4430 | No | ||
| 4 | PSMA2 | ENSG00000106588.11 | 19562 | 0.050 | -0.4423 | No | ||
| 5 | PSMB10 | ENSG00000205220.12 | 19937 | 0.040 | -0.4478 | No | ||
| 6 | PSMC4 | ENSG00000013275.8 | 20810 | 0.018 | -0.4674 | No | ||
| 7 | UBC | ENSG00000150991.15 | 20948 | 0.013 | -0.4695 | No | ||
| 8 | PSMA6 | ENSG00000100902.11 | 20976 | 0.013 | -0.4690 | No | ||
| 9 | PSMB4 | ENSG00000159377.11 | 21127 | 0.007 | -0.4719 | No | ||
| 10 | UBB | ENSG00000170315.14 | 22145 | -0.016 | -0.4952 | No | ||
| 11 | UBA52 | ENSG00000221983.8 | 22256 | -0.020 | -0.4960 | No | ||
| 12 | HNRNPD | ENSG00000138668.19 | 22348 | -0.022 | -0.4962 | No | ||
| 13 | PSMD3 | ENSG00000108344.15 | 23004 | -0.040 | -0.5085 | No | ||
| 14 | PSME1 | ENSG00000092010.15 | 23061 | -0.041 | -0.5061 | No | ||
| 15 | PSMA1 | ENSG00000129084.17 | 23337 | -0.048 | -0.5084 | No | ||
| 16 | PSMB9 | ENSG00000240065.8 | 23347 | -0.048 | -0.5042 | No | ||
| 17 | SEM1 | ENSG00000127922.9 | 24231 | -0.071 | -0.5191 | No | ||
| 18 | PSMB7 | ENSG00000136930.13 | 24754 | -0.084 | -0.5241 | No | ||
| 19 | RPS27A | ENSG00000143947.13 | 24801 | -0.086 | -0.5174 | No | ||
| 20 | PSMB1 | ENSG00000008018.9 | 25518 | -0.106 | -0.5250 | Yes | ||
| 21 | PSMB6 | ENSG00000142507.10 | 25718 | -0.112 | -0.5196 | Yes | ||
| 22 | PSMD8 | ENSG00000099341.11 | 25815 | -0.114 | -0.5115 | Yes | ||
| 23 | PSMB2 | ENSG00000126067.12 | 26219 | -0.126 | -0.5097 | Yes | ||
| 24 | PSMD2 | ENSG00000175166.17 | 26310 | -0.129 | -0.5000 | Yes | ||
| 25 | PSMD9 | ENSG00000110801.14 | 26323 | -0.129 | -0.4884 | Yes | ||
| 26 | PSMA7 | ENSG00000101182.15 | 26454 | -0.134 | -0.4793 | Yes | ||
| 27 | PSMC1 | ENSG00000100764.14 | 26467 | -0.134 | -0.4673 | Yes | ||
| 28 | PSMF1 | ENSG00000125818.18 | 26669 | -0.140 | -0.4593 | Yes | ||
| 29 | PSMD4 | ENSG00000159352.15 | 26816 | -0.144 | -0.4496 | Yes | ||
| 30 | PABPC1 | ENSG00000070756.16 | 26850 | -0.145 | -0.4371 | Yes | ||
| 31 | PSMD13 | ENSG00000185627.18 | 27620 | -0.168 | -0.4404 | Yes | ||
| 32 | PSMD11 | ENSG00000108671.11 | 27717 | -0.171 | -0.4270 | Yes | ||
| 33 | PSMC5 | ENSG00000087191.13 | 27761 | -0.172 | -0.4122 | Yes | ||
| 34 | PSMB3 | ENSG00000277791.5 | 27956 | -0.179 | -0.4005 | Yes | ||
| 35 | PSMD6 | ENSG00000163636.10 | 28425 | -0.194 | -0.3941 | Yes | ||
| 36 | PSMC6 | ENSG00000100519.12 | 28449 | -0.195 | -0.3768 | Yes | ||
| 37 | PSMC3 | ENSG00000165916.8 | 28625 | -0.200 | -0.3626 | Yes | ||
| 38 | HSPB1 | ENSG00000106211.9 | 29472 | -0.226 | -0.3624 | Yes | ||
| 39 | PSMD7 | ENSG00000103035.11 | 30001 | -0.244 | -0.3528 | Yes | ||
| 40 | PSMA4 | ENSG00000041357.16 | 30107 | -0.247 | -0.3327 | Yes | ||
| 41 | PSMA8 | ENSG00000154611.14 | 31324 | -0.285 | -0.3360 | Yes | ||
| 42 | EIF4G1 | ENSG00000114867.20 | 31786 | -0.289 | -0.3207 | Yes | ||
| 43 | PSMD10 | ENSG00000101843.19 | 32694 | -0.324 | -0.3130 | Yes | ||
| 44 | PSMD12 | ENSG00000197170.10 | 32756 | -0.326 | -0.2845 | Yes | ||
| 45 | PSMB5 | ENSG00000100804.19 | 35039 | -0.367 | -0.3063 | Yes | ||
| 46 | PSMD5 | ENSG00000095261.14 | 35294 | -0.378 | -0.2777 | Yes | ||
| 47 | PSME3 | ENSG00000131467.11 | 35430 | -0.385 | -0.2456 | Yes | ||
| 48 | HSPA1A | ENSG00000204389.10 | 35629 | -0.395 | -0.2142 | Yes | ||
| 49 | PSMA5 | ENSG00000143106.13 | 35640 | -0.395 | -0.1781 | Yes | ||
| 50 | PSMC2 | ENSG00000161057.12 | 35948 | -0.403 | -0.1485 | Yes | ||
| 51 | PSMD14 | ENSG00000115233.12 | 36419 | -0.422 | -0.1212 | Yes | ||
| 52 | PSMD1 | ENSG00000173692.13 | 37725 | -0.504 | -0.1066 | Yes | ||
| 53 | HSPA8 | ENSG00000109971.14 | 37882 | -0.519 | -0.0627 | Yes | ||
| 54 | PSME4 | ENSG00000068878.15 | 38072 | -0.532 | -0.0184 | Yes | ||
| 55 | PSMB11 | ENSG00000222028.5 | 40874 | -1.018 | 0.0070 | Yes |