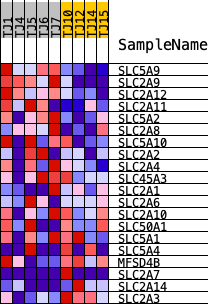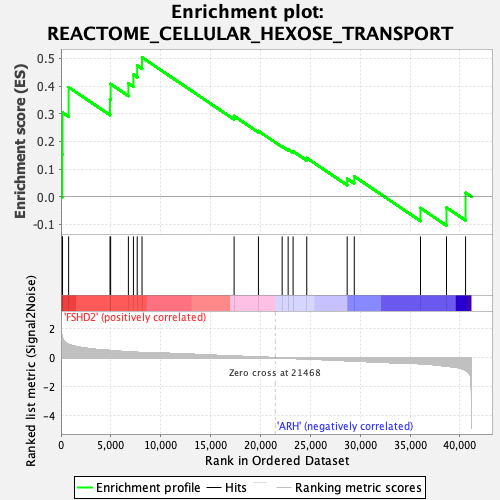
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_CELLULAR_HEXOSE_TRANSPORT |
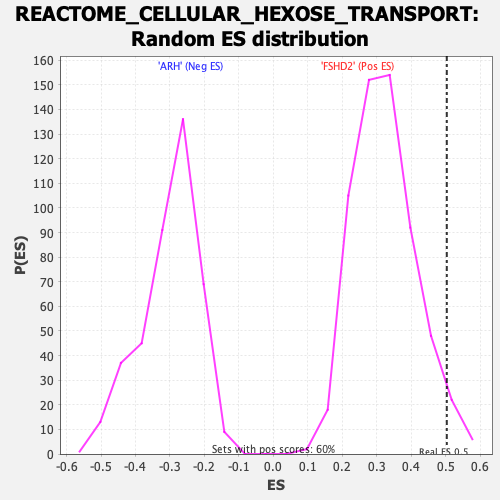
| Enrichment Score (ES) | 0.5036097 |
| Normalized Enrichment Score (NES) | 1.5581893 |
| Nominal p-value | 0.031719532 |
| FDR q-value | 0.22489277 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC5A9 | ENSG00000117834.12 | 127 | 1.355 | 0.1532 | Yes | ||
| 2 | SLC2A9 | ENSG00000109667.12 | 153 | 1.317 | 0.3044 | Yes | ||
| 3 | SLC2A12 | ENSG00000146411.6 | 765 | 0.922 | 0.3959 | Yes | ||
| 4 | SLC2A11 | ENSG00000133460.19 | 4916 | 0.496 | 0.3522 | Yes | ||
| 5 | SLC5A2 | ENSG00000140675.13 | 4979 | 0.492 | 0.4075 | Yes | ||
| 6 | SLC2A8 | ENSG00000136856.18 | 6757 | 0.398 | 0.4102 | Yes | ||
| 7 | SLC5A10 | ENSG00000154025.15 | 7263 | 0.377 | 0.4413 | Yes | ||
| 8 | SLC2A2 | ENSG00000163581.14 | 7642 | 0.365 | 0.4742 | Yes | ||
| 9 | SLC2A4 | ENSG00000181856.15 | 8126 | 0.356 | 0.5036 | Yes | ||
| 10 | SLC45A3 | ENSG00000158715.6 | 17358 | 0.108 | 0.2917 | No | ||
| 11 | SLC2A1 | ENSG00000117394.23 | 19798 | 0.043 | 0.2375 | No | ||
| 12 | SLC2A6 | ENSG00000160326.14 | 22176 | -0.017 | 0.1817 | No | ||
| 13 | SLC2A10 | ENSG00000197496.6 | 22777 | -0.033 | 0.1709 | No | ||
| 14 | SLC50A1 | ENSG00000169241.19 | 23273 | -0.046 | 0.1641 | No | ||
| 15 | SLC5A1 | ENSG00000100170.10 | 24643 | -0.081 | 0.1402 | No | ||
| 16 | SLC5A4 | ENSG00000100191.5 | 28696 | -0.202 | 0.0650 | No | ||
| 17 | MFSD4B | ENSG00000173214.6 | 29408 | -0.225 | 0.0736 | No | ||
| 18 | SLC2A7 | ENSG00000197241.3 | 36049 | -0.406 | -0.0410 | No | ||
| 19 | SLC2A14 | ENSG00000173262.11 | 38655 | -0.567 | -0.0389 | No | ||
| 20 | SLC2A3 | ENSG00000059804.16 | 40569 | -0.866 | 0.0144 | No |