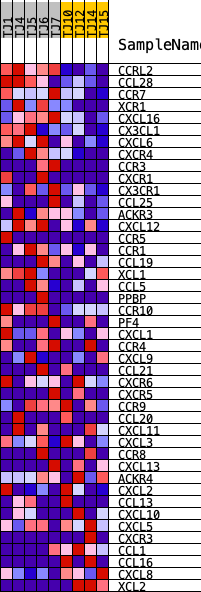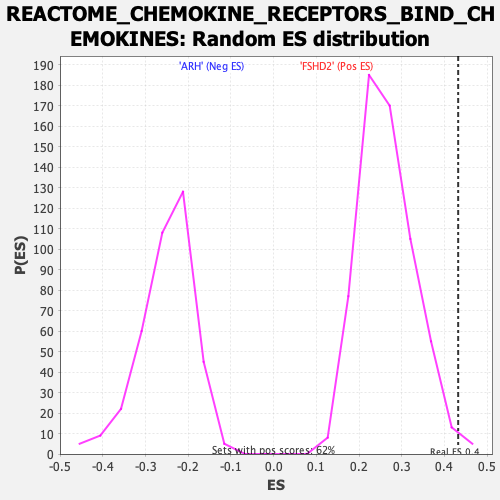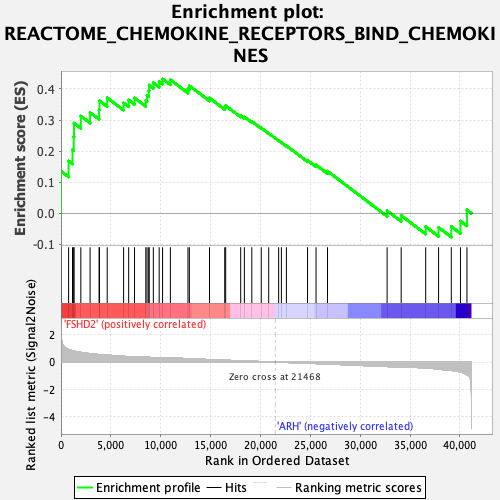
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_CHEMOKINE_RECEPTORS_BIND_CHEMOKINES |
| Enrichment Score (ES) | 0.43224216 |
| Normalized Enrichment Score (NES) | 1.6316983 |
| Nominal p-value | 0.008090615 |
| FDR q-value | 0.18785647 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CCRL2 | ENSG00000121797.10 | 4 | 2.444 | 0.1355 | Yes | ||
| 2 | CCL28 | ENSG00000151882.11 | 764 | 0.923 | 0.1683 | Yes | ||
| 3 | CCR7 | ENSG00000126353.3 | 1153 | 0.825 | 0.2047 | Yes | ||
| 4 | XCR1 | ENSG00000173578.7 | 1277 | 0.805 | 0.2463 | Yes | ||
| 5 | CXCL16 | ENSG00000161921.15 | 1306 | 0.801 | 0.2901 | Yes | ||
| 6 | CX3CL1 | ENSG00000006210.7 | 1988 | 0.707 | 0.3128 | Yes | ||
| 7 | CXCL6 | ENSG00000124875.10 | 2918 | 0.612 | 0.3242 | Yes | ||
| 8 | CXCR4 | ENSG00000121966.6 | 3819 | 0.548 | 0.3327 | Yes | ||
| 9 | CCR3 | ENSG00000183625.15 | 3860 | 0.547 | 0.3620 | Yes | ||
| 10 | CXCR1 | ENSG00000163464.8 | 4633 | 0.512 | 0.3717 | Yes | ||
| 11 | CX3CR1 | ENSG00000168329.13 | 6279 | 0.419 | 0.3549 | Yes | ||
| 12 | CCL25 | ENSG00000131142.13 | 6786 | 0.397 | 0.3646 | Yes | ||
| 13 | ACKR3 | ENSG00000144476.6 | 7372 | 0.373 | 0.3711 | Yes | ||
| 14 | CXCL12 | ENSG00000107562.16 | 8504 | 0.351 | 0.3631 | Yes | ||
| 15 | CCR5 | ENSG00000160791.13 | 8627 | 0.348 | 0.3794 | Yes | ||
| 16 | CCR1 | ENSG00000163823.4 | 8802 | 0.344 | 0.3943 | Yes | ||
| 17 | CCL19 | ENSG00000172724.12 | 8838 | 0.342 | 0.4124 | Yes | ||
| 18 | XCL1 | ENSG00000143184.5 | 9258 | 0.325 | 0.4202 | Yes | ||
| 19 | CCL5 | ENSG00000271503.6 | 9843 | 0.317 | 0.4236 | Yes | ||
| 20 | PPBP | ENSG00000163736.4 | 10187 | 0.305 | 0.4322 | Yes | ||
| 21 | CCR10 | ENSG00000184451.5 | 10967 | 0.292 | 0.4295 | No | ||
| 22 | PF4 | ENSG00000163737.4 | 12734 | 0.247 | 0.4002 | No | ||
| 23 | CXCL1 | ENSG00000163739.5 | 12872 | 0.242 | 0.4103 | No | ||
| 24 | CCR4 | ENSG00000183813.7 | 14891 | 0.177 | 0.3711 | No | ||
| 25 | CXCL9 | ENSG00000138755.6 | 16417 | 0.133 | 0.3414 | No | ||
| 26 | CCL21 | ENSG00000137077.8 | 16515 | 0.131 | 0.3463 | No | ||
| 27 | CXCR6 | ENSG00000172215.6 | 18020 | 0.090 | 0.3147 | No | ||
| 28 | CXCR5 | ENSG00000160683.5 | 18387 | 0.082 | 0.3103 | No | ||
| 29 | CCR9 | ENSG00000173585.16 | 19135 | 0.061 | 0.2956 | No | ||
| 30 | CCL20 | ENSG00000115009.13 | 20079 | 0.037 | 0.2747 | No | ||
| 31 | CXCL11 | ENSG00000169248.13 | 20828 | 0.017 | 0.2574 | No | ||
| 32 | CXCL3 | ENSG00000163734.4 | 21826 | -0.010 | 0.2338 | No | ||
| 33 | CCR8 | ENSG00000179934.7 | 22100 | -0.016 | 0.2280 | No | ||
| 34 | CXCL13 | ENSG00000156234.7 | 22603 | -0.029 | 0.2174 | No | ||
| 35 | ACKR4 | ENSG00000129048.7 | 24721 | -0.083 | 0.1705 | No | ||
| 36 | CXCL2 | ENSG00000081041.9 | 25575 | -0.107 | 0.1557 | No | ||
| 37 | CCL13 | ENSG00000181374.8 | 26725 | -0.141 | 0.1356 | No | ||
| 38 | CXCL10 | ENSG00000169245.6 | 32700 | -0.324 | 0.0083 | No | ||
| 39 | CXCL5 | ENSG00000163735.7 | 34111 | -0.346 | -0.0068 | No | ||
| 40 | CXCR3 | ENSG00000186810.8 | 36572 | -0.430 | -0.0428 | No | ||
| 41 | CCL1 | ENSG00000108702.4 | 37863 | -0.517 | -0.0454 | No | ||
| 42 | CCL16 | ENSG00000275152.5 | 39137 | -0.614 | -0.0423 | No | ||
| 43 | CXCL8 | ENSG00000169429.11 | 40055 | -0.722 | -0.0246 | No | ||
| 44 | XCL2 | ENSG00000143185.4 | 40706 | -0.927 | 0.0111 | No |