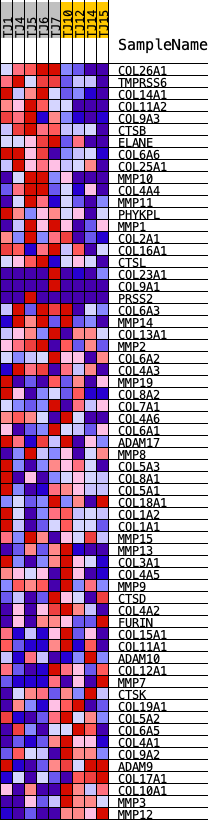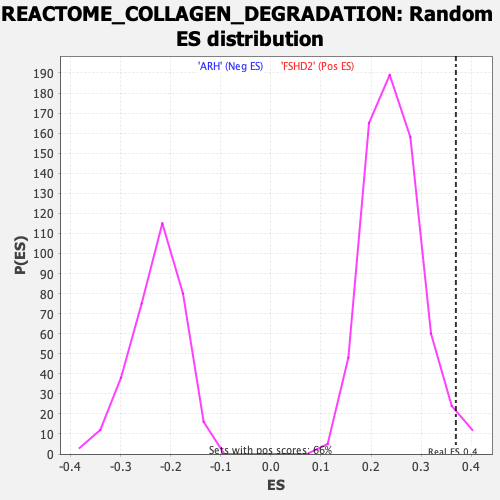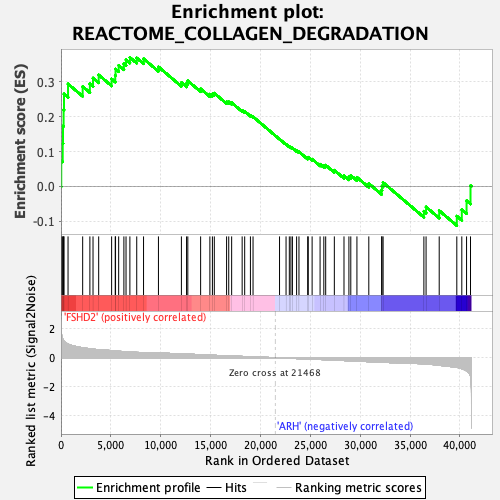
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_COLLAGEN_DEGRADATION |
| Enrichment Score (ES) | 0.3694199 |
| Normalized Enrichment Score (NES) | 1.5033642 |
| Nominal p-value | 0.024205748 |
| FDR q-value | 0.2374084 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COL26A1 | ENSG00000160963.14 | 26 | 1.811 | 0.0738 | Yes | ||
| 2 | TMPRSS6 | ENSG00000187045.18 | 170 | 1.289 | 0.1232 | Yes | ||
| 3 | COL14A1 | ENSG00000187955.12 | 202 | 1.233 | 0.1731 | Yes | ||
| 4 | COL11A2 | ENSG00000204248.10 | 261 | 1.164 | 0.2195 | Yes | ||
| 5 | COL9A3 | ENSG00000092758.18 | 301 | 1.131 | 0.2650 | Yes | ||
| 6 | CTSB | ENSG00000164733.21 | 706 | 0.938 | 0.2937 | Yes | ||
| 7 | ELANE | ENSG00000197561.7 | 2173 | 0.686 | 0.2862 | Yes | ||
| 8 | COL6A6 | ENSG00000206384.10 | 2896 | 0.614 | 0.2938 | Yes | ||
| 9 | COL25A1 | ENSG00000188517.16 | 3211 | 0.590 | 0.3104 | Yes | ||
| 10 | MMP10 | ENSG00000166670.10 | 3778 | 0.552 | 0.3193 | Yes | ||
| 11 | COL4A4 | ENSG00000081052.12 | 5076 | 0.485 | 0.3077 | Yes | ||
| 12 | MMP11 | ENSG00000099953.10 | 5438 | 0.463 | 0.3179 | Yes | ||
| 13 | PHYKPL | ENSG00000175309.15 | 5469 | 0.461 | 0.3361 | Yes | ||
| 14 | MMP1 | ENSG00000196611.5 | 5783 | 0.444 | 0.3467 | Yes | ||
| 15 | COL2A1 | ENSG00000139219.19 | 6306 | 0.417 | 0.3511 | Yes | ||
| 16 | COL16A1 | ENSG00000084636.18 | 6521 | 0.407 | 0.3627 | Yes | ||
| 17 | CTSL | ENSG00000135047.15 | 6908 | 0.393 | 0.3694 | Yes | ||
| 18 | COL23A1 | ENSG00000050767.17 | 7594 | 0.367 | 0.3678 | No | ||
| 19 | COL9A1 | ENSG00000112280.16 | 8277 | 0.352 | 0.3657 | No | ||
| 20 | PRSS2 | ENSG00000275896.5 | 9771 | 0.319 | 0.3425 | No | ||
| 21 | COL6A3 | ENSG00000163359.15 | 12073 | 0.268 | 0.2975 | No | ||
| 22 | MMP14 | ENSG00000157227.13 | 12590 | 0.252 | 0.2953 | No | ||
| 23 | COL13A1 | ENSG00000197467.14 | 12717 | 0.247 | 0.3024 | No | ||
| 24 | MMP2 | ENSG00000087245.13 | 14001 | 0.206 | 0.2796 | No | ||
| 25 | COL6A2 | ENSG00000142173.15 | 14929 | 0.176 | 0.2643 | No | ||
| 26 | COL4A3 | ENSG00000169031.19 | 15183 | 0.168 | 0.2650 | No | ||
| 27 | MMP19 | ENSG00000123342.16 | 15373 | 0.162 | 0.2671 | No | ||
| 28 | COL8A2 | ENSG00000171812.13 | 16605 | 0.128 | 0.2424 | No | ||
| 29 | COL7A1 | ENSG00000114270.17 | 16820 | 0.122 | 0.2422 | No | ||
| 30 | COL4A6 | ENSG00000197565.15 | 17112 | 0.114 | 0.2398 | No | ||
| 31 | COL6A1 | ENSG00000142156.14 | 18165 | 0.087 | 0.2178 | No | ||
| 32 | ADAM17 | ENSG00000151694.14 | 18419 | 0.082 | 0.2150 | No | ||
| 33 | MMP8 | ENSG00000118113.12 | 18994 | 0.065 | 0.2037 | No | ||
| 34 | COL5A3 | ENSG00000080573.7 | 19257 | 0.058 | 0.1997 | No | ||
| 35 | COL8A1 | ENSG00000144810.16 | 21914 | -0.012 | 0.1356 | No | ||
| 36 | COL5A1 | ENSG00000130635.15 | 22567 | -0.028 | 0.1208 | No | ||
| 37 | COL18A1 | ENSG00000182871.16 | 22895 | -0.036 | 0.1144 | No | ||
| 38 | COL1A2 | ENSG00000164692.17 | 23053 | -0.041 | 0.1122 | No | ||
| 39 | COL1A1 | ENSG00000108821.13 | 23206 | -0.045 | 0.1104 | No | ||
| 40 | MMP15 | ENSG00000102996.5 | 23626 | -0.056 | 0.1025 | No | ||
| 41 | MMP13 | ENSG00000137745.12 | 23862 | -0.062 | 0.0993 | No | ||
| 42 | COL3A1 | ENSG00000168542.14 | 24737 | -0.084 | 0.0815 | No | ||
| 43 | COL4A5 | ENSG00000188153.13 | 24793 | -0.085 | 0.0836 | No | ||
| 44 | MMP9 | ENSG00000100985.7 | 25181 | -0.096 | 0.0781 | No | ||
| 45 | CTSD | ENSG00000117984.14 | 25979 | -0.120 | 0.0637 | No | ||
| 46 | COL4A2 | ENSG00000134871.19 | 26354 | -0.131 | 0.0599 | No | ||
| 47 | FURIN | ENSG00000140564.12 | 26542 | -0.136 | 0.0610 | No | ||
| 48 | COL15A1 | ENSG00000204291.11 | 27409 | -0.161 | 0.0465 | No | ||
| 49 | COL11A1 | ENSG00000060718.22 | 28377 | -0.192 | 0.0309 | No | ||
| 50 | ADAM10 | ENSG00000137845.15 | 28870 | -0.207 | 0.0274 | No | ||
| 51 | COL12A1 | ENSG00000111799.21 | 29059 | -0.214 | 0.0316 | No | ||
| 52 | MMP7 | ENSG00000137673.9 | 29674 | -0.233 | 0.0263 | No | ||
| 53 | CTSK | ENSG00000143387.13 | 30865 | -0.272 | 0.0085 | No | ||
| 54 | COL19A1 | ENSG00000082293.13 | 32153 | -0.302 | -0.0104 | No | ||
| 55 | COL5A2 | ENSG00000204262.14 | 32186 | -0.304 | 0.0013 | No | ||
| 56 | COL6A5 | ENSG00000172752.14 | 32299 | -0.309 | 0.0113 | No | ||
| 57 | COL4A1 | ENSG00000187498.16 | 36388 | -0.420 | -0.0709 | No | ||
| 58 | COL9A2 | ENSG00000049089.15 | 36608 | -0.432 | -0.0585 | No | ||
| 59 | ADAM9 | ENSG00000168615.12 | 37925 | -0.522 | -0.0691 | No | ||
| 60 | COL17A1 | ENSG00000065618.21 | 39690 | -0.661 | -0.0849 | No | ||
| 61 | COL10A1 | ENSG00000123500.10 | 40191 | -0.750 | -0.0662 | No | ||
| 62 | MMP3 | ENSG00000149968.12 | 40668 | -0.906 | -0.0406 | No | ||
| 63 | MMP12 | ENSG00000262406.3 | 41064 | -1.280 | 0.0024 | No |