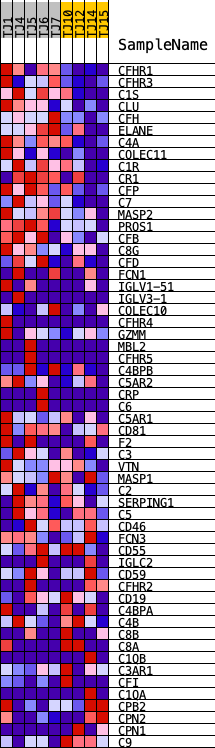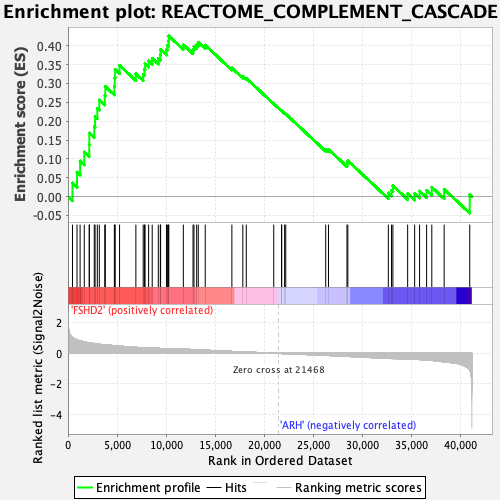
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
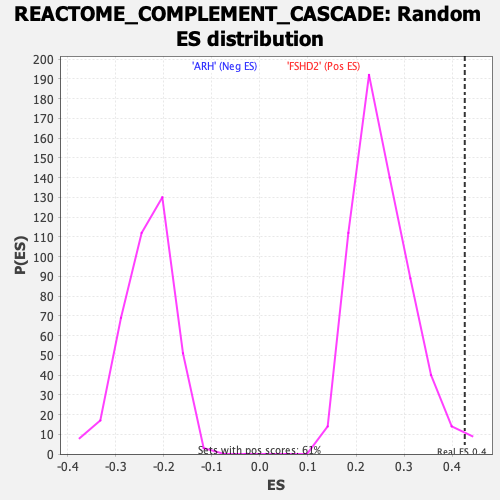
| GeneSet | REACTOME_COMPLEMENT_CASCADE |
| Enrichment Score (ES) | 0.4261623 |
| Normalized Enrichment Score (NES) | 1.6718168 |
| Nominal p-value | 0.011475409 |
| FDR q-value | 0.16877401 |
| FWER p-Value | 0.997 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CFHR1 | ENSG00000244414.6 | 457 | 1.033 | 0.0361 | Yes | ||
| 2 | CFHR3 | ENSG00000116785.13 | 920 | 0.878 | 0.0649 | Yes | ||
| 3 | C1S | ENSG00000182326.15 | 1233 | 0.811 | 0.0944 | Yes | ||
| 4 | CLU | ENSG00000120885.22 | 1659 | 0.752 | 0.1184 | Yes | ||
| 5 | CFH | ENSG00000000971.15 | 2168 | 0.686 | 0.1374 | Yes | ||
| 6 | ELANE | ENSG00000197561.7 | 2173 | 0.686 | 0.1687 | Yes | ||
| 7 | C4A | ENSG00000244731.8 | 2677 | 0.636 | 0.1855 | Yes | ||
| 8 | COLEC11 | ENSG00000118004.17 | 2760 | 0.627 | 0.2122 | Yes | ||
| 9 | C1R | ENSG00000159403.18 | 2987 | 0.607 | 0.2344 | Yes | ||
| 10 | CR1 | ENSG00000203710.11 | 3191 | 0.592 | 0.2565 | Yes | ||
| 11 | CFP | ENSG00000126759.13 | 3748 | 0.554 | 0.2682 | Yes | ||
| 12 | C7 | ENSG00000112936.19 | 3802 | 0.550 | 0.2921 | Yes | ||
| 13 | MASP2 | ENSG00000009724.17 | 4725 | 0.509 | 0.2929 | Yes | ||
| 14 | PROS1 | ENSG00000184500.15 | 4767 | 0.507 | 0.3150 | Yes | ||
| 15 | CFB | ENSG00000243649.8 | 4800 | 0.504 | 0.3373 | Yes | ||
| 16 | C8G | ENSG00000176919.13 | 5255 | 0.474 | 0.3479 | Yes | ||
| 17 | CFD | ENSG00000197766.8 | 6918 | 0.393 | 0.3254 | Yes | ||
| 18 | FCN1 | ENSG00000085265.11 | 7665 | 0.364 | 0.3239 | Yes | ||
| 19 | IGLV1-51 | ENSG00000211644.3 | 7791 | 0.363 | 0.3374 | Yes | ||
| 20 | IGLV3-1 | ENSG00000211673.2 | 7856 | 0.362 | 0.3524 | Yes | ||
| 21 | COLEC10 | ENSG00000184374.3 | 8219 | 0.353 | 0.3597 | Yes | ||
| 22 | CFHR4 | ENSG00000134365.13 | 8583 | 0.348 | 0.3668 | Yes | ||
| 23 | GZMM | ENSG00000197540.8 | 9204 | 0.327 | 0.3666 | Yes | ||
| 24 | MBL2 | ENSG00000165471.6 | 9409 | 0.319 | 0.3763 | Yes | ||
| 25 | CFHR5 | ENSG00000134389.9 | 9439 | 0.319 | 0.3901 | Yes | ||
| 26 | C4BPB | ENSG00000123843.13 | 10036 | 0.310 | 0.3898 | Yes | ||
| 27 | C5AR2 | ENSG00000134830.6 | 10132 | 0.306 | 0.4015 | Yes | ||
| 28 | CRP | ENSG00000132693.12 | 10259 | 0.305 | 0.4124 | Yes | ||
| 29 | C6 | ENSG00000039537.13 | 10266 | 0.305 | 0.4262 | Yes | ||
| 30 | C5AR1 | ENSG00000197405.8 | 11756 | 0.280 | 0.4027 | No | ||
| 31 | CD81 | ENSG00000110651.11 | 12744 | 0.247 | 0.3900 | No | ||
| 32 | F2 | ENSG00000180210.14 | 12841 | 0.243 | 0.3987 | No | ||
| 33 | C3 | ENSG00000125730.17 | 13115 | 0.235 | 0.4028 | No | ||
| 34 | VTN | ENSG00000109072.14 | 13279 | 0.229 | 0.4093 | No | ||
| 35 | MASP1 | ENSG00000127241.16 | 13988 | 0.206 | 0.4015 | No | ||
| 36 | C2 | ENSG00000166278.15 | 16704 | 0.125 | 0.3412 | No | ||
| 37 | SERPING1 | ENSG00000149131.15 | 17821 | 0.096 | 0.3184 | No | ||
| 38 | C5 | ENSG00000106804.7 | 18173 | 0.086 | 0.3138 | No | ||
| 39 | CD46 | ENSG00000117335.19 | 20971 | 0.013 | 0.2464 | No | ||
| 40 | FCN3 | ENSG00000142748.12 | 21773 | -0.008 | 0.2273 | No | ||
| 41 | CD55 | ENSG00000196352.15 | 21778 | -0.009 | 0.2276 | No | ||
| 42 | IGLC2 | ENSG00000211677.2 | 22079 | -0.016 | 0.2210 | No | ||
| 43 | CD59 | ENSG00000085063.16 | 22198 | -0.018 | 0.2189 | No | ||
| 44 | CFHR2 | ENSG00000080910.13 | 26270 | -0.128 | 0.1257 | No | ||
| 45 | CD19 | ENSG00000177455.14 | 26551 | -0.137 | 0.1252 | No | ||
| 46 | C4BPA | ENSG00000123838.11 | 28435 | -0.194 | 0.0882 | No | ||
| 47 | C4B | ENSG00000224389.9 | 28524 | -0.197 | 0.0951 | No | ||
| 48 | C8B | ENSG00000021852.13 | 32651 | -0.322 | 0.0095 | No | ||
| 49 | C8A | ENSG00000157131.10 | 32976 | -0.335 | 0.0169 | No | ||
| 50 | C1QB | ENSG00000173369.16 | 33116 | -0.336 | 0.0289 | No | ||
| 51 | C3AR1 | ENSG00000171860.5 | 34621 | -0.353 | 0.0084 | No | ||
| 52 | CFI | ENSG00000205403.13 | 35335 | -0.380 | 0.0084 | No | ||
| 53 | C1QA | ENSG00000173372.17 | 35837 | -0.402 | 0.0146 | No | ||
| 54 | CPB2 | ENSG00000080618.16 | 36554 | -0.430 | 0.0168 | No | ||
| 55 | CPN2 | ENSG00000178772.7 | 37083 | -0.457 | 0.0249 | No | ||
| 56 | CPN1 | ENSG00000120054.12 | 38342 | -0.555 | 0.0196 | No | ||
| 57 | C9 | ENSG00000113600.11 | 40949 | -1.072 | 0.0052 | No |