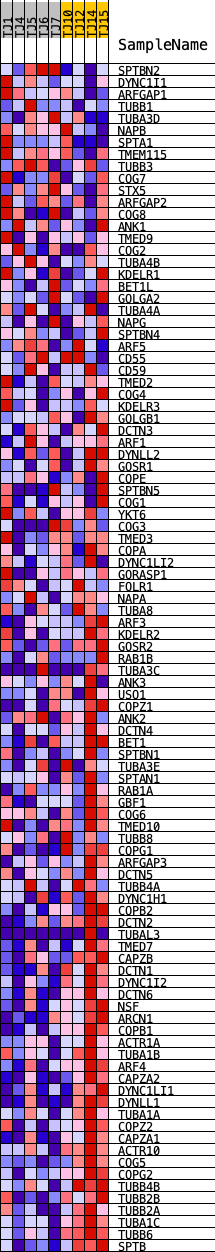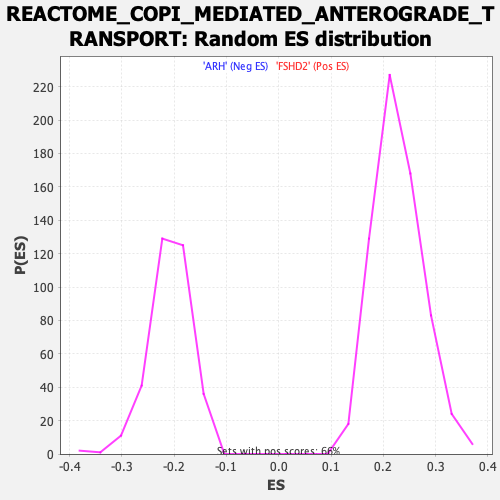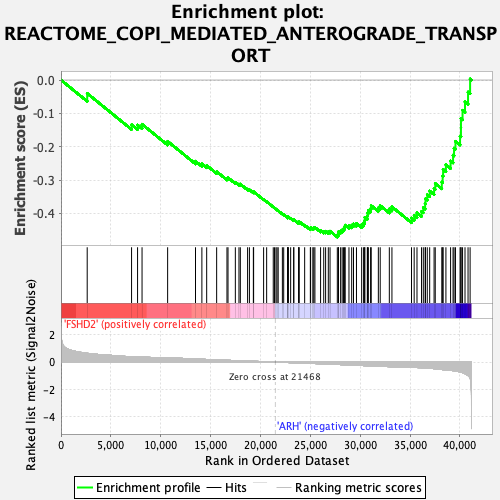
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_COPI_MEDIATED_ANTEROGRADE_TRANSPORT |
| Enrichment Score (ES) | -0.4710829 |
| Normalized Enrichment Score (NES) | -2.2487395 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.9037893E-4 |
| FWER p-Value | 0.009 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SPTBN2 | ENSG00000173898.13 | 2627 | 0.640 | -0.0400 | No | ||
| 2 | DYNC1I1 | ENSG00000158560.14 | 7076 | 0.386 | -0.1339 | No | ||
| 3 | ARFGAP1 | ENSG00000101199.13 | 7678 | 0.364 | -0.1350 | No | ||
| 4 | TUBB1 | ENSG00000101162.3 | 8127 | 0.356 | -0.1326 | No | ||
| 5 | TUBA3D | ENSG00000075886.11 | 10693 | 0.303 | -0.1837 | No | ||
| 6 | NAPB | ENSG00000125814.17 | 13487 | 0.222 | -0.2434 | No | ||
| 7 | SPTA1 | ENSG00000163554.15 | 14127 | 0.202 | -0.2515 | No | ||
| 8 | TMEM115 | ENSG00000126062.4 | 14608 | 0.187 | -0.2562 | No | ||
| 9 | TUBB3 | ENSG00000258947.7 | 15608 | 0.156 | -0.2747 | No | ||
| 10 | COG7 | ENSG00000168434.13 | 16642 | 0.127 | -0.2951 | No | ||
| 11 | STX5 | ENSG00000162236.12 | 16747 | 0.124 | -0.2930 | No | ||
| 12 | ARFGAP2 | ENSG00000149182.15 | 17482 | 0.104 | -0.3069 | No | ||
| 13 | COG8 | ENSG00000213380.15 | 17846 | 0.095 | -0.3122 | No | ||
| 14 | ANK1 | ENSG00000029534.20 | 17982 | 0.091 | -0.3121 | No | ||
| 15 | TMED9 | ENSG00000184840.12 | 18711 | 0.073 | -0.3271 | No | ||
| 16 | COG2 | ENSG00000135775.14 | 18893 | 0.068 | -0.3290 | No | ||
| 17 | TUBA4B | ENSG00000243910.8 | 19289 | 0.057 | -0.3365 | No | ||
| 18 | KDELR1 | ENSG00000105438.9 | 19315 | 0.056 | -0.3350 | No | ||
| 19 | BET1L | ENSG00000177951.17 | 20317 | 0.030 | -0.3582 | No | ||
| 20 | GOLGA2 | ENSG00000167110.17 | 20622 | 0.022 | -0.3648 | No | ||
| 21 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | -0.3808 | No | ||
| 22 | NAPG | ENSG00000134265.13 | 21412 | 0.001 | -0.3838 | No | ||
| 23 | SPTBN4 | ENSG00000160460.16 | 21556 | -0.003 | -0.3872 | No | ||
| 24 | ARF5 | ENSG00000004059.11 | 21663 | -0.006 | -0.3896 | No | ||
| 25 | CD55 | ENSG00000196352.15 | 21778 | -0.009 | -0.3920 | No | ||
| 26 | CD59 | ENSG00000085063.16 | 22198 | -0.018 | -0.4016 | No | ||
| 27 | TMED2 | ENSG00000086598.11 | 22331 | -0.022 | -0.4040 | No | ||
| 28 | COG4 | ENSG00000103051.19 | 22739 | -0.032 | -0.4127 | No | ||
| 29 | KDELR3 | ENSG00000100196.11 | 22740 | -0.032 | -0.4115 | No | ||
| 30 | GOLGB1 | ENSG00000173230.15 | 22743 | -0.032 | -0.4104 | No | ||
| 31 | DCTN3 | ENSG00000137100.16 | 22796 | -0.033 | -0.4104 | No | ||
| 32 | ARF1 | ENSG00000143761.16 | 23029 | -0.040 | -0.4145 | No | ||
| 33 | DYNLL2 | ENSG00000264364.3 | 23336 | -0.048 | -0.4202 | No | ||
| 34 | GOSR1 | ENSG00000108587.15 | 23341 | -0.048 | -0.4185 | No | ||
| 35 | COPE | ENSG00000105669.14 | 23839 | -0.062 | -0.4283 | No | ||
| 36 | SPTBN5 | ENSG00000137877.10 | 23841 | -0.062 | -0.4260 | No | ||
| 37 | COG1 | ENSG00000166685.12 | 23874 | -0.062 | -0.4245 | No | ||
| 38 | YKT6 | ENSG00000106636.8 | 24433 | -0.075 | -0.4352 | No | ||
| 39 | COG3 | ENSG00000136152.15 | 25003 | -0.092 | -0.4457 | No | ||
| 40 | TMED3 | ENSG00000166557.13 | 25030 | -0.093 | -0.4428 | No | ||
| 41 | COPA | ENSG00000122218.16 | 25255 | -0.097 | -0.4447 | No | ||
| 42 | DYNC1LI2 | ENSG00000135720.13 | 25352 | -0.101 | -0.4432 | No | ||
| 43 | GORASP1 | ENSG00000114745.13 | 25461 | -0.104 | -0.4420 | No | ||
| 44 | FOLR1 | ENSG00000110195.13 | 26030 | -0.121 | -0.4513 | No | ||
| 45 | NAPA | ENSG00000105402.8 | 26346 | -0.131 | -0.4540 | No | ||
| 46 | TUBA8 | ENSG00000183785.15 | 26534 | -0.136 | -0.4535 | No | ||
| 47 | ARF3 | ENSG00000134287.10 | 26805 | -0.144 | -0.4547 | No | ||
| 48 | KDELR2 | ENSG00000136240.10 | 26978 | -0.149 | -0.4534 | No | ||
| 49 | GOSR2 | ENSG00000108433.16 | 27707 | -0.171 | -0.4647 | Yes | ||
| 50 | RAB1B | ENSG00000174903.16 | 27815 | -0.174 | -0.4608 | Yes | ||
| 51 | TUBA3C | ENSG00000198033.12 | 27829 | -0.175 | -0.4546 | Yes | ||
| 52 | ANK3 | ENSG00000151150.22 | 28038 | -0.182 | -0.4529 | Yes | ||
| 53 | USO1 | ENSG00000138768.14 | 28192 | -0.186 | -0.4496 | Yes | ||
| 54 | COPZ1 | ENSG00000111481.10 | 28332 | -0.190 | -0.4459 | Yes | ||
| 55 | ANK2 | ENSG00000145362.19 | 28436 | -0.194 | -0.4412 | Yes | ||
| 56 | DCTN4 | ENSG00000132912.12 | 28497 | -0.197 | -0.4353 | Yes | ||
| 57 | BET1 | ENSG00000105829.13 | 28876 | -0.207 | -0.4367 | Yes | ||
| 58 | SPTBN1 | ENSG00000115306.16 | 29149 | -0.216 | -0.4353 | Yes | ||
| 59 | TUBA3E | ENSG00000152086.9 | 29330 | -0.222 | -0.4313 | Yes | ||
| 60 | SPTAN1 | ENSG00000197694.16 | 29628 | -0.232 | -0.4299 | Yes | ||
| 61 | RAB1A | ENSG00000138069.18 | 30150 | -0.249 | -0.4333 | Yes | ||
| 62 | GBF1 | ENSG00000107862.4 | 30343 | -0.255 | -0.4284 | Yes | ||
| 63 | COG6 | ENSG00000133103.17 | 30452 | -0.258 | -0.4214 | Yes | ||
| 64 | TMED10 | ENSG00000170348.9 | 30459 | -0.259 | -0.4119 | Yes | ||
| 65 | TUBB8 | ENSG00000261456.5 | 30717 | -0.267 | -0.4082 | Yes | ||
| 66 | COPG1 | ENSG00000181789.14 | 30735 | -0.267 | -0.3986 | Yes | ||
| 67 | ARFGAP3 | ENSG00000242247.11 | 30822 | -0.270 | -0.3906 | Yes | ||
| 68 | DCTN5 | ENSG00000166847.10 | 31044 | -0.279 | -0.3856 | Yes | ||
| 69 | TUBB4A | ENSG00000104833.12 | 31106 | -0.281 | -0.3766 | Yes | ||
| 70 | DYNC1H1 | ENSG00000197102.12 | 31822 | -0.290 | -0.3831 | Yes | ||
| 71 | COPB2 | ENSG00000184432.10 | 32009 | -0.297 | -0.3766 | Yes | ||
| 72 | DCTN2 | ENSG00000175203.15 | 32923 | -0.333 | -0.3864 | Yes | ||
| 73 | TUBAL3 | ENSG00000178462.12 | 33185 | -0.336 | -0.3802 | Yes | ||
| 74 | TMED7 | ENSG00000134970.14 | 35148 | -0.371 | -0.4141 | Yes | ||
| 75 | CAPZB | ENSG00000077549.18 | 35414 | -0.384 | -0.4062 | Yes | ||
| 76 | DCTN1 | ENSG00000204843.12 | 35697 | -0.397 | -0.3982 | Yes | ||
| 77 | DYNC1I2 | ENSG00000077380.15 | 36153 | -0.411 | -0.3939 | Yes | ||
| 78 | DCTN6 | ENSG00000104671.8 | 36330 | -0.417 | -0.3826 | Yes | ||
| 79 | NSF | ENSG00000073969.18 | 36513 | -0.428 | -0.3710 | Yes | ||
| 80 | ARCN1 | ENSG00000095139.14 | 36550 | -0.430 | -0.3558 | Yes | ||
| 81 | COPB1 | ENSG00000129083.12 | 36728 | -0.437 | -0.3438 | Yes | ||
| 82 | ACTR1A | ENSG00000138107.13 | 36970 | -0.450 | -0.3329 | Yes | ||
| 83 | TUBA1B | ENSG00000123416.15 | 37411 | -0.478 | -0.3257 | Yes | ||
| 84 | ARF4 | ENSG00000168374.11 | 37528 | -0.487 | -0.3103 | Yes | ||
| 85 | CAPZA2 | ENSG00000198898.14 | 38185 | -0.541 | -0.3061 | Yes | ||
| 86 | DYNC1LI1 | ENSG00000144635.9 | 38271 | -0.548 | -0.2877 | Yes | ||
| 87 | DYNLL1 | ENSG00000088986.11 | 38317 | -0.554 | -0.2681 | Yes | ||
| 88 | TUBA1A | ENSG00000167552.14 | 38598 | -0.561 | -0.2539 | Yes | ||
| 89 | COPZ2 | ENSG00000005243.10 | 39066 | -0.608 | -0.2426 | Yes | ||
| 90 | CAPZA1 | ENSG00000116489.13 | 39314 | -0.624 | -0.2253 | Yes | ||
| 91 | ACTR10 | ENSG00000131966.14 | 39413 | -0.627 | -0.2042 | Yes | ||
| 92 | COG5 | ENSG00000164597.13 | 39556 | -0.642 | -0.1837 | Yes | ||
| 93 | COPG2 | ENSG00000158623.14 | 40017 | -0.716 | -0.1682 | Yes | ||
| 94 | TUBB4B | ENSG00000188229.6 | 40106 | -0.733 | -0.1429 | Yes | ||
| 95 | TUBB2B | ENSG00000137285.10 | 40109 | -0.733 | -0.1156 | Yes | ||
| 96 | TUBB2A | ENSG00000137267.6 | 40266 | -0.768 | -0.0907 | Yes | ||
| 97 | TUBA1C | ENSG00000167553.15 | 40512 | -0.843 | -0.0651 | Yes | ||
| 98 | TUBB6 | ENSG00000176014.13 | 40817 | -0.979 | -0.0359 | Yes | ||
| 99 | SPTB | ENSG00000070182.21 | 41024 | -1.185 | 0.0033 | Yes |