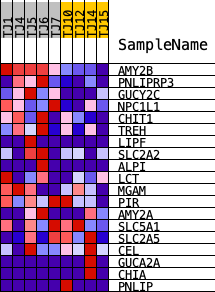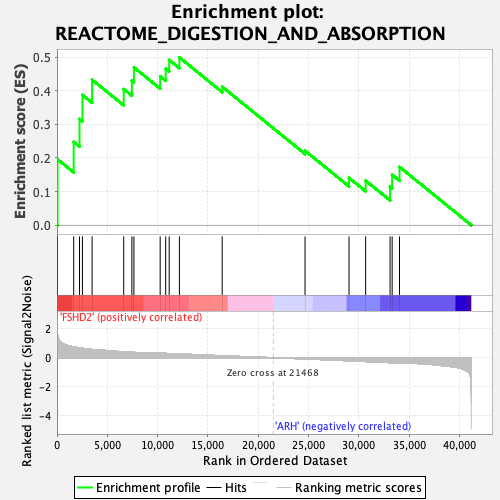
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_DIGESTION_AND_ABSORPTION |
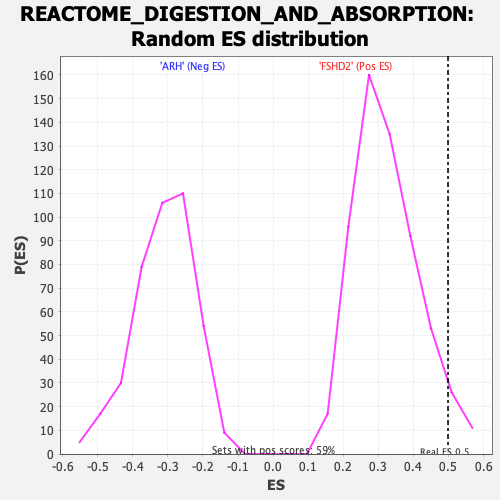
| Enrichment Score (ES) | 0.4987871 |
| Normalized Enrichment Score (NES) | 1.5430739 |
| Nominal p-value | 0.050847456 |
| FDR q-value | 0.2254824 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AMY2B | ENSG00000240038.6 | 50 | 1.633 | 0.1959 | Yes | ||
| 2 | PNLIPRP3 | ENSG00000203837.5 | 1658 | 0.752 | 0.2476 | Yes | ||
| 3 | GUCY2C | ENSG00000070019.5 | 2238 | 0.680 | 0.3156 | Yes | ||
| 4 | NPC1L1 | ENSG00000015520.14 | 2525 | 0.651 | 0.3873 | Yes | ||
| 5 | CHIT1 | ENSG00000133063.16 | 3487 | 0.568 | 0.4325 | Yes | ||
| 6 | TREH | ENSG00000118094.12 | 6628 | 0.402 | 0.4047 | Yes | ||
| 7 | LIPF | ENSG00000182333.14 | 7440 | 0.372 | 0.4299 | Yes | ||
| 8 | SLC2A2 | ENSG00000163581.14 | 7642 | 0.365 | 0.4691 | Yes | ||
| 9 | ALPI | ENSG00000163295.5 | 10250 | 0.305 | 0.4426 | Yes | ||
| 10 | LCT | ENSG00000115850.10 | 10798 | 0.299 | 0.4653 | Yes | ||
| 11 | MGAM | ENSG00000257335.8 | 11146 | 0.286 | 0.4914 | Yes | ||
| 12 | PIR | ENSG00000087842.11 | 12161 | 0.265 | 0.4988 | Yes | ||
| 13 | AMY2A | ENSG00000243480.7 | 16412 | 0.133 | 0.4116 | No | ||
| 14 | SLC5A1 | ENSG00000100170.10 | 24643 | -0.081 | 0.2213 | No | ||
| 15 | SLC2A5 | ENSG00000142583.18 | 29013 | -0.212 | 0.1408 | No | ||
| 16 | CEL | ENSG00000170835.14 | 30665 | -0.265 | 0.1326 | No | ||
| 17 | GUCA2A | ENSG00000197273.4 | 33091 | -0.336 | 0.1142 | No | ||
| 18 | CHIA | ENSG00000134216.19 | 33299 | -0.336 | 0.1498 | No | ||
| 19 | PNLIP | ENSG00000175535.6 | 34030 | -0.342 | 0.1733 | No |