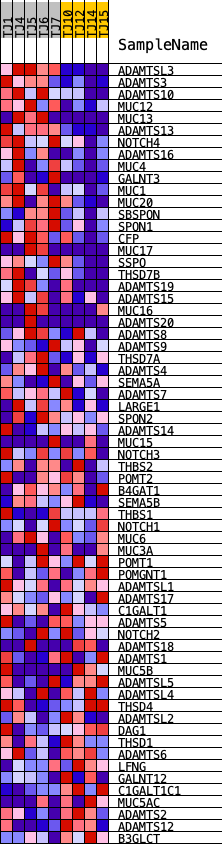
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
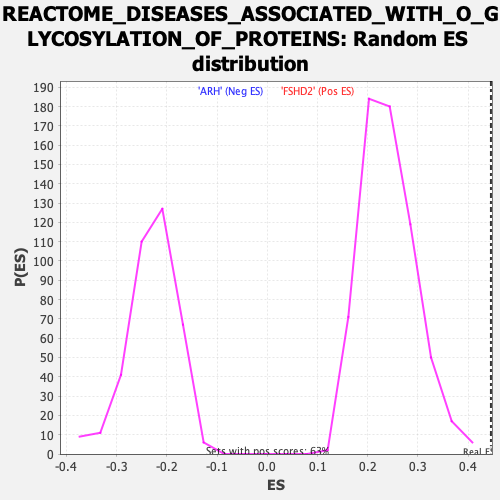
| GeneSet | REACTOME_DISEASES_ASSOCIATED_WITH_O_GLYCOSYLATION_OF_PROTEINS |
| Enrichment Score (ES) | 0.44522503 |
| Normalized Enrichment Score (NES) | 1.846524 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07447787 |
| FWER p-Value | 0.71 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADAMTSL3 | ENSG00000156218.13 | 3 | 2.451 | 0.0940 | Yes | ||
| 2 | ADAMTS3 | ENSG00000156140.10 | 329 | 1.107 | 0.1285 | Yes | ||
| 3 | ADAMTS10 | ENSG00000142303.14 | 413 | 1.054 | 0.1670 | Yes | ||
| 4 | MUC12 | ENSG00000205277.9 | 488 | 1.021 | 0.2043 | Yes | ||
| 5 | MUC13 | ENSG00000173702.7 | 724 | 0.933 | 0.2344 | Yes | ||
| 6 | ADAMTS13 | ENSG00000160323.18 | 974 | 0.865 | 0.2615 | Yes | ||
| 7 | NOTCH4 | ENSG00000204301.6 | 2057 | 0.698 | 0.2620 | Yes | ||
| 8 | ADAMTS16 | ENSG00000145536.15 | 2139 | 0.689 | 0.2865 | Yes | ||
| 9 | MUC4 | ENSG00000145113.22 | 2316 | 0.671 | 0.3079 | Yes | ||
| 10 | GALNT3 | ENSG00000115339.14 | 2460 | 0.658 | 0.3297 | Yes | ||
| 11 | MUC1 | ENSG00000185499.16 | 2664 | 0.637 | 0.3492 | Yes | ||
| 12 | MUC20 | ENSG00000176945.17 | 2795 | 0.623 | 0.3699 | Yes | ||
| 13 | SBSPON | ENSG00000164764.11 | 3341 | 0.581 | 0.3789 | Yes | ||
| 14 | SPON1 | ENSG00000262655.4 | 3535 | 0.565 | 0.3959 | Yes | ||
| 15 | CFP | ENSG00000126759.13 | 3748 | 0.554 | 0.4120 | Yes | ||
| 16 | MUC17 | ENSG00000169876.14 | 4070 | 0.538 | 0.4249 | Yes | ||
| 17 | SSPO | ENSG00000197558.12 | 4521 | 0.518 | 0.4338 | Yes | ||
| 18 | THSD7B | ENSG00000144229.12 | 4843 | 0.501 | 0.4452 | Yes | ||
| 19 | ADAMTS19 | ENSG00000145808.10 | 6613 | 0.403 | 0.4176 | No | ||
| 20 | ADAMTS15 | ENSG00000166106.3 | 8807 | 0.343 | 0.3775 | No | ||
| 21 | MUC16 | ENSG00000181143.15 | 8966 | 0.337 | 0.3865 | No | ||
| 22 | ADAMTS20 | ENSG00000173157.17 | 9514 | 0.319 | 0.3855 | No | ||
| 23 | ADAMTS8 | ENSG00000134917.10 | 10081 | 0.309 | 0.3836 | No | ||
| 24 | ADAMTS9 | ENSG00000163638.13 | 11107 | 0.287 | 0.3696 | No | ||
| 25 | THSD7A | ENSG00000005108.16 | 11769 | 0.279 | 0.3643 | No | ||
| 26 | ADAMTS4 | ENSG00000158859.10 | 12138 | 0.266 | 0.3655 | No | ||
| 27 | SEMA5A | ENSG00000112902.12 | 13382 | 0.226 | 0.3439 | No | ||
| 28 | ADAMTS7 | ENSG00000136378.15 | 13412 | 0.225 | 0.3519 | No | ||
| 29 | LARGE1 | ENSG00000133424.20 | 13853 | 0.211 | 0.3492 | No | ||
| 30 | SPON2 | ENSG00000159674.12 | 15536 | 0.158 | 0.3144 | No | ||
| 31 | ADAMTS14 | ENSG00000138316.11 | 17197 | 0.112 | 0.2783 | No | ||
| 32 | MUC15 | ENSG00000169550.14 | 17306 | 0.110 | 0.2799 | No | ||
| 33 | NOTCH3 | ENSG00000074181.9 | 17558 | 0.102 | 0.2777 | No | ||
| 34 | THBS2 | ENSG00000186340.15 | 19463 | 0.052 | 0.2333 | No | ||
| 35 | POMT2 | ENSG00000009830.11 | 20787 | 0.018 | 0.2018 | No | ||
| 36 | B4GAT1 | ENSG00000174684.7 | 20914 | 0.014 | 0.1993 | No | ||
| 37 | SEMA5B | ENSG00000082684.16 | 21095 | 0.009 | 0.1953 | No | ||
| 38 | THBS1 | ENSG00000137801.11 | 22033 | -0.015 | 0.1731 | No | ||
| 39 | NOTCH1 | ENSG00000148400.12 | 22768 | -0.032 | 0.1565 | No | ||
| 40 | MUC6 | ENSG00000184956.16 | 22845 | -0.034 | 0.1559 | No | ||
| 41 | MUC3A | ENSG00000169894.18 | 23648 | -0.057 | 0.1386 | No | ||
| 42 | POMT1 | ENSG00000130714.16 | 25396 | -0.102 | 0.1000 | No | ||
| 43 | POMGNT1 | ENSG00000085998.14 | 26001 | -0.120 | 0.0899 | No | ||
| 44 | ADAMTSL1 | ENSG00000178031.17 | 27031 | -0.150 | 0.0706 | No | ||
| 45 | ADAMTS17 | ENSG00000140470.14 | 27067 | -0.151 | 0.0756 | No | ||
| 46 | C1GALT1 | ENSG00000106392.10 | 27118 | -0.153 | 0.0802 | No | ||
| 47 | ADAMTS5 | ENSG00000154736.6 | 27545 | -0.166 | 0.0762 | No | ||
| 48 | NOTCH2 | ENSG00000134250.20 | 28133 | -0.185 | 0.0690 | No | ||
| 49 | ADAMTS18 | ENSG00000140873.16 | 29086 | -0.215 | 0.0541 | No | ||
| 50 | ADAMTS1 | ENSG00000154734.15 | 29777 | -0.237 | 0.0464 | No | ||
| 51 | MUC5B | ENSG00000117983.17 | 30117 | -0.248 | 0.0477 | No | ||
| 52 | ADAMTSL5 | ENSG00000185761.11 | 30748 | -0.268 | 0.0426 | No | ||
| 53 | ADAMTSL4 | ENSG00000143382.14 | 30945 | -0.275 | 0.0484 | No | ||
| 54 | THSD4 | ENSG00000187720.14 | 31761 | -0.288 | 0.0396 | No | ||
| 55 | ADAMTSL2 | ENSG00000197859.10 | 32179 | -0.303 | 0.0411 | No | ||
| 56 | DAG1 | ENSG00000173402.11 | 32891 | -0.332 | 0.0365 | No | ||
| 57 | THSD1 | ENSG00000136114.17 | 34085 | -0.344 | 0.0207 | No | ||
| 58 | ADAMTS6 | ENSG00000049192.15 | 35718 | -0.398 | -0.0037 | No | ||
| 59 | LFNG | ENSG00000106003.13 | 36633 | -0.433 | -0.0094 | No | ||
| 60 | GALNT12 | ENSG00000119514.7 | 37272 | -0.469 | -0.0069 | No | ||
| 61 | C1GALT1C1 | ENSG00000171155.8 | 37611 | -0.494 | 0.0038 | No | ||
| 62 | MUC5AC | ENSG00000215182.8 | 37754 | -0.508 | 0.0199 | No | ||
| 63 | ADAMTS2 | ENSG00000087116.16 | 37945 | -0.524 | 0.0353 | No | ||
| 64 | ADAMTS12 | ENSG00000151388.11 | 38028 | -0.530 | 0.0537 | No | ||
| 65 | B3GLCT | ENSG00000187676.8 | 38874 | -0.587 | 0.0556 | No |