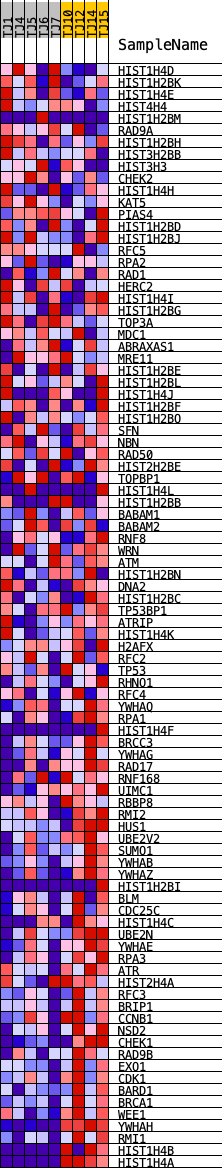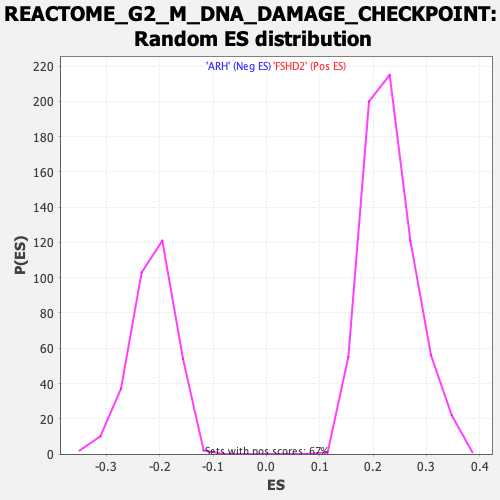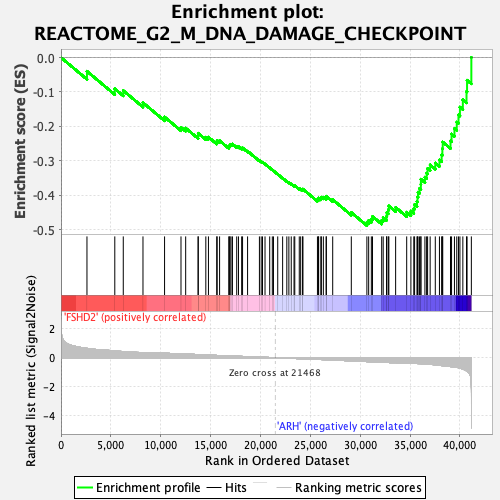
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_G2_M_DNA_DAMAGE_CHECKPOINT |
| Enrichment Score (ES) | -0.48816934 |
| Normalized Enrichment Score (NES) | -2.2719893 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.215072E-4 |
| FWER p-Value | 0.007 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HIST1H4D | ENSG00000277157.1 | 2603 | 0.643 | -0.0398 | No | ||
| 2 | HIST1H2BK | ENSG00000197903.7 | 5394 | 0.465 | -0.0907 | No | ||
| 3 | HIST1H4E | ENSG00000276966.2 | 6244 | 0.421 | -0.0960 | No | ||
| 4 | HIST4H4 | ENSG00000197837.3 | 8220 | 0.353 | -0.1311 | No | ||
| 5 | HIST1H2BM | ENSG00000273703.1 | 10385 | 0.305 | -0.1727 | No | ||
| 6 | RAD9A | ENSG00000172613.8 | 12032 | 0.270 | -0.2029 | No | ||
| 7 | HIST1H2BH | ENSG00000275713.2 | 12502 | 0.254 | -0.2050 | No | ||
| 8 | HIST3H2BB | ENSG00000196890.4 | 13749 | 0.214 | -0.2275 | No | ||
| 9 | HIST3H3 | ENSG00000168148.3 | 13761 | 0.214 | -0.2199 | No | ||
| 10 | CHEK2 | ENSG00000183765.22 | 14522 | 0.190 | -0.2314 | No | ||
| 11 | HIST1H4H | ENSG00000158406.4 | 14785 | 0.181 | -0.2312 | No | ||
| 12 | KAT5 | ENSG00000172977.13 | 15617 | 0.156 | -0.2457 | No | ||
| 13 | PIAS4 | ENSG00000105229.7 | 15657 | 0.155 | -0.2410 | No | ||
| 14 | HIST1H2BD | ENSG00000158373.8 | 15903 | 0.147 | -0.2416 | No | ||
| 15 | HIST1H2BJ | ENSG00000124635.8 | 16824 | 0.122 | -0.2595 | No | ||
| 16 | RFC5 | ENSG00000111445.14 | 16883 | 0.120 | -0.2565 | No | ||
| 17 | RPA2 | ENSG00000117748.10 | 16945 | 0.118 | -0.2537 | No | ||
| 18 | RAD1 | ENSG00000113456.19 | 17050 | 0.116 | -0.2519 | No | ||
| 19 | HERC2 | ENSG00000128731.17 | 17193 | 0.112 | -0.2513 | No | ||
| 20 | HIST1H4I | ENSG00000276180.1 | 17620 | 0.100 | -0.2580 | No | ||
| 21 | HIST1H2BG | ENSG00000273802.2 | 17796 | 0.096 | -0.2587 | No | ||
| 22 | TOP3A | ENSG00000177302.15 | 18116 | 0.088 | -0.2633 | No | ||
| 23 | MDC1 | ENSG00000137337.15 | 18206 | 0.085 | -0.2623 | No | ||
| 24 | ABRAXAS1 | ENSG00000163322.14 | 18710 | 0.073 | -0.2719 | No | ||
| 25 | MRE11 | ENSG00000020922.12 | 19896 | 0.041 | -0.2992 | No | ||
| 26 | HIST1H2BE | ENSG00000274290.2 | 20074 | 0.038 | -0.3022 | No | ||
| 27 | HIST1H2BL | ENSG00000185130.5 | 20206 | 0.034 | -0.3041 | No | ||
| 28 | HIST1H4J | ENSG00000197238.4 | 20458 | 0.026 | -0.3093 | No | ||
| 29 | HIST1H2BF | ENSG00000277224.2 | 20928 | 0.014 | -0.3202 | No | ||
| 30 | HIST1H2BO | ENSG00000274641.1 | 21190 | 0.007 | -0.3263 | No | ||
| 31 | SFN | ENSG00000175793.12 | 21306 | 0.004 | -0.3289 | No | ||
| 32 | NBN | ENSG00000104320.13 | 21733 | -0.007 | -0.3391 | No | ||
| 33 | RAD50 | ENSG00000113522.14 | 22224 | -0.019 | -0.3503 | No | ||
| 34 | HIST2H2BE | ENSG00000184678.10 | 22644 | -0.030 | -0.3594 | No | ||
| 35 | TOPBP1 | ENSG00000163781.14 | 22850 | -0.035 | -0.3631 | No | ||
| 36 | HIST1H4L | ENSG00000275126.1 | 23088 | -0.042 | -0.3674 | No | ||
| 37 | HIST1H2BB | ENSG00000276410.3 | 23366 | -0.049 | -0.3723 | No | ||
| 38 | BABAM1 | ENSG00000105393.16 | 23442 | -0.051 | -0.3723 | No | ||
| 39 | BABAM2 | ENSG00000158019.20 | 23893 | -0.063 | -0.3809 | No | ||
| 40 | RNF8 | ENSG00000112130.17 | 24033 | -0.065 | -0.3819 | No | ||
| 41 | WRN | ENSG00000165392.10 | 24224 | -0.071 | -0.3840 | No | ||
| 42 | ATM | ENSG00000149311.18 | 24258 | -0.072 | -0.3822 | No | ||
| 43 | HIST1H2BN | ENSG00000233822.4 | 25715 | -0.111 | -0.4135 | No | ||
| 44 | DNA2 | ENSG00000138346.15 | 25812 | -0.114 | -0.4117 | No | ||
| 45 | HIST1H2BC | ENSG00000180596.7 | 25836 | -0.115 | -0.4081 | No | ||
| 46 | TP53BP1 | ENSG00000067369.13 | 26057 | -0.123 | -0.4089 | No | ||
| 47 | ATRIP | ENSG00000164053.21 | 26108 | -0.124 | -0.4056 | No | ||
| 48 | HIST1H4K | ENSG00000273542.1 | 26308 | -0.129 | -0.4057 | No | ||
| 49 | H2AFX | ENSG00000188486.3 | 26590 | -0.138 | -0.4075 | No | ||
| 50 | RFC2 | ENSG00000049541.11 | 26606 | -0.138 | -0.4028 | No | ||
| 51 | TP53 | ENSG00000141510.17 | 27245 | -0.156 | -0.4127 | No | ||
| 52 | RHNO1 | ENSG00000171792.11 | 29110 | -0.215 | -0.4502 | No | ||
| 53 | RFC4 | ENSG00000163918.10 | 30672 | -0.265 | -0.4785 | Yes | ||
| 54 | YWHAQ | ENSG00000134308.14 | 30852 | -0.271 | -0.4729 | Yes | ||
| 55 | RPA1 | ENSG00000132383.12 | 31119 | -0.282 | -0.4691 | Yes | ||
| 56 | HIST1H4F | ENSG00000274618.1 | 31237 | -0.285 | -0.4615 | Yes | ||
| 57 | BRCC3 | ENSG00000185515.14 | 32158 | -0.302 | -0.4728 | Yes | ||
| 58 | YWHAG | ENSG00000170027.7 | 32308 | -0.310 | -0.4651 | Yes | ||
| 59 | RAD17 | ENSG00000152942.19 | 32645 | -0.322 | -0.4615 | Yes | ||
| 60 | RNF168 | ENSG00000163961.4 | 32669 | -0.323 | -0.4502 | Yes | ||
| 61 | UIMC1 | ENSG00000087206.17 | 32823 | -0.329 | -0.4419 | Yes | ||
| 62 | RBBP8 | ENSG00000101773.19 | 32876 | -0.331 | -0.4310 | Yes | ||
| 63 | RMI2 | ENSG00000175643.10 | 33556 | -0.338 | -0.4352 | Yes | ||
| 64 | HUS1 | ENSG00000136273.13 | 34665 | -0.355 | -0.4492 | Yes | ||
| 65 | UBE2V2 | ENSG00000169139.12 | 35086 | -0.368 | -0.4459 | Yes | ||
| 66 | SUMO1 | ENSG00000116030.16 | 35376 | -0.382 | -0.4390 | Yes | ||
| 67 | YWHAB | ENSG00000166913.13 | 35457 | -0.386 | -0.4268 | Yes | ||
| 68 | YWHAZ | ENSG00000164924.18 | 35696 | -0.397 | -0.4180 | Yes | ||
| 69 | HIST1H2BI | ENSG00000278588.1 | 35748 | -0.400 | -0.4046 | Yes | ||
| 70 | BLM | ENSG00000197299.12 | 35803 | -0.402 | -0.3912 | Yes | ||
| 71 | CDC25C | ENSG00000158402.20 | 35954 | -0.403 | -0.3801 | Yes | ||
| 72 | HIST1H4C | ENSG00000197061.4 | 36084 | -0.408 | -0.3682 | Yes | ||
| 73 | UBE2N | ENSG00000177889.10 | 36090 | -0.408 | -0.3534 | Yes | ||
| 74 | YWHAE | ENSG00000108953.17 | 36492 | -0.426 | -0.3476 | Yes | ||
| 75 | RPA3 | ENSG00000106399.11 | 36661 | -0.434 | -0.3358 | Yes | ||
| 76 | ATR | ENSG00000175054.16 | 36761 | -0.438 | -0.3221 | Yes | ||
| 77 | HIST2H4A | ENSG00000270882.2 | 37017 | -0.453 | -0.3118 | Yes | ||
| 78 | RFC3 | ENSG00000133119.13 | 37532 | -0.487 | -0.3064 | Yes | ||
| 79 | BRIP1 | ENSG00000136492.8 | 37949 | -0.524 | -0.2974 | Yes | ||
| 80 | CCNB1 | ENSG00000134057.15 | 38169 | -0.540 | -0.2829 | Yes | ||
| 81 | NSD2 | ENSG00000109685.18 | 38242 | -0.546 | -0.2647 | Yes | ||
| 82 | CHEK1 | ENSG00000149554.13 | 38279 | -0.549 | -0.2455 | Yes | ||
| 83 | RAD9B | ENSG00000151164.18 | 39057 | -0.607 | -0.2422 | Yes | ||
| 84 | EXO1 | ENSG00000174371.17 | 39163 | -0.615 | -0.2222 | Yes | ||
| 85 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.2061 | Yes | ||
| 86 | BARD1 | ENSG00000138376.11 | 39685 | -0.660 | -0.1876 | Yes | ||
| 87 | BRCA1 | ENSG00000012048.22 | 39857 | -0.687 | -0.1666 | Yes | ||
| 88 | WEE1 | ENSG00000166483.11 | 40002 | -0.714 | -0.1439 | Yes | ||
| 89 | YWHAH | ENSG00000128245.15 | 40295 | -0.774 | -0.1227 | Yes | ||
| 90 | RMI1 | ENSG00000178966.16 | 40663 | -0.903 | -0.0986 | Yes | ||
| 91 | HIST1H4B | ENSG00000278705.1 | 40718 | -0.927 | -0.0659 | Yes | ||
| 92 | HIST1H4A | ENSG00000278637.1 | 41148 | -2.093 | 0.0003 | Yes |