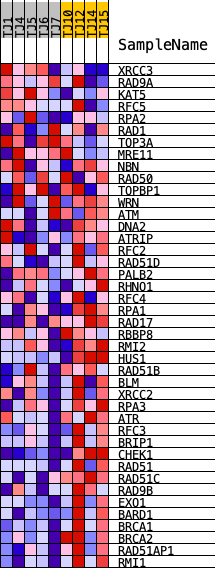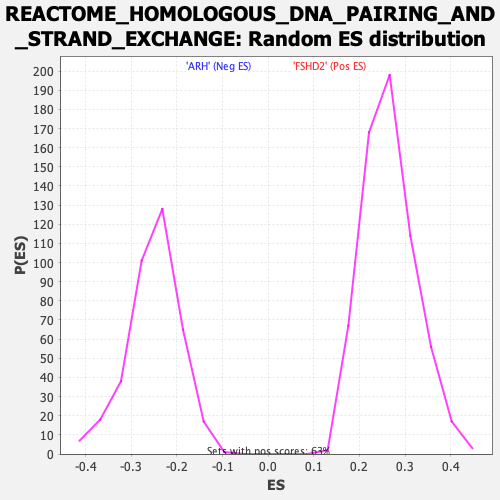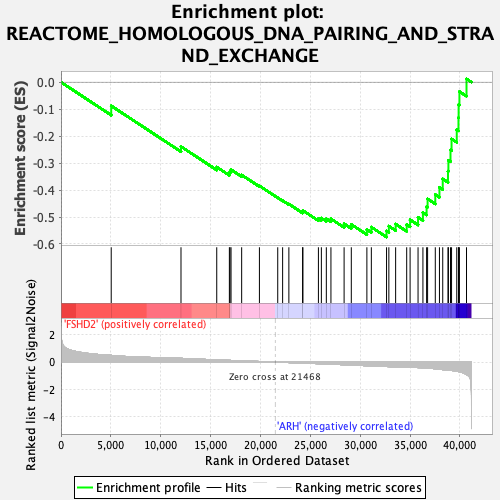
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_HOMOLOGOUS_DNA_PAIRING_AND_STRAND_EXCHANGE |
| Enrichment Score (ES) | -0.5744901 |
| Normalized Enrichment Score (NES) | -2.28841 |
| Nominal p-value | 0.0 |
| FDR q-value | 2.9503606E-4 |
| FWER p-Value | 0.006 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | XRCC3 | ENSG00000126215.14 | 5035 | 0.488 | -0.0877 | No | ||
| 2 | RAD9A | ENSG00000172613.8 | 12032 | 0.270 | -0.2387 | No | ||
| 3 | KAT5 | ENSG00000172977.13 | 15617 | 0.156 | -0.3148 | No | ||
| 4 | RFC5 | ENSG00000111445.14 | 16883 | 0.120 | -0.3370 | No | ||
| 5 | RPA2 | ENSG00000117748.10 | 16945 | 0.118 | -0.3301 | No | ||
| 6 | RAD1 | ENSG00000113456.19 | 17050 | 0.116 | -0.3244 | No | ||
| 7 | TOP3A | ENSG00000177302.15 | 18116 | 0.088 | -0.3440 | No | ||
| 8 | MRE11 | ENSG00000020922.12 | 19896 | 0.041 | -0.3844 | No | ||
| 9 | NBN | ENSG00000104320.13 | 21733 | -0.007 | -0.4285 | No | ||
| 10 | RAD50 | ENSG00000113522.14 | 22224 | -0.019 | -0.4391 | No | ||
| 11 | TOPBP1 | ENSG00000163781.14 | 22850 | -0.035 | -0.4519 | No | ||
| 12 | WRN | ENSG00000165392.10 | 24224 | -0.071 | -0.4803 | No | ||
| 13 | ATM | ENSG00000149311.18 | 24258 | -0.072 | -0.4760 | No | ||
| 14 | DNA2 | ENSG00000138346.15 | 25812 | -0.114 | -0.5056 | No | ||
| 15 | ATRIP | ENSG00000164053.21 | 26108 | -0.124 | -0.5040 | No | ||
| 16 | RFC2 | ENSG00000049541.11 | 26606 | -0.138 | -0.5063 | No | ||
| 17 | RAD51D | ENSG00000185379.20 | 27068 | -0.151 | -0.5068 | No | ||
| 18 | PALB2 | ENSG00000083093.9 | 28388 | -0.192 | -0.5252 | No | ||
| 19 | RHNO1 | ENSG00000171792.11 | 29110 | -0.215 | -0.5274 | No | ||
| 20 | RFC4 | ENSG00000163918.10 | 30672 | -0.265 | -0.5466 | No | ||
| 21 | RPA1 | ENSG00000132383.12 | 31119 | -0.282 | -0.5374 | No | ||
| 22 | RAD17 | ENSG00000152942.19 | 32645 | -0.322 | -0.5516 | Yes | ||
| 23 | RBBP8 | ENSG00000101773.19 | 32876 | -0.331 | -0.5337 | Yes | ||
| 24 | RMI2 | ENSG00000175643.10 | 33556 | -0.338 | -0.5262 | Yes | ||
| 25 | HUS1 | ENSG00000136273.13 | 34665 | -0.355 | -0.5279 | Yes | ||
| 26 | RAD51B | ENSG00000182185.18 | 35006 | -0.365 | -0.5103 | Yes | ||
| 27 | BLM | ENSG00000197299.12 | 35803 | -0.402 | -0.5010 | Yes | ||
| 28 | XRCC2 | ENSG00000196584.3 | 36292 | -0.416 | -0.4834 | Yes | ||
| 29 | RPA3 | ENSG00000106399.11 | 36661 | -0.434 | -0.4615 | Yes | ||
| 30 | ATR | ENSG00000175054.16 | 36761 | -0.438 | -0.4328 | Yes | ||
| 31 | RFC3 | ENSG00000133119.13 | 37532 | -0.487 | -0.4169 | Yes | ||
| 32 | BRIP1 | ENSG00000136492.8 | 37949 | -0.524 | -0.3898 | Yes | ||
| 33 | CHEK1 | ENSG00000149554.13 | 38279 | -0.549 | -0.3587 | Yes | ||
| 34 | RAD51 | ENSG00000051180.17 | 38806 | -0.582 | -0.3302 | Yes | ||
| 35 | RAD51C | ENSG00000108384.15 | 38848 | -0.585 | -0.2896 | Yes | ||
| 36 | RAD9B | ENSG00000151164.18 | 39057 | -0.607 | -0.2516 | Yes | ||
| 37 | EXO1 | ENSG00000174371.17 | 39163 | -0.615 | -0.2104 | Yes | ||
| 38 | BARD1 | ENSG00000138376.11 | 39685 | -0.660 | -0.1762 | Yes | ||
| 39 | BRCA1 | ENSG00000012048.22 | 39857 | -0.687 | -0.1316 | Yes | ||
| 40 | BRCA2 | ENSG00000139618.15 | 39873 | -0.690 | -0.0830 | Yes | ||
| 41 | RAD51AP1 | ENSG00000111247.14 | 39953 | -0.704 | -0.0348 | Yes | ||
| 42 | RMI1 | ENSG00000178966.16 | 40663 | -0.903 | 0.0121 | Yes |