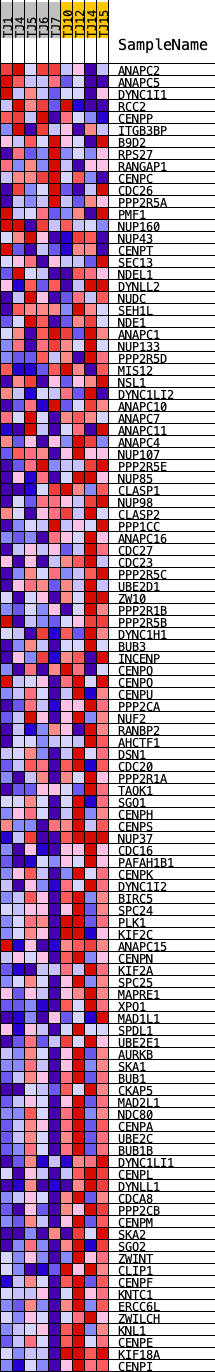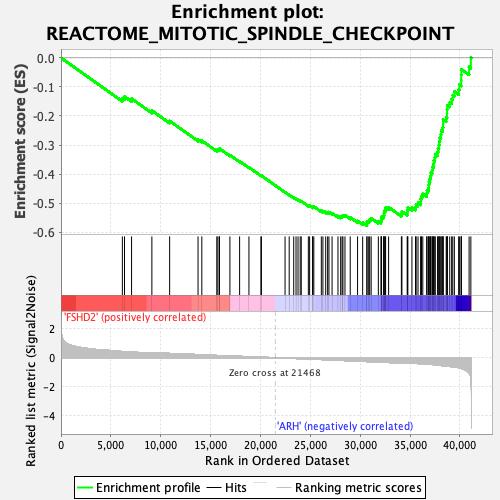
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_MITOTIC_SPINDLE_CHECKPOINT |
| Enrichment Score (ES) | -0.5772716 |
| Normalized Enrichment Score (NES) | -2.8589137 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ANAPC2 | ENSG00000176248.9 | 6153 | 0.426 | -0.1388 | No | ||
| 2 | ANAPC5 | ENSG00000089053.13 | 6368 | 0.414 | -0.1333 | No | ||
| 3 | DYNC1I1 | ENSG00000158560.14 | 7076 | 0.386 | -0.1405 | No | ||
| 4 | RCC2 | ENSG00000179051.14 | 9112 | 0.331 | -0.1815 | No | ||
| 5 | CENPP | ENSG00000188312.14 | 10894 | 0.295 | -0.2172 | No | ||
| 6 | ITGB3BP | ENSG00000142856.16 | 13741 | 0.215 | -0.2809 | No | ||
| 7 | B9D2 | ENSG00000123810.8 | 14121 | 0.202 | -0.2849 | No | ||
| 8 | RPS27 | ENSG00000177954.14 | 15622 | 0.156 | -0.3174 | No | ||
| 9 | RANGAP1 | ENSG00000100401.19 | 15733 | 0.152 | -0.3161 | No | ||
| 10 | CENPC | ENSG00000145241.11 | 15881 | 0.148 | -0.3159 | No | ||
| 11 | CDC26 | ENSG00000176386.9 | 15890 | 0.148 | -0.3122 | No | ||
| 12 | PPP2R5A | ENSG00000066027.12 | 16937 | 0.118 | -0.3346 | No | ||
| 13 | PMF1 | ENSG00000160783.19 | 17911 | 0.093 | -0.3559 | No | ||
| 14 | NUP160 | ENSG00000030066.13 | 18845 | 0.069 | -0.3768 | No | ||
| 15 | NUP43 | ENSG00000120253.14 | 20047 | 0.038 | -0.4051 | No | ||
| 16 | CENPT | ENSG00000102901.13 | 20096 | 0.037 | -0.4053 | No | ||
| 17 | SEC13 | ENSG00000157020.18 | 22468 | -0.026 | -0.4624 | No | ||
| 18 | NDEL1 | ENSG00000166579.15 | 22882 | -0.036 | -0.4715 | No | ||
| 19 | DYNLL2 | ENSG00000264364.3 | 23336 | -0.048 | -0.4813 | No | ||
| 20 | NUDC | ENSG00000090273.14 | 23568 | -0.054 | -0.4855 | No | ||
| 21 | SEH1L | ENSG00000085415.16 | 23768 | -0.059 | -0.4888 | No | ||
| 22 | NDE1 | ENSG00000072864.15 | 23985 | -0.064 | -0.4924 | No | ||
| 23 | ANAPC1 | ENSG00000153107.13 | 24092 | -0.067 | -0.4933 | No | ||
| 24 | NUP133 | ENSG00000069248.12 | 24802 | -0.086 | -0.5083 | No | ||
| 25 | PPP2R5D | ENSG00000112640.15 | 24932 | -0.090 | -0.5092 | No | ||
| 26 | MIS12 | ENSG00000167842.15 | 25231 | -0.097 | -0.5139 | No | ||
| 27 | NSL1 | ENSG00000117697.15 | 25239 | -0.097 | -0.5115 | No | ||
| 28 | DYNC1LI2 | ENSG00000135720.13 | 25352 | -0.101 | -0.5117 | No | ||
| 29 | ANAPC10 | ENSG00000164162.14 | 26104 | -0.124 | -0.5267 | No | ||
| 30 | ANAPC7 | ENSG00000196510.12 | 26263 | -0.128 | -0.5273 | No | ||
| 31 | ANAPC11 | ENSG00000141552.17 | 26543 | -0.136 | -0.5305 | No | ||
| 32 | ANAPC4 | ENSG00000053900.10 | 26737 | -0.142 | -0.5315 | No | ||
| 33 | NUP107 | ENSG00000111581.10 | 26867 | -0.145 | -0.5309 | No | ||
| 34 | PPP2R5E | ENSG00000154001.14 | 27182 | -0.155 | -0.5345 | No | ||
| 35 | NUP85 | ENSG00000125450.11 | 27774 | -0.173 | -0.5444 | No | ||
| 36 | CLASP1 | ENSG00000074054.18 | 28043 | -0.182 | -0.5463 | No | ||
| 37 | NUP98 | ENSG00000110713.17 | 28185 | -0.186 | -0.5449 | No | ||
| 38 | CLASP2 | ENSG00000163539.17 | 28270 | -0.189 | -0.5420 | No | ||
| 39 | PPP1CC | ENSG00000186298.12 | 28475 | -0.195 | -0.5419 | No | ||
| 40 | ANAPC16 | ENSG00000166295.9 | 29001 | -0.212 | -0.5492 | No | ||
| 41 | CDC27 | ENSG00000004897.12 | 29738 | -0.236 | -0.5610 | No | ||
| 42 | CDC23 | ENSG00000094880.11 | 30249 | -0.252 | -0.5669 | No | ||
| 43 | PPP2R5C | ENSG00000078304.19 | 30677 | -0.265 | -0.5704 | Yes | ||
| 44 | UBE2D1 | ENSG00000072401.15 | 30679 | -0.265 | -0.5635 | Yes | ||
| 45 | ZW10 | ENSG00000086827.9 | 30856 | -0.271 | -0.5608 | Yes | ||
| 46 | PPP2R1B | ENSG00000137713.16 | 30954 | -0.276 | -0.5560 | Yes | ||
| 47 | PPP2R5B | ENSG00000068971.14 | 31101 | -0.281 | -0.5522 | Yes | ||
| 48 | DYNC1H1 | ENSG00000197102.12 | 31822 | -0.290 | -0.5622 | Yes | ||
| 49 | BUB3 | ENSG00000154473.18 | 32087 | -0.300 | -0.5609 | Yes | ||
| 50 | INCENP | ENSG00000149503.13 | 32126 | -0.301 | -0.5540 | Yes | ||
| 51 | CENPQ | ENSG00000031691.7 | 32142 | -0.302 | -0.5465 | Yes | ||
| 52 | CENPO | ENSG00000138092.11 | 32352 | -0.312 | -0.5435 | Yes | ||
| 53 | CENPU | ENSG00000151725.12 | 32395 | -0.313 | -0.5364 | Yes | ||
| 54 | PPP2CA | ENSG00000113575.10 | 32408 | -0.314 | -0.5285 | Yes | ||
| 55 | NUF2 | ENSG00000143228.13 | 32503 | -0.317 | -0.5226 | Yes | ||
| 56 | RANBP2 | ENSG00000153201.16 | 32550 | -0.318 | -0.5154 | Yes | ||
| 57 | AHCTF1 | ENSG00000153207.15 | 32851 | -0.330 | -0.5142 | Yes | ||
| 58 | DSN1 | ENSG00000149636.15 | 34130 | -0.346 | -0.5363 | Yes | ||
| 59 | CDC20 | ENSG00000117399.14 | 34187 | -0.349 | -0.5286 | Yes | ||
| 60 | PPP2R1A | ENSG00000105568.18 | 34725 | -0.357 | -0.5324 | Yes | ||
| 61 | TAOK1 | ENSG00000160551.11 | 34732 | -0.358 | -0.5233 | Yes | ||
| 62 | SGO1 | ENSG00000129810.15 | 34782 | -0.360 | -0.5151 | Yes | ||
| 63 | CENPH | ENSG00000153044.10 | 35190 | -0.373 | -0.5153 | Yes | ||
| 64 | CENPS | ENSG00000175279.22 | 35550 | -0.391 | -0.5139 | Yes | ||
| 65 | NUP37 | ENSG00000075188.9 | 35611 | -0.394 | -0.5052 | Yes | ||
| 66 | CDC16 | ENSG00000130177.16 | 35800 | -0.402 | -0.4993 | Yes | ||
| 67 | PAFAH1B1 | ENSG00000007168.13 | 36076 | -0.408 | -0.4954 | Yes | ||
| 68 | CENPK | ENSG00000123219.13 | 36086 | -0.408 | -0.4850 | Yes | ||
| 69 | DYNC1I2 | ENSG00000077380.15 | 36153 | -0.411 | -0.4759 | Yes | ||
| 70 | BIRC5 | ENSG00000089685.15 | 36279 | -0.415 | -0.4682 | Yes | ||
| 71 | SPC24 | ENSG00000161888.11 | 36669 | -0.434 | -0.4664 | Yes | ||
| 72 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.4560 | Yes | ||
| 73 | KIF2C | ENSG00000142945.13 | 36852 | -0.443 | -0.4480 | Yes | ||
| 74 | ANAPC15 | ENSG00000110200.8 | 36886 | -0.445 | -0.4372 | Yes | ||
| 75 | CENPN | ENSG00000166451.13 | 36906 | -0.447 | -0.4261 | Yes | ||
| 76 | KIF2A | ENSG00000068796.16 | 36994 | -0.452 | -0.4165 | Yes | ||
| 77 | SPC25 | ENSG00000152253.9 | 37054 | -0.455 | -0.4061 | Yes | ||
| 78 | MAPRE1 | ENSG00000101367.9 | 37101 | -0.458 | -0.3953 | Yes | ||
| 79 | XPO1 | ENSG00000082898.17 | 37227 | -0.465 | -0.3863 | Yes | ||
| 80 | MAD1L1 | ENSG00000002822.15 | 37265 | -0.468 | -0.3750 | Yes | ||
| 81 | SPDL1 | ENSG00000040275.17 | 37361 | -0.475 | -0.3650 | Yes | ||
| 82 | UBE2E1 | ENSG00000170142.12 | 37384 | -0.477 | -0.3531 | Yes | ||
| 83 | AURKB | ENSG00000178999.13 | 37485 | -0.484 | -0.3430 | Yes | ||
| 84 | SKA1 | ENSG00000154839.10 | 37525 | -0.487 | -0.3313 | Yes | ||
| 85 | BUB1 | ENSG00000169679.15 | 37763 | -0.508 | -0.3239 | Yes | ||
| 86 | CKAP5 | ENSG00000175216.15 | 37814 | -0.514 | -0.3118 | Yes | ||
| 87 | MAD2L1 | ENSG00000164109.14 | 37893 | -0.520 | -0.3002 | Yes | ||
| 88 | NDC80 | ENSG00000080986.13 | 37927 | -0.522 | -0.2874 | Yes | ||
| 89 | CENPA | ENSG00000115163.15 | 37993 | -0.527 | -0.2753 | Yes | ||
| 90 | UBE2C | ENSG00000175063.17 | 38091 | -0.534 | -0.2638 | Yes | ||
| 91 | BUB1B | ENSG00000156970.12 | 38129 | -0.537 | -0.2508 | Yes | ||
| 92 | DYNC1LI1 | ENSG00000144635.9 | 38271 | -0.548 | -0.2400 | Yes | ||
| 93 | CENPL | ENSG00000120334.15 | 38315 | -0.553 | -0.2266 | Yes | ||
| 94 | DYNLL1 | ENSG00000088986.11 | 38317 | -0.554 | -0.2123 | Yes | ||
| 95 | CDCA8 | ENSG00000134690.11 | 38629 | -0.564 | -0.2052 | Yes | ||
| 96 | PPP2CB | ENSG00000104695.13 | 38708 | -0.572 | -0.1922 | Yes | ||
| 97 | CENPM | ENSG00000100162.15 | 38711 | -0.572 | -0.1774 | Yes | ||
| 98 | SKA2 | ENSG00000182628.13 | 38739 | -0.574 | -0.1632 | Yes | ||
| 99 | SGO2 | ENSG00000163535.18 | 38980 | -0.599 | -0.1535 | Yes | ||
| 100 | ZWINT | ENSG00000122952.17 | 39173 | -0.616 | -0.1421 | Yes | ||
| 101 | CLIP1 | ENSG00000130779.20 | 39284 | -0.622 | -0.1287 | Yes | ||
| 102 | CENPF | ENSG00000117724.13 | 39444 | -0.631 | -0.1161 | Yes | ||
| 103 | KNTC1 | ENSG00000184445.12 | 39875 | -0.690 | -0.1087 | Yes | ||
| 104 | ERCC6L | ENSG00000186871.7 | 39947 | -0.704 | -0.0922 | Yes | ||
| 105 | ZWILCH | ENSG00000174442.12 | 40134 | -0.739 | -0.0775 | Yes | ||
| 106 | KNL1 | ENSG00000137812.19 | 40136 | -0.739 | -0.0583 | Yes | ||
| 107 | CENPE | ENSG00000138778.12 | 40145 | -0.740 | -0.0393 | Yes | ||
| 108 | KIF18A | ENSG00000121621.7 | 40921 | -1.038 | -0.0312 | Yes | ||
| 109 | CENPI | ENSG00000102384.13 | 41099 | -1.425 | 0.0015 | Yes |