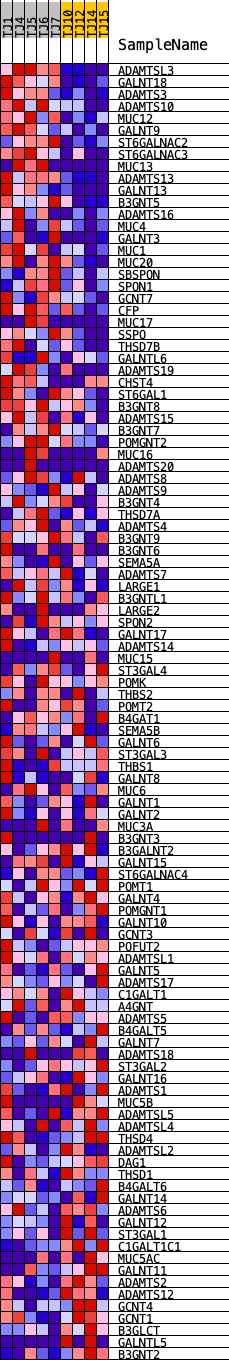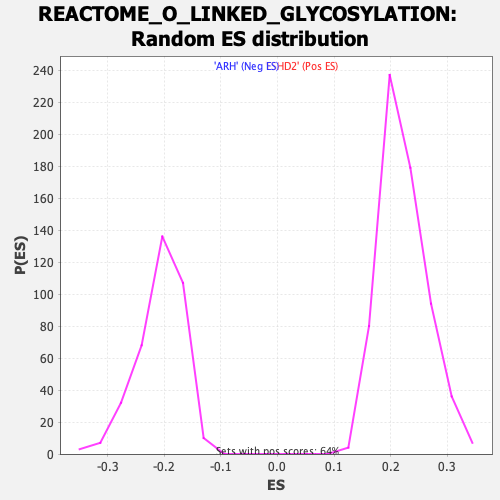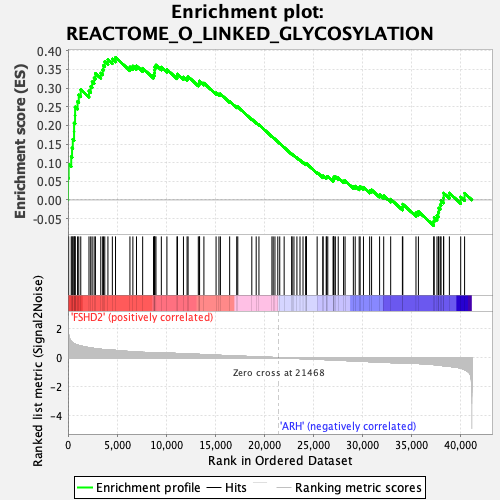
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_O_LINKED_GLYCOSYLATION |
| Enrichment Score (ES) | 0.38226092 |
| Normalized Enrichment Score (NES) | 1.720815 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14910132 |
| FWER p-Value | 0.981 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ADAMTSL3 | ENSG00000156218.13 | 3 | 2.451 | 0.0594 | Yes | ||
| 2 | GALNT18 | ENSG00000110328.6 | 68 | 1.559 | 0.0956 | Yes | ||
| 3 | ADAMTS3 | ENSG00000156140.10 | 329 | 1.107 | 0.1161 | Yes | ||
| 4 | ADAMTS10 | ENSG00000142303.14 | 413 | 1.054 | 0.1397 | Yes | ||
| 5 | MUC12 | ENSG00000205277.9 | 488 | 1.021 | 0.1626 | Yes | ||
| 6 | GALNT9 | ENSG00000182870.13 | 618 | 0.971 | 0.1830 | Yes | ||
| 7 | ST6GALNAC2 | ENSG00000070731.11 | 621 | 0.969 | 0.2065 | Yes | ||
| 8 | ST6GALNAC3 | ENSG00000184005.11 | 719 | 0.934 | 0.2268 | Yes | ||
| 9 | MUC13 | ENSG00000173702.7 | 724 | 0.933 | 0.2493 | Yes | ||
| 10 | ADAMTS13 | ENSG00000160323.18 | 974 | 0.865 | 0.2642 | Yes | ||
| 11 | GALNT13 | ENSG00000144278.15 | 1098 | 0.838 | 0.2816 | Yes | ||
| 12 | B3GNT5 | ENSG00000176597.12 | 1301 | 0.802 | 0.2961 | Yes | ||
| 13 | ADAMTS16 | ENSG00000145536.15 | 2139 | 0.689 | 0.2924 | Yes | ||
| 14 | MUC4 | ENSG00000145113.22 | 2316 | 0.671 | 0.3044 | Yes | ||
| 15 | GALNT3 | ENSG00000115339.14 | 2460 | 0.658 | 0.3169 | Yes | ||
| 16 | MUC1 | ENSG00000185499.16 | 2664 | 0.637 | 0.3274 | Yes | ||
| 17 | MUC20 | ENSG00000176945.17 | 2795 | 0.623 | 0.3393 | Yes | ||
| 18 | SBSPON | ENSG00000164764.11 | 3341 | 0.581 | 0.3401 | Yes | ||
| 19 | SPON1 | ENSG00000262655.4 | 3535 | 0.565 | 0.3491 | Yes | ||
| 20 | GCNT7 | ENSG00000124091.9 | 3615 | 0.563 | 0.3609 | Yes | ||
| 21 | CFP | ENSG00000126759.13 | 3748 | 0.554 | 0.3711 | Yes | ||
| 22 | MUC17 | ENSG00000169876.14 | 4070 | 0.538 | 0.3763 | Yes | ||
| 23 | SSPO | ENSG00000197558.12 | 4521 | 0.518 | 0.3779 | Yes | ||
| 24 | THSD7B | ENSG00000144229.12 | 4843 | 0.501 | 0.3823 | Yes | ||
| 25 | GALNTL6 | ENSG00000174473.16 | 6313 | 0.416 | 0.3566 | No | ||
| 26 | ADAMTS19 | ENSG00000145808.10 | 6613 | 0.403 | 0.3591 | No | ||
| 27 | CHST4 | ENSG00000140835.9 | 6981 | 0.390 | 0.3596 | No | ||
| 28 | ST6GAL1 | ENSG00000073849.15 | 7610 | 0.366 | 0.3532 | No | ||
| 29 | B3GNT8 | ENSG00000177191.2 | 8714 | 0.347 | 0.3347 | No | ||
| 30 | ADAMTS15 | ENSG00000166106.3 | 8807 | 0.343 | 0.3408 | No | ||
| 31 | B3GNT7 | ENSG00000156966.7 | 8836 | 0.342 | 0.3484 | No | ||
| 32 | POMGNT2 | ENSG00000144647.6 | 8847 | 0.342 | 0.3565 | No | ||
| 33 | MUC16 | ENSG00000181143.15 | 8966 | 0.337 | 0.3618 | No | ||
| 34 | ADAMTS20 | ENSG00000173157.17 | 9514 | 0.319 | 0.3562 | No | ||
| 35 | ADAMTS8 | ENSG00000134917.10 | 10081 | 0.309 | 0.3499 | No | ||
| 36 | ADAMTS9 | ENSG00000163638.13 | 11107 | 0.287 | 0.3319 | No | ||
| 37 | B3GNT4 | ENSG00000176383.9 | 11159 | 0.285 | 0.3376 | No | ||
| 38 | THSD7A | ENSG00000005108.16 | 11769 | 0.279 | 0.3295 | No | ||
| 39 | ADAMTS4 | ENSG00000158859.10 | 12138 | 0.266 | 0.3270 | No | ||
| 40 | B3GNT9 | ENSG00000237172.4 | 12241 | 0.263 | 0.3309 | No | ||
| 41 | B3GNT6 | ENSG00000198488.10 | 13282 | 0.229 | 0.3111 | No | ||
| 42 | SEMA5A | ENSG00000112902.12 | 13382 | 0.226 | 0.3142 | No | ||
| 43 | ADAMTS7 | ENSG00000136378.15 | 13412 | 0.225 | 0.3189 | No | ||
| 44 | LARGE1 | ENSG00000133424.20 | 13853 | 0.211 | 0.3133 | No | ||
| 45 | B3GNTL1 | ENSG00000175711.8 | 15093 | 0.171 | 0.2873 | No | ||
| 46 | LARGE2 | ENSG00000165905.18 | 15387 | 0.162 | 0.2841 | No | ||
| 47 | SPON2 | ENSG00000159674.12 | 15536 | 0.158 | 0.2843 | No | ||
| 48 | GALNT17 | ENSG00000185274.12 | 16478 | 0.131 | 0.2646 | No | ||
| 49 | ADAMTS14 | ENSG00000138316.11 | 17197 | 0.112 | 0.2498 | No | ||
| 50 | MUC15 | ENSG00000169550.14 | 17306 | 0.110 | 0.2498 | No | ||
| 51 | ST3GAL4 | ENSG00000110080.18 | 18736 | 0.072 | 0.2168 | No | ||
| 52 | POMK | ENSG00000185900.10 | 19182 | 0.060 | 0.2074 | No | ||
| 53 | THBS2 | ENSG00000186340.15 | 19463 | 0.052 | 0.2018 | No | ||
| 54 | POMT2 | ENSG00000009830.11 | 20787 | 0.018 | 0.1701 | No | ||
| 55 | B4GAT1 | ENSG00000174684.7 | 20914 | 0.014 | 0.1673 | No | ||
| 56 | SEMA5B | ENSG00000082684.16 | 21095 | 0.009 | 0.1632 | No | ||
| 57 | GALNT6 | ENSG00000139629.16 | 21384 | 0.002 | 0.1562 | No | ||
| 58 | ST3GAL3 | ENSG00000126091.20 | 21583 | -0.004 | 0.1515 | No | ||
| 59 | THBS1 | ENSG00000137801.11 | 22033 | -0.015 | 0.1409 | No | ||
| 60 | GALNT8 | ENSG00000130035.8 | 22800 | -0.033 | 0.1230 | No | ||
| 61 | MUC6 | ENSG00000184956.16 | 22845 | -0.034 | 0.1228 | No | ||
| 62 | GALNT1 | ENSG00000141429.13 | 23031 | -0.040 | 0.1193 | No | ||
| 63 | GALNT2 | ENSG00000143641.10 | 23333 | -0.048 | 0.1131 | No | ||
| 64 | MUC3A | ENSG00000169894.18 | 23648 | -0.057 | 0.1068 | No | ||
| 65 | B3GNT3 | ENSG00000179913.11 | 23955 | -0.064 | 0.1009 | No | ||
| 66 | B3GALNT2 | ENSG00000162885.13 | 24225 | -0.071 | 0.0961 | No | ||
| 67 | GALNT15 | ENSG00000131386.19 | 24287 | -0.073 | 0.0963 | No | ||
| 68 | ST6GALNAC4 | ENSG00000136840.19 | 24307 | -0.073 | 0.0977 | No | ||
| 69 | POMT1 | ENSG00000130714.16 | 25396 | -0.102 | 0.0736 | No | ||
| 70 | GALNT4 | ENSG00000257594.4 | 25976 | -0.120 | 0.0624 | No | ||
| 71 | POMGNT1 | ENSG00000085998.14 | 26001 | -0.120 | 0.0648 | No | ||
| 72 | GALNT10 | ENSG00000164574.16 | 26318 | -0.129 | 0.0602 | No | ||
| 73 | GCNT3 | ENSG00000140297.13 | 26366 | -0.131 | 0.0622 | No | ||
| 74 | POFUT2 | ENSG00000186866.16 | 26480 | -0.134 | 0.0627 | No | ||
| 75 | ADAMTSL1 | ENSG00000178031.17 | 27031 | -0.150 | 0.0530 | No | ||
| 76 | GALNT5 | ENSG00000136542.9 | 27046 | -0.151 | 0.0563 | No | ||
| 77 | ADAMTS17 | ENSG00000140470.14 | 27067 | -0.151 | 0.0595 | No | ||
| 78 | C1GALT1 | ENSG00000106392.10 | 27118 | -0.153 | 0.0620 | No | ||
| 79 | A4GNT | ENSG00000118017.4 | 27252 | -0.156 | 0.0625 | No | ||
| 80 | ADAMTS5 | ENSG00000154736.6 | 27545 | -0.166 | 0.0594 | No | ||
| 81 | B4GALT5 | ENSG00000158470.5 | 28090 | -0.184 | 0.0506 | No | ||
| 82 | GALNT7 | ENSG00000109586.12 | 28242 | -0.188 | 0.0515 | No | ||
| 83 | ADAMTS18 | ENSG00000140873.16 | 29086 | -0.215 | 0.0362 | No | ||
| 84 | ST3GAL2 | ENSG00000157350.13 | 29275 | -0.220 | 0.0369 | No | ||
| 85 | GALNT16 | ENSG00000100626.17 | 29679 | -0.233 | 0.0328 | No | ||
| 86 | ADAMTS1 | ENSG00000154734.15 | 29777 | -0.237 | 0.0362 | No | ||
| 87 | MUC5B | ENSG00000117983.17 | 30117 | -0.248 | 0.0339 | No | ||
| 88 | ADAMTSL5 | ENSG00000185761.11 | 30748 | -0.268 | 0.0251 | No | ||
| 89 | ADAMTSL4 | ENSG00000143382.14 | 30945 | -0.275 | 0.0270 | No | ||
| 90 | THSD4 | ENSG00000187720.14 | 31761 | -0.288 | 0.0141 | No | ||
| 91 | ADAMTSL2 | ENSG00000197859.10 | 32179 | -0.303 | 0.0113 | No | ||
| 92 | DAG1 | ENSG00000173402.11 | 32891 | -0.332 | 0.0020 | No | ||
| 93 | THSD1 | ENSG00000136114.17 | 34085 | -0.344 | -0.0187 | No | ||
| 94 | B4GALT6 | ENSG00000118276.11 | 34127 | -0.346 | -0.0113 | No | ||
| 95 | GALNT14 | ENSG00000158089.15 | 35470 | -0.387 | -0.0346 | No | ||
| 96 | ADAMTS6 | ENSG00000049192.15 | 35718 | -0.398 | -0.0310 | No | ||
| 97 | GALNT12 | ENSG00000119514.7 | 37272 | -0.469 | -0.0574 | No | ||
| 98 | ST3GAL1 | ENSG00000008513.16 | 37327 | -0.473 | -0.0472 | No | ||
| 99 | C1GALT1C1 | ENSG00000171155.8 | 37611 | -0.494 | -0.0422 | No | ||
| 100 | MUC5AC | ENSG00000215182.8 | 37754 | -0.508 | -0.0333 | No | ||
| 101 | GALNT11 | ENSG00000178234.13 | 37794 | -0.512 | -0.0218 | No | ||
| 102 | ADAMTS2 | ENSG00000087116.16 | 37945 | -0.524 | -0.0128 | No | ||
| 103 | ADAMTS12 | ENSG00000151388.11 | 38028 | -0.530 | -0.0019 | No | ||
| 104 | GCNT4 | ENSG00000176928.6 | 38284 | -0.550 | 0.0052 | No | ||
| 105 | GCNT1 | ENSG00000187210.14 | 38293 | -0.551 | 0.0183 | No | ||
| 106 | B3GLCT | ENSG00000187676.8 | 38874 | -0.587 | 0.0185 | No | ||
| 107 | GALNTL5 | ENSG00000106648.14 | 40028 | -0.718 | 0.0078 | No | ||
| 108 | B3GNT2 | ENSG00000170340.11 | 40425 | -0.816 | 0.0179 | No |