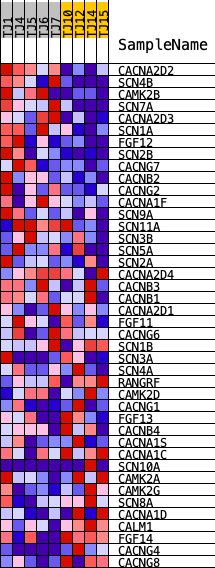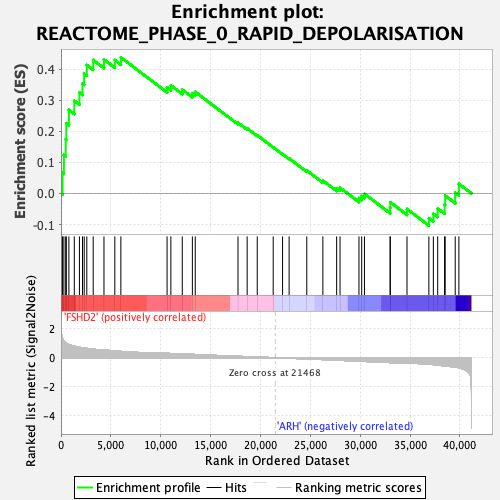
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
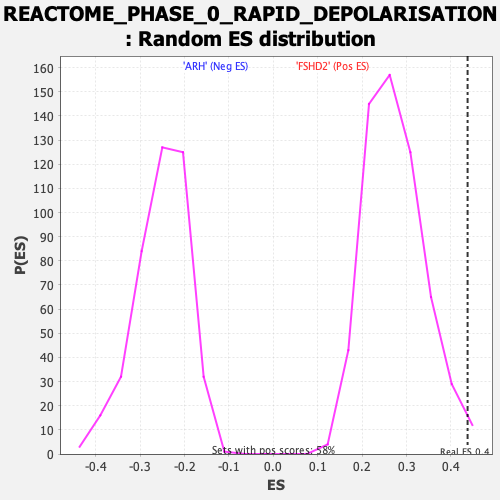
| GeneSet | REACTOME_PHASE_0_RAPID_DEPOLARISATION |
| Enrichment Score (ES) | 0.43751985 |
| Normalized Enrichment Score (NES) | 1.5999498 |
| Nominal p-value | 0.010344828 |
| FDR q-value | 0.18852091 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA2D2 | ENSG00000007402.11 | 156 | 1.309 | 0.0661 | Yes | ||
| 2 | SCN4B | ENSG00000177098.8 | 290 | 1.139 | 0.1236 | Yes | ||
| 3 | CAMK2B | ENSG00000058404.20 | 471 | 1.027 | 0.1741 | Yes | ||
| 4 | SCN7A | ENSG00000136546.16 | 533 | 1.002 | 0.2261 | Yes | ||
| 5 | CACNA2D3 | ENSG00000157445.15 | 791 | 0.915 | 0.2687 | Yes | ||
| 6 | SCN1A | ENSG00000144285.21 | 1332 | 0.796 | 0.2981 | Yes | ||
| 7 | FGF12 | ENSG00000114279.13 | 1848 | 0.725 | 0.3242 | Yes | ||
| 8 | SCN2B | ENSG00000149575.6 | 2157 | 0.688 | 0.3534 | Yes | ||
| 9 | CACNG7 | ENSG00000105605.7 | 2325 | 0.669 | 0.3851 | Yes | ||
| 10 | CACNB2 | ENSG00000165995.20 | 2587 | 0.645 | 0.4132 | Yes | ||
| 11 | CACNG2 | ENSG00000166862.6 | 3228 | 0.589 | 0.4291 | Yes | ||
| 12 | CACNA1F | ENSG00000102001.12 | 4309 | 0.523 | 0.4307 | Yes | ||
| 13 | SCN9A | ENSG00000169432.16 | 5398 | 0.465 | 0.4291 | Yes | ||
| 14 | SCN11A | ENSG00000168356.12 | 6003 | 0.433 | 0.4375 | Yes | ||
| 15 | SCN3B | ENSG00000166257.9 | 10636 | 0.305 | 0.3412 | No | ||
| 16 | SCN5A | ENSG00000183873.15 | 11016 | 0.290 | 0.3474 | No | ||
| 17 | SCN2A | ENSG00000136531.16 | 12159 | 0.265 | 0.3338 | No | ||
| 18 | CACNA2D4 | ENSG00000151062.15 | 13173 | 0.233 | 0.3216 | No | ||
| 19 | CACNB3 | ENSG00000167535.8 | 13461 | 0.223 | 0.3265 | No | ||
| 20 | CACNB1 | ENSG00000067191.16 | 17750 | 0.098 | 0.2275 | No | ||
| 21 | CACNA2D1 | ENSG00000153956.16 | 18667 | 0.074 | 0.2092 | No | ||
| 22 | FGF11 | ENSG00000161958.11 | 19678 | 0.047 | 0.1871 | No | ||
| 23 | CACNG6 | ENSG00000130433.7 | 21278 | 0.004 | 0.1485 | No | ||
| 24 | SCN1B | ENSG00000105711.12 | 22218 | -0.018 | 0.1266 | No | ||
| 25 | SCN3A | ENSG00000153253.18 | 22874 | -0.035 | 0.1126 | No | ||
| 26 | SCN4A | ENSG00000007314.12 | 24646 | -0.081 | 0.0738 | No | ||
| 27 | RANGRF | ENSG00000108961.14 | 26258 | -0.128 | 0.0415 | No | ||
| 28 | CAMK2D | ENSG00000145349.17 | 27641 | -0.169 | 0.0169 | No | ||
| 29 | CACNG1 | ENSG00000108878.5 | 27982 | -0.180 | 0.0182 | No | ||
| 30 | FGF13 | ENSG00000129682.16 | 29879 | -0.241 | -0.0151 | No | ||
| 31 | CACNB4 | ENSG00000182389.19 | 30152 | -0.249 | -0.0084 | No | ||
| 32 | CACNA1S | ENSG00000081248.11 | 30428 | -0.258 | -0.0013 | No | ||
| 33 | CACNA1C | ENSG00000151067.22 | 32998 | -0.336 | -0.0459 | No | ||
| 34 | SCN10A | ENSG00000185313.8 | 33022 | -0.336 | -0.0285 | No | ||
| 35 | CAMK2A | ENSG00000070808.16 | 34695 | -0.356 | -0.0501 | No | ||
| 36 | CAMK2G | ENSG00000148660.20 | 36889 | -0.445 | -0.0797 | No | ||
| 37 | SCN8A | ENSG00000196876.17 | 37337 | -0.474 | -0.0653 | No | ||
| 38 | CACNA1D | ENSG00000157388.19 | 37775 | -0.510 | -0.0487 | No | ||
| 39 | CALM1 | ENSG00000198668.12 | 38464 | -0.558 | -0.0356 | No | ||
| 40 | FGF14 | ENSG00000102466.15 | 38522 | -0.559 | -0.0071 | No | ||
| 41 | CACNG4 | ENSG00000075461.6 | 39529 | -0.640 | 0.0026 | No | ||
| 42 | CACNG8 | ENSG00000142408.4 | 39898 | -0.695 | 0.0307 | No |