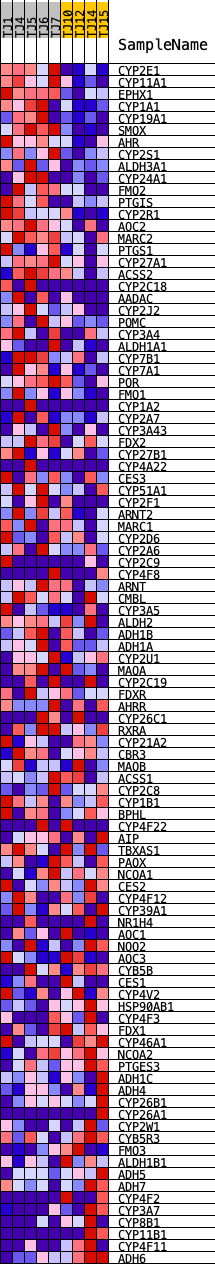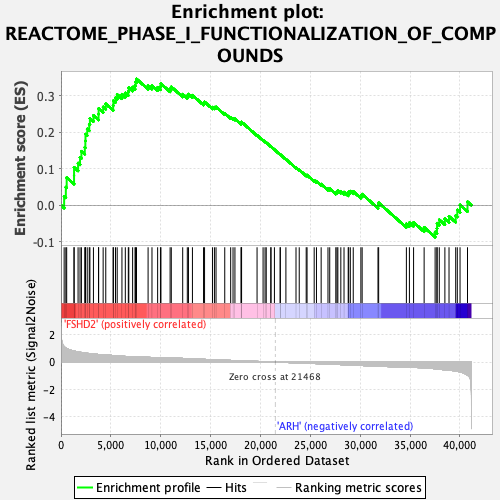
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
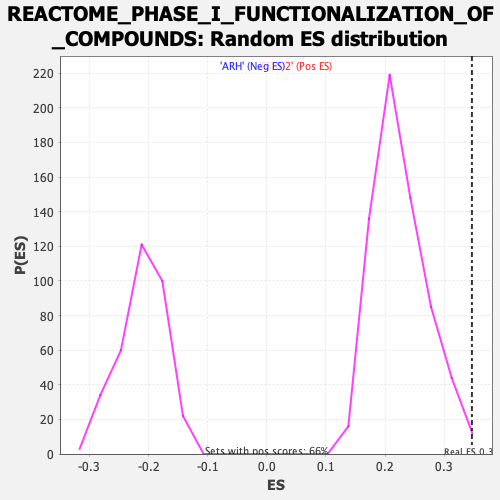
| GeneSet | REACTOME_PHASE_I_FUNCTIONALIZATION_OF_COMPOUNDS |
| Enrichment Score (ES) | 0.346659 |
| Normalized Enrichment Score (NES) | 1.53726 |
| Nominal p-value | 0.0045454544 |
| FDR q-value | 0.22412154 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CYP2E1 | ENSG00000130649.10 | 315 | 1.122 | 0.0245 | Yes | ||
| 2 | CYP11A1 | ENSG00000140459.18 | 485 | 1.022 | 0.0497 | Yes | ||
| 3 | EPHX1 | ENSG00000143819.12 | 570 | 0.990 | 0.0761 | Yes | ||
| 4 | CYP1A1 | ENSG00000140465.14 | 1309 | 0.801 | 0.0811 | Yes | ||
| 5 | CYP19A1 | ENSG00000137869.15 | 1311 | 0.800 | 0.1040 | Yes | ||
| 6 | SMOX | ENSG00000088826.18 | 1712 | 0.745 | 0.1157 | Yes | ||
| 7 | AHR | ENSG00000106546.14 | 1908 | 0.715 | 0.1315 | Yes | ||
| 8 | CYP2S1 | ENSG00000167600.14 | 2047 | 0.699 | 0.1482 | Yes | ||
| 9 | ALDH3A1 | ENSG00000108602.18 | 2386 | 0.664 | 0.1590 | Yes | ||
| 10 | CYP24A1 | ENSG00000019186.10 | 2442 | 0.659 | 0.1766 | Yes | ||
| 11 | FMO2 | ENSG00000094963.14 | 2456 | 0.658 | 0.1951 | Yes | ||
| 12 | PTGIS | ENSG00000124212.6 | 2629 | 0.640 | 0.2093 | Yes | ||
| 13 | CYP2R1 | ENSG00000186104.10 | 2833 | 0.620 | 0.2222 | Yes | ||
| 14 | AOC2 | ENSG00000131480.9 | 2898 | 0.614 | 0.2382 | Yes | ||
| 15 | MARC2 | ENSG00000117791.16 | 3252 | 0.587 | 0.2465 | Yes | ||
| 16 | PTGS1 | ENSG00000095303.17 | 3763 | 0.553 | 0.2499 | Yes | ||
| 17 | CYP27A1 | ENSG00000135929.9 | 3774 | 0.552 | 0.2656 | Yes | ||
| 18 | ACSS2 | ENSG00000131069.20 | 4226 | 0.529 | 0.2697 | Yes | ||
| 19 | CYP2C18 | ENSG00000108242.13 | 4489 | 0.518 | 0.2782 | Yes | ||
| 20 | AADAC | ENSG00000114771.14 | 5228 | 0.475 | 0.2739 | Yes | ||
| 21 | CYP2J2 | ENSG00000134716.11 | 5250 | 0.474 | 0.2870 | Yes | ||
| 22 | POMC | ENSG00000115138.11 | 5463 | 0.461 | 0.2951 | Yes | ||
| 23 | CYP3A4 | ENSG00000160868.15 | 5635 | 0.451 | 0.3039 | Yes | ||
| 24 | ALDH1A1 | ENSG00000165092.13 | 6111 | 0.428 | 0.3046 | Yes | ||
| 25 | CYP7B1 | ENSG00000172817.4 | 6449 | 0.410 | 0.3081 | Yes | ||
| 26 | CYP7A1 | ENSG00000167910.4 | 6750 | 0.398 | 0.3123 | Yes | ||
| 27 | POR | ENSG00000127948.16 | 6799 | 0.396 | 0.3225 | Yes | ||
| 28 | FMO1 | ENSG00000010932.17 | 7182 | 0.381 | 0.3241 | Yes | ||
| 29 | CYP1A2 | ENSG00000140505.7 | 7424 | 0.372 | 0.3289 | Yes | ||
| 30 | CYP2A7 | ENSG00000198077.10 | 7480 | 0.372 | 0.3383 | Yes | ||
| 31 | CYP3A43 | ENSG00000021461.16 | 7570 | 0.368 | 0.3467 | Yes | ||
| 32 | FDX2 | ENSG00000267673.6 | 8733 | 0.346 | 0.3283 | No | ||
| 33 | CYP27B1 | ENSG00000111012.10 | 9123 | 0.330 | 0.3283 | No | ||
| 34 | CYP4A22 | ENSG00000162365.12 | 9690 | 0.319 | 0.3237 | No | ||
| 35 | CES3 | ENSG00000172828.13 | 9973 | 0.313 | 0.3258 | No | ||
| 36 | CYP51A1 | ENSG00000001630.17 | 10022 | 0.310 | 0.3335 | No | ||
| 37 | CYP2F1 | ENSG00000197446.9 | 10941 | 0.293 | 0.3196 | No | ||
| 38 | ARNT2 | ENSG00000172379.21 | 11059 | 0.289 | 0.3250 | No | ||
| 39 | MARC1 | ENSG00000186205.13 | 12203 | 0.264 | 0.3048 | No | ||
| 40 | CYP2D6 | ENSG00000100197.22 | 12665 | 0.249 | 0.3007 | No | ||
| 41 | CYP2A6 | ENSG00000255974.8 | 12781 | 0.245 | 0.3049 | No | ||
| 42 | CYP2C9 | ENSG00000138109.11 | 13179 | 0.233 | 0.3019 | No | ||
| 43 | CYP4F8 | ENSG00000186526.13 | 14293 | 0.197 | 0.2805 | No | ||
| 44 | ARNT | ENSG00000143437.21 | 14395 | 0.194 | 0.2836 | No | ||
| 45 | CMBL | ENSG00000164237.9 | 15188 | 0.168 | 0.2692 | No | ||
| 46 | CYP3A5 | ENSG00000106258.15 | 15384 | 0.162 | 0.2691 | No | ||
| 47 | ALDH2 | ENSG00000111275.13 | 15545 | 0.158 | 0.2697 | No | ||
| 48 | ADH1B | ENSG00000196616.14 | 16420 | 0.133 | 0.2522 | No | ||
| 49 | ADH1A | ENSG00000187758.8 | 17014 | 0.117 | 0.2412 | No | ||
| 50 | CYP2U1 | ENSG00000155016.18 | 17251 | 0.111 | 0.2386 | No | ||
| 51 | MAOA | ENSG00000189221.9 | 17441 | 0.106 | 0.2370 | No | ||
| 52 | CYP2C19 | ENSG00000165841.11 | 18054 | 0.089 | 0.2247 | No | ||
| 53 | FDXR | ENSG00000161513.12 | 18088 | 0.088 | 0.2264 | No | ||
| 54 | AHRR | ENSG00000063438.17 | 18108 | 0.088 | 0.2284 | No | ||
| 55 | CYP26C1 | ENSG00000187553.10 | 19660 | 0.047 | 0.1920 | No | ||
| 56 | RXRA | ENSG00000186350.12 | 20264 | 0.032 | 0.1783 | No | ||
| 57 | CYP21A2 | ENSG00000231852.9 | 20461 | 0.026 | 0.1742 | No | ||
| 58 | CBR3 | ENSG00000159231.6 | 20580 | 0.023 | 0.1720 | No | ||
| 59 | MAOB | ENSG00000069535.14 | 21045 | 0.010 | 0.1610 | No | ||
| 60 | ACSS1 | ENSG00000154930.15 | 21052 | 0.010 | 0.1612 | No | ||
| 61 | CYP2C8 | ENSG00000138115.14 | 21401 | 0.001 | 0.1527 | No | ||
| 62 | CYP1B1 | ENSG00000138061.12 | 21975 | -0.014 | 0.1392 | No | ||
| 63 | BPHL | ENSG00000137274.13 | 21993 | -0.014 | 0.1392 | No | ||
| 64 | CYP4F22 | ENSG00000171954.13 | 22552 | -0.027 | 0.1264 | No | ||
| 65 | AIP | ENSG00000110711.10 | 23565 | -0.054 | 0.1033 | No | ||
| 66 | TBXAS1 | ENSG00000059377.17 | 23889 | -0.063 | 0.0972 | No | ||
| 67 | PAOX | ENSG00000148832.16 | 24587 | -0.080 | 0.0825 | No | ||
| 68 | NCOA1 | ENSG00000084676.15 | 24673 | -0.082 | 0.0828 | No | ||
| 69 | CES2 | ENSG00000172831.13 | 25388 | -0.102 | 0.0684 | No | ||
| 70 | CYP4F12 | ENSG00000186204.15 | 25617 | -0.109 | 0.0659 | No | ||
| 71 | CYP39A1 | ENSG00000146233.8 | 26087 | -0.123 | 0.0580 | No | ||
| 72 | NR1H4 | ENSG00000012504.15 | 26772 | -0.143 | 0.0455 | No | ||
| 73 | AOC1 | ENSG00000002726.20 | 26947 | -0.148 | 0.0455 | No | ||
| 74 | NQO2 | ENSG00000124588.20 | 27556 | -0.166 | 0.0355 | No | ||
| 75 | AOC3 | ENSG00000131471.7 | 27672 | -0.170 | 0.0375 | No | ||
| 76 | CYB5B | ENSG00000103018.16 | 27762 | -0.172 | 0.0403 | No | ||
| 77 | CES1 | ENSG00000198848.12 | 28060 | -0.183 | 0.0383 | No | ||
| 78 | CYP4V2 | ENSG00000145476.16 | 28380 | -0.192 | 0.0361 | No | ||
| 79 | HSP90AB1 | ENSG00000096384.20 | 28800 | -0.205 | 0.0317 | No | ||
| 80 | CYP4F3 | ENSG00000186529.16 | 28849 | -0.207 | 0.0365 | No | ||
| 81 | FDX1 | ENSG00000137714.3 | 29012 | -0.212 | 0.0387 | No | ||
| 82 | CYP46A1 | ENSG00000036530.9 | 29305 | -0.221 | 0.0379 | No | ||
| 83 | NCOA2 | ENSG00000140396.13 | 30070 | -0.246 | 0.0264 | No | ||
| 84 | PTGES3 | ENSG00000110958.16 | 30197 | -0.250 | 0.0305 | No | ||
| 85 | ADH1C | ENSG00000248144.6 | 31787 | -0.289 | 0.0001 | No | ||
| 86 | ADH4 | ENSG00000198099.8 | 31853 | -0.292 | 0.0069 | No | ||
| 87 | CYP26B1 | ENSG00000003137.8 | 34633 | -0.353 | -0.0507 | No | ||
| 88 | CYP26A1 | ENSG00000095596.12 | 34937 | -0.363 | -0.0476 | No | ||
| 89 | CYP2W1 | ENSG00000073067.14 | 35355 | -0.381 | -0.0468 | No | ||
| 90 | CYB5R3 | ENSG00000100243.21 | 36413 | -0.421 | -0.0605 | No | ||
| 91 | FMO3 | ENSG00000007933.13 | 37515 | -0.486 | -0.0734 | No | ||
| 92 | ALDH1B1 | ENSG00000137124.8 | 37698 | -0.502 | -0.0634 | No | ||
| 93 | ADH5 | ENSG00000197894.11 | 37712 | -0.503 | -0.0493 | No | ||
| 94 | ADH7 | ENSG00000196344.11 | 37933 | -0.523 | -0.0396 | No | ||
| 95 | CYP4F2 | ENSG00000186115.13 | 38494 | -0.558 | -0.0372 | No | ||
| 96 | CYP3A7 | ENSG00000160870.14 | 38909 | -0.591 | -0.0303 | No | ||
| 97 | CYP8B1 | ENSG00000180432.6 | 39579 | -0.645 | -0.0281 | No | ||
| 98 | CYP11B1 | ENSG00000160882.11 | 39739 | -0.666 | -0.0129 | No | ||
| 99 | CYP4F11 | ENSG00000171903.16 | 40014 | -0.715 | 0.0010 | No | ||
| 100 | ADH6 | ENSG00000172955.17 | 40762 | -0.939 | 0.0097 | No |