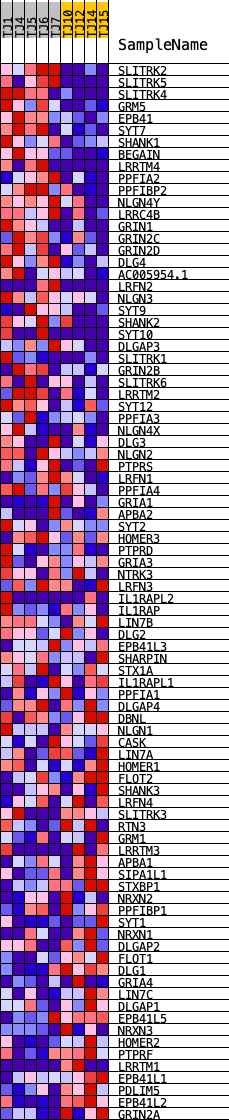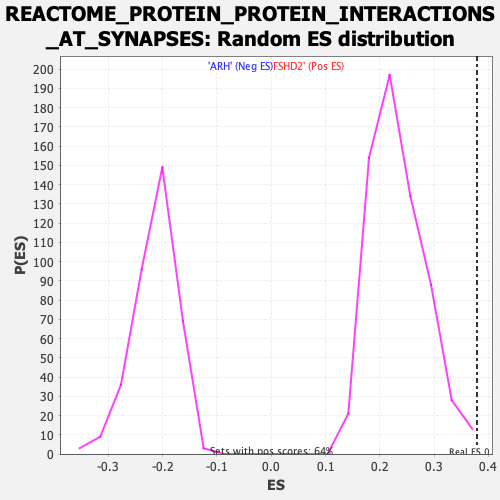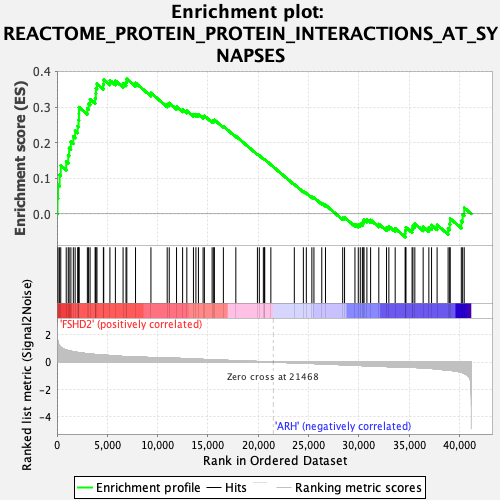
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_PROTEIN_PROTEIN_INTERACTIONS_AT_SYNAPSES |
| Enrichment Score (ES) | 0.37891006 |
| Normalized Enrichment Score (NES) | 1.6264288 |
| Nominal p-value | 0.0031496063 |
| FDR q-value | 0.16778718 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLITRK2 | ENSG00000185985.10 | 52 | 1.628 | 0.0434 | Yes | ||
| 2 | SLITRK5 | ENSG00000165300.7 | 105 | 1.414 | 0.0809 | Yes | ||
| 3 | SLITRK4 | ENSG00000179542.16 | 259 | 1.164 | 0.1091 | Yes | ||
| 4 | GRM5 | ENSG00000168959.14 | 374 | 1.077 | 0.1358 | Yes | ||
| 5 | EPB41 | ENSG00000159023.21 | 916 | 0.879 | 0.1467 | Yes | ||
| 6 | SYT7 | ENSG00000011347.9 | 1114 | 0.834 | 0.1648 | Yes | ||
| 7 | SHANK1 | ENSG00000161681.15 | 1204 | 0.817 | 0.1851 | Yes | ||
| 8 | BEGAIN | ENSG00000183092.16 | 1379 | 0.791 | 0.2025 | Yes | ||
| 9 | LRRTM4 | ENSG00000176204.13 | 1622 | 0.756 | 0.2173 | Yes | ||
| 10 | PPFIA2 | ENSG00000139220.16 | 1802 | 0.733 | 0.2331 | Yes | ||
| 11 | PPFIBP2 | ENSG00000166387.13 | 2053 | 0.698 | 0.2461 | Yes | ||
| 12 | NLGN4Y | ENSG00000165246.14 | 2146 | 0.688 | 0.2628 | Yes | ||
| 13 | LRRC4B | ENSG00000131409.13 | 2177 | 0.686 | 0.2809 | Yes | ||
| 14 | GRIN1 | ENSG00000176884.15 | 2182 | 0.686 | 0.2995 | Yes | ||
| 15 | GRIN2C | ENSG00000161509.14 | 3016 | 0.605 | 0.2958 | Yes | ||
| 16 | GRIN2D | ENSG00000105464.3 | 3138 | 0.596 | 0.3092 | Yes | ||
| 17 | DLG4 | ENSG00000132535.19 | 3287 | 0.584 | 0.3217 | Yes | ||
| 18 | AC005954.1 | ENSG00000267138.1 | 3787 | 0.551 | 0.3246 | Yes | ||
| 19 | LRFN2 | ENSG00000156564.9 | 3853 | 0.547 | 0.3380 | Yes | ||
| 20 | NLGN3 | ENSG00000196338.12 | 3869 | 0.547 | 0.3526 | Yes | ||
| 21 | SYT9 | ENSG00000170743.17 | 3970 | 0.542 | 0.3651 | Yes | ||
| 22 | SHANK2 | ENSG00000162105.19 | 4607 | 0.513 | 0.3636 | Yes | ||
| 23 | SYT10 | ENSG00000110975.8 | 4643 | 0.512 | 0.3768 | Yes | ||
| 24 | DLGAP3 | ENSG00000116544.12 | 5257 | 0.474 | 0.3749 | Yes | ||
| 25 | SLITRK1 | ENSG00000178235.7 | 5801 | 0.443 | 0.3738 | Yes | ||
| 26 | GRIN2B | ENSG00000273079.5 | 6565 | 0.405 | 0.3664 | Yes | ||
| 27 | SLITRK6 | ENSG00000184564.11 | 6857 | 0.394 | 0.3701 | Yes | ||
| 28 | LRRTM2 | ENSG00000146006.8 | 6937 | 0.392 | 0.3789 | Yes | ||
| 29 | SYT12 | ENSG00000173227.14 | 7802 | 0.362 | 0.3678 | No | ||
| 30 | PPFIA3 | ENSG00000177380.14 | 9332 | 0.322 | 0.3394 | No | ||
| 31 | NLGN4X | ENSG00000146938.16 | 10948 | 0.293 | 0.3081 | No | ||
| 32 | DLG3 | ENSG00000082458.12 | 11151 | 0.285 | 0.3110 | No | ||
| 33 | NLGN2 | ENSG00000169992.10 | 11877 | 0.275 | 0.3009 | No | ||
| 34 | PTPRS | ENSG00000105426.16 | 12480 | 0.255 | 0.2933 | No | ||
| 35 | LRFN1 | ENSG00000128011.4 | 12899 | 0.241 | 0.2897 | No | ||
| 36 | PPFIA4 | ENSG00000143847.15 | 13557 | 0.220 | 0.2797 | No | ||
| 37 | GRIA1 | ENSG00000155511.18 | 13800 | 0.213 | 0.2797 | No | ||
| 38 | APBA2 | ENSG00000034053.14 | 14052 | 0.204 | 0.2792 | No | ||
| 39 | SYT2 | ENSG00000143858.12 | 14514 | 0.190 | 0.2732 | No | ||
| 40 | HOMER3 | ENSG00000051128.19 | 14643 | 0.186 | 0.2751 | No | ||
| 41 | PTPRD | ENSG00000153707.17 | 15413 | 0.161 | 0.2608 | No | ||
| 42 | GRIA3 | ENSG00000125675.18 | 15568 | 0.157 | 0.2614 | No | ||
| 43 | NTRK3 | ENSG00000140538.16 | 15638 | 0.155 | 0.2640 | No | ||
| 44 | LRFN3 | ENSG00000126243.8 | 16530 | 0.130 | 0.2459 | No | ||
| 45 | IL1RAPL2 | ENSG00000189108.13 | 17767 | 0.097 | 0.2184 | No | ||
| 46 | IL1RAP | ENSG00000196083.10 | 19912 | 0.041 | 0.1674 | No | ||
| 47 | LIN7B | ENSG00000104863.12 | 20102 | 0.037 | 0.1638 | No | ||
| 48 | DLG2 | ENSG00000150672.17 | 20501 | 0.024 | 0.1547 | No | ||
| 49 | EPB41L3 | ENSG00000082397.17 | 20603 | 0.022 | 0.1529 | No | ||
| 50 | SHARPIN | ENSG00000179526.17 | 20639 | 0.021 | 0.1526 | No | ||
| 51 | STX1A | ENSG00000106089.12 | 21247 | 0.005 | 0.1380 | No | ||
| 52 | IL1RAPL1 | ENSG00000169306.10 | 23578 | -0.055 | 0.0828 | No | ||
| 53 | PPFIA1 | ENSG00000131626.18 | 24477 | -0.077 | 0.0630 | No | ||
| 54 | DLGAP4 | ENSG00000080845.17 | 24771 | -0.085 | 0.0582 | No | ||
| 55 | DBNL | ENSG00000136279.20 | 25322 | -0.100 | 0.0475 | No | ||
| 56 | NLGN1 | ENSG00000169760.17 | 25527 | -0.106 | 0.0455 | No | ||
| 57 | CASK | ENSG00000147044.21 | 26307 | -0.129 | 0.0300 | No | ||
| 58 | LIN7A | ENSG00000111052.7 | 26675 | -0.140 | 0.0249 | No | ||
| 59 | HOMER1 | ENSG00000152413.15 | 28384 | -0.192 | -0.0114 | No | ||
| 60 | FLOT2 | ENSG00000132589.16 | 28570 | -0.198 | -0.0105 | No | ||
| 61 | SHANK3 | ENSG00000251322.8 | 29602 | -0.230 | -0.0292 | No | ||
| 62 | LRFN4 | ENSG00000173621.9 | 29933 | -0.242 | -0.0306 | No | ||
| 63 | SLITRK3 | ENSG00000121871.4 | 30133 | -0.248 | -0.0287 | No | ||
| 64 | RTN3 | ENSG00000133318.14 | 30339 | -0.255 | -0.0267 | No | ||
| 65 | GRM1 | ENSG00000152822.14 | 30417 | -0.257 | -0.0215 | No | ||
| 66 | LRRTM3 | ENSG00000198739.11 | 30506 | -0.260 | -0.0165 | No | ||
| 67 | APBA1 | ENSG00000107282.8 | 30789 | -0.269 | -0.0160 | No | ||
| 68 | SIPA1L1 | ENSG00000197555.9 | 31158 | -0.283 | -0.0172 | No | ||
| 69 | STXBP1 | ENSG00000136854.21 | 31972 | -0.295 | -0.0289 | No | ||
| 70 | NRXN2 | ENSG00000110076.18 | 32739 | -0.325 | -0.0386 | No | ||
| 71 | PPFIBP1 | ENSG00000110841.14 | 32972 | -0.335 | -0.0351 | No | ||
| 72 | SYT1 | ENSG00000067715.14 | 33600 | -0.340 | -0.0410 | No | ||
| 73 | NRXN1 | ENSG00000179915.23 | 34602 | -0.352 | -0.0558 | No | ||
| 74 | DLGAP2 | ENSG00000198010.12 | 34606 | -0.352 | -0.0462 | No | ||
| 75 | FLOT1 | ENSG00000137312.15 | 34673 | -0.355 | -0.0381 | No | ||
| 76 | DLG1 | ENSG00000075711.21 | 35280 | -0.377 | -0.0425 | No | ||
| 77 | GRIA4 | ENSG00000152578.13 | 35362 | -0.382 | -0.0340 | No | ||
| 78 | LIN7C | ENSG00000148943.12 | 35551 | -0.391 | -0.0278 | No | ||
| 79 | DLGAP1 | ENSG00000170579.17 | 36379 | -0.420 | -0.0365 | No | ||
| 80 | EPB41L5 | ENSG00000115109.14 | 36945 | -0.449 | -0.0379 | No | ||
| 81 | NRXN3 | ENSG00000021645.18 | 37213 | -0.464 | -0.0317 | No | ||
| 82 | HOMER2 | ENSG00000103942.13 | 37764 | -0.508 | -0.0311 | No | ||
| 83 | PTPRF | ENSG00000142949.17 | 38859 | -0.586 | -0.0417 | No | ||
| 84 | LRRTM1 | ENSG00000162951.11 | 39016 | -0.602 | -0.0290 | No | ||
| 85 | EPB41L1 | ENSG00000088367.23 | 39064 | -0.608 | -0.0135 | No | ||
| 86 | PDLIM5 | ENSG00000163110.15 | 40169 | -0.746 | -0.0199 | No | ||
| 87 | EPB41L2 | ENSG00000079819.19 | 40303 | -0.778 | -0.0018 | No | ||
| 88 | GRIN2A | ENSG00000183454.17 | 40459 | -0.827 | 0.0171 | No |