Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_REGULATION_OF_PLK1_ACTIVITY_AT_G2_M_TRANSITION |
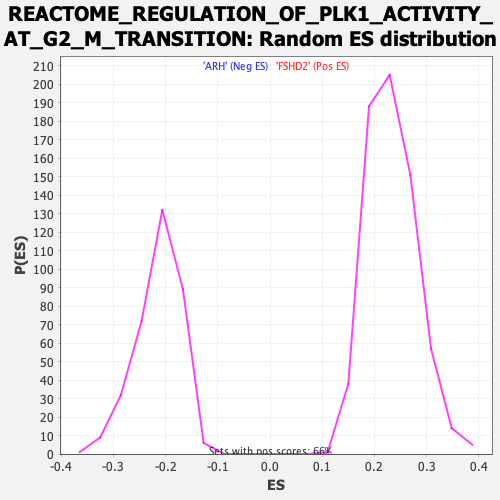
| Enrichment Score (ES) | -0.492044 |
| Normalized Enrichment Score (NES) | -2.3001006 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.397385E-4 |
| FWER p-Value | 0.006 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HAUS7 | ENSG00000213397.10 | 1436 | 0.783 | -0.0055 | No | ||
| 2 | CDK5RAP2 | ENSG00000136861.18 | 3658 | 0.560 | -0.0384 | No | ||
| 3 | NINL | ENSG00000101004.15 | 3818 | 0.548 | -0.0217 | No | ||
| 4 | HAUS5 | ENSG00000249115.9 | 5985 | 0.434 | -0.0581 | No | ||
| 5 | CEP131 | ENSG00000141577.14 | 6755 | 0.398 | -0.0618 | No | ||
| 6 | PRKAR2B | ENSG00000005249.13 | 12170 | 0.265 | -0.1836 | No | ||
| 7 | CSNK1E | ENSG00000213923.12 | 12504 | 0.254 | -0.1821 | No | ||
| 8 | SFI1 | ENSG00000198089.16 | 13183 | 0.233 | -0.1899 | No | ||
| 9 | CNTRL | ENSG00000119397.16 | 13560 | 0.220 | -0.1907 | No | ||
| 10 | ODF2 | ENSG00000136811.16 | 15981 | 0.146 | -0.2441 | No | ||
| 11 | CEP135 | ENSG00000174799.11 | 16064 | 0.144 | -0.2407 | No | ||
| 12 | CEP192 | ENSG00000101639.18 | 17033 | 0.117 | -0.2599 | No | ||
| 13 | CEP57 | ENSG00000166037.11 | 17042 | 0.117 | -0.2557 | No | ||
| 14 | CEP164 | ENSG00000110274.16 | 17447 | 0.105 | -0.2616 | No | ||
| 15 | CSNK1D | ENSG00000141551.14 | 18935 | 0.067 | -0.2952 | No | ||
| 16 | HAUS1 | ENSG00000152240.13 | 19046 | 0.064 | -0.2955 | No | ||
| 17 | AKAP9 | ENSG00000127914.16 | 19199 | 0.060 | -0.2970 | No | ||
| 18 | HAUS3 | ENSG00000214367.8 | 19258 | 0.058 | -0.2962 | No | ||
| 19 | UBC | ENSG00000150991.15 | 20948 | 0.013 | -0.3368 | No | ||
| 20 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | -0.3448 | No | ||
| 21 | FGFR1OP | ENSG00000213066.13 | 21442 | 0.000 | -0.3486 | No | ||
| 22 | UBB | ENSG00000170315.14 | 22145 | -0.016 | -0.3651 | No | ||
| 23 | OFD1 | ENSG00000046651.15 | 22172 | -0.017 | -0.3651 | No | ||
| 24 | UBA52 | ENSG00000221983.8 | 22256 | -0.020 | -0.3664 | No | ||
| 25 | DCTN3 | ENSG00000137100.16 | 22796 | -0.033 | -0.3782 | No | ||
| 26 | CEP70 | ENSG00000114107.9 | 23190 | -0.045 | -0.3861 | No | ||
| 27 | CEP76 | ENSG00000101624.11 | 23293 | -0.047 | -0.3868 | No | ||
| 28 | PCNT | ENSG00000160299.17 | 23855 | -0.062 | -0.3982 | No | ||
| 29 | NDE1 | ENSG00000072864.15 | 23985 | -0.064 | -0.3989 | No | ||
| 30 | CEP41 | ENSG00000106477.19 | 24138 | -0.068 | -0.4000 | No | ||
| 31 | ALMS1 | ENSG00000116127.18 | 24464 | -0.076 | -0.4051 | No | ||
| 32 | PRKACA | ENSG00000072062.14 | 24521 | -0.077 | -0.4035 | No | ||
| 33 | CUL1 | ENSG00000055130.17 | 24550 | -0.079 | -0.4013 | No | ||
| 34 | SSNA1 | ENSG00000176101.12 | 24700 | -0.083 | -0.4018 | No | ||
| 35 | RPS27A | ENSG00000143947.13 | 24801 | -0.086 | -0.4010 | No | ||
| 36 | BTRC | ENSG00000166167.18 | 25965 | -0.119 | -0.4248 | No | ||
| 37 | CENPJ | ENSG00000151849.15 | 26281 | -0.128 | -0.4277 | No | ||
| 38 | HAUS8 | ENSG00000131351.15 | 26530 | -0.136 | -0.4286 | No | ||
| 39 | HAUS4 | ENSG00000092036.18 | 26579 | -0.137 | -0.4246 | No | ||
| 40 | PPP1R12B | ENSG00000077157.22 | 27498 | -0.164 | -0.4407 | No | ||
| 41 | PCM1 | ENSG00000078674.17 | 27675 | -0.170 | -0.4386 | No | ||
| 42 | CLASP1 | ENSG00000074054.18 | 28043 | -0.182 | -0.4407 | No | ||
| 43 | SKP1 | ENSG00000113558.18 | 28827 | -0.206 | -0.4520 | No | ||
| 44 | RAB8A | ENSG00000167461.12 | 29225 | -0.218 | -0.4535 | No | ||
| 45 | AJUBA | ENSG00000129474.16 | 29753 | -0.236 | -0.4574 | No | ||
| 46 | CEP72 | ENSG00000112877.8 | 29993 | -0.243 | -0.4540 | No | ||
| 47 | SDCCAG8 | ENSG00000054282.16 | 30251 | -0.252 | -0.4508 | No | ||
| 48 | CEP63 | ENSG00000182923.17 | 30525 | -0.260 | -0.4477 | No | ||
| 49 | TUBB4A | ENSG00000104833.12 | 31106 | -0.281 | -0.4512 | No | ||
| 50 | DYNC1H1 | ENSG00000197102.12 | 31822 | -0.290 | -0.4577 | No | ||
| 51 | YWHAG | ENSG00000170027.7 | 32308 | -0.310 | -0.4578 | No | ||
| 52 | CEP78 | ENSG00000148019.13 | 32557 | -0.319 | -0.4518 | No | ||
| 53 | DCTN2 | ENSG00000175203.15 | 32923 | -0.333 | -0.4482 | No | ||
| 54 | PPP2R1A | ENSG00000105568.18 | 34725 | -0.357 | -0.4786 | Yes | ||
| 55 | CEP250 | ENSG00000126001.16 | 35181 | -0.373 | -0.4756 | Yes | ||
| 56 | FBXW11 | ENSG00000072803.17 | 35526 | -0.390 | -0.4693 | Yes | ||
| 57 | CCP110 | ENSG00000103540.16 | 35632 | -0.395 | -0.4570 | Yes | ||
| 58 | DCTN1 | ENSG00000204843.12 | 35697 | -0.397 | -0.4436 | Yes | ||
| 59 | PAFAH1B1 | ENSG00000007168.13 | 36076 | -0.408 | -0.4374 | Yes | ||
| 60 | DYNC1I2 | ENSG00000077380.15 | 36153 | -0.411 | -0.4238 | Yes | ||
| 61 | PPP1CB | ENSG00000213639.10 | 36325 | -0.417 | -0.4122 | Yes | ||
| 62 | YWHAE | ENSG00000108953.17 | 36492 | -0.426 | -0.4002 | Yes | ||
| 63 | CCNB2 | ENSG00000157456.8 | 36613 | -0.432 | -0.3869 | Yes | ||
| 64 | PLK1 | ENSG00000166851.15 | 36707 | -0.436 | -0.3727 | Yes | ||
| 65 | HAUS2 | ENSG00000137814.11 | 36796 | -0.439 | -0.3583 | Yes | ||
| 66 | ACTR1A | ENSG00000138107.13 | 36970 | -0.450 | -0.3456 | Yes | ||
| 67 | MAPRE1 | ENSG00000101367.9 | 37101 | -0.458 | -0.3315 | Yes | ||
| 68 | NEDD1 | ENSG00000139350.11 | 37180 | -0.462 | -0.3160 | Yes | ||
| 69 | CEP152 | ENSG00000103995.14 | 37402 | -0.478 | -0.3034 | Yes | ||
| 70 | BORA | ENSG00000136122.17 | 37603 | -0.494 | -0.2896 | Yes | ||
| 71 | CKAP5 | ENSG00000175216.15 | 37814 | -0.514 | -0.2754 | Yes | ||
| 72 | HSP90AA1 | ENSG00000080824.18 | 37823 | -0.514 | -0.2562 | Yes | ||
| 73 | NEK2 | ENSG00000117650.13 | 37958 | -0.525 | -0.2397 | Yes | ||
| 74 | CETN2 | ENSG00000147400.9 | 38135 | -0.537 | -0.2238 | Yes | ||
| 75 | CCNB1 | ENSG00000134057.15 | 38169 | -0.540 | -0.2043 | Yes | ||
| 76 | HAUS6 | ENSG00000147874.11 | 38210 | -0.543 | -0.1848 | Yes | ||
| 77 | TUBG1 | ENSG00000131462.8 | 38291 | -0.550 | -0.1660 | Yes | ||
| 78 | DYNLL1 | ENSG00000088986.11 | 38317 | -0.554 | -0.1457 | Yes | ||
| 79 | TUBA1A | ENSG00000167552.14 | 38598 | -0.561 | -0.1314 | Yes | ||
| 80 | CEP290 | ENSG00000198707.16 | 38626 | -0.564 | -0.1108 | Yes | ||
| 81 | AURKA | ENSG00000087586.17 | 38638 | -0.565 | -0.0898 | Yes | ||
| 82 | TUBB | ENSG00000196230.13 | 39081 | -0.611 | -0.0776 | Yes | ||
| 83 | PLK4 | ENSG00000142731.11 | 39267 | -0.620 | -0.0587 | Yes | ||
| 84 | CDK1 | ENSG00000170312.16 | 39453 | -0.632 | -0.0394 | Yes | ||
| 85 | OPTN | ENSG00000123240.17 | 39634 | -0.651 | -0.0193 | Yes | ||
| 86 | TUBB4B | ENSG00000188229.6 | 40106 | -0.733 | -0.0032 | Yes | ||
| 87 | PPP1R12A | ENSG00000058272.19 | 40248 | -0.765 | 0.0222 | Yes |