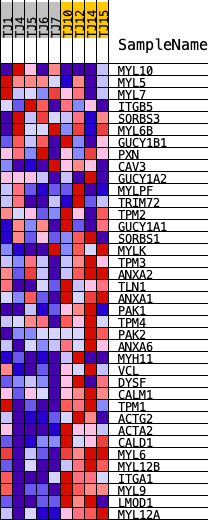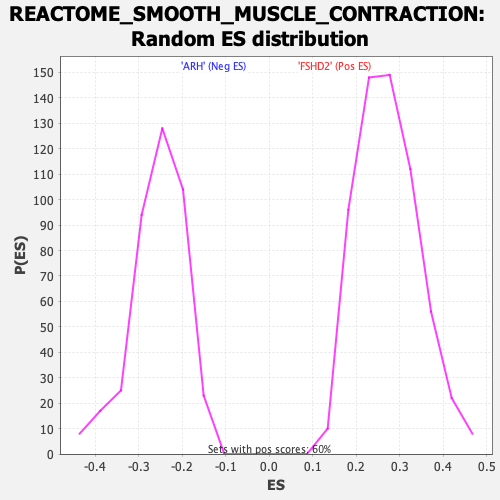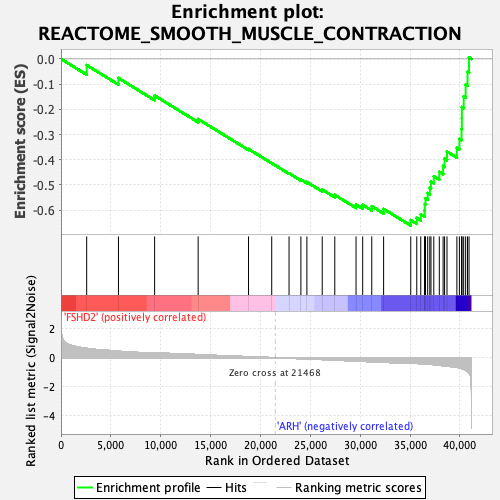
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_SMOOTH_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | -0.66116625 |
| Normalized Enrichment Score (NES) | -2.5992444 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MYL10 | ENSG00000106436.7 | 2575 | 0.646 | -0.0246 | No | ||
| 2 | MYL5 | ENSG00000215375.6 | 5761 | 0.445 | -0.0759 | No | ||
| 3 | MYL7 | ENSG00000106631.8 | 9388 | 0.320 | -0.1452 | No | ||
| 4 | ITGB5 | ENSG00000082781.12 | 13753 | 0.214 | -0.2387 | No | ||
| 5 | SORBS3 | ENSG00000120896.13 | 18804 | 0.070 | -0.3574 | No | ||
| 6 | MYL6B | ENSG00000196465.10 | 21130 | 0.007 | -0.4135 | No | ||
| 7 | GUCY1B1 | ENSG00000061918.13 | 22867 | -0.035 | -0.4536 | No | ||
| 8 | PXN | ENSG00000089159.16 | 24055 | -0.066 | -0.4786 | No | ||
| 9 | CAV3 | ENSG00000182533.6 | 24657 | -0.081 | -0.4884 | No | ||
| 10 | GUCY1A2 | ENSG00000152402.10 | 26195 | -0.126 | -0.5184 | No | ||
| 11 | MYLPF | ENSG00000180209.12 | 27454 | -0.163 | -0.5394 | No | ||
| 12 | TRIM72 | ENSG00000177238.14 | 29585 | -0.230 | -0.5777 | No | ||
| 13 | TPM2 | ENSG00000198467.15 | 30252 | -0.252 | -0.5791 | No | ||
| 14 | GUCY1A1 | ENSG00000164116.16 | 31161 | -0.283 | -0.5845 | No | ||
| 15 | SORBS1 | ENSG00000095637.22 | 32350 | -0.311 | -0.5950 | No | ||
| 16 | MYLK | ENSG00000065534.18 | 35071 | -0.368 | -0.6395 | Yes | ||
| 17 | TPM3 | ENSG00000143549.21 | 35675 | -0.397 | -0.6308 | Yes | ||
| 18 | ANXA2 | ENSG00000182718.16 | 36074 | -0.407 | -0.6165 | Yes | ||
| 19 | TLN1 | ENSG00000137076.21 | 36459 | -0.424 | -0.6009 | Yes | ||
| 20 | ANXA1 | ENSG00000135046.14 | 36480 | -0.425 | -0.5763 | Yes | ||
| 21 | PAK1 | ENSG00000149269.9 | 36549 | -0.430 | -0.5527 | Yes | ||
| 22 | TPM4 | ENSG00000167460.17 | 36771 | -0.438 | -0.5323 | Yes | ||
| 23 | PAK2 | ENSG00000180370.10 | 36983 | -0.451 | -0.5108 | Yes | ||
| 24 | ANXA6 | ENSG00000197043.14 | 37098 | -0.457 | -0.4867 | Yes | ||
| 25 | MYH11 | ENSG00000133392.18 | 37382 | -0.477 | -0.4655 | Yes | ||
| 26 | VCL | ENSG00000035403.17 | 37935 | -0.523 | -0.4481 | Yes | ||
| 27 | DYSF | ENSG00000135636.14 | 38316 | -0.553 | -0.4248 | Yes | ||
| 28 | CALM1 | ENSG00000198668.12 | 38464 | -0.558 | -0.3955 | Yes | ||
| 29 | TPM1 | ENSG00000140416.21 | 38707 | -0.572 | -0.3677 | Yes | ||
| 30 | ACTG2 | ENSG00000163017.14 | 39698 | -0.662 | -0.3528 | Yes | ||
| 31 | ACTA2 | ENSG00000107796.13 | 39950 | -0.704 | -0.3175 | Yes | ||
| 32 | CALD1 | ENSG00000122786.20 | 40163 | -0.745 | -0.2788 | Yes | ||
| 33 | MYL6 | ENSG00000092841.18 | 40195 | -0.751 | -0.2353 | Yes | ||
| 34 | MYL12B | ENSG00000118680.13 | 40197 | -0.752 | -0.1911 | Yes | ||
| 35 | ITGA1 | ENSG00000213949.10 | 40378 | -0.799 | -0.1484 | Yes | ||
| 36 | MYL9 | ENSG00000101335.10 | 40576 | -0.868 | -0.1021 | Yes | ||
| 37 | LMOD1 | ENSG00000163431.13 | 40764 | -0.941 | -0.0513 | Yes | ||
| 38 | MYL12A | ENSG00000101608.12 | 40916 | -1.034 | 0.0060 | Yes |