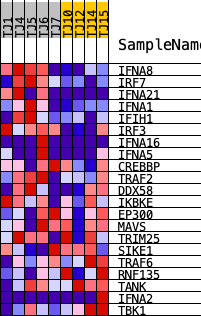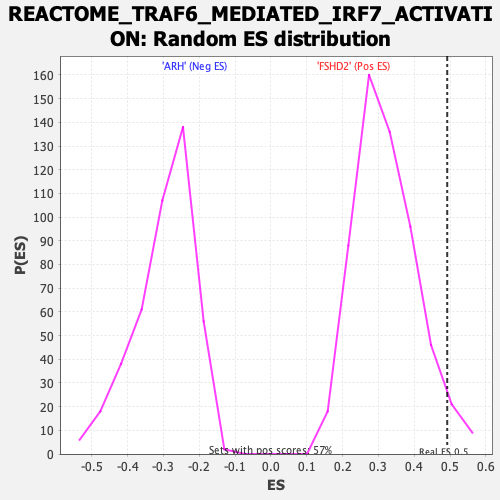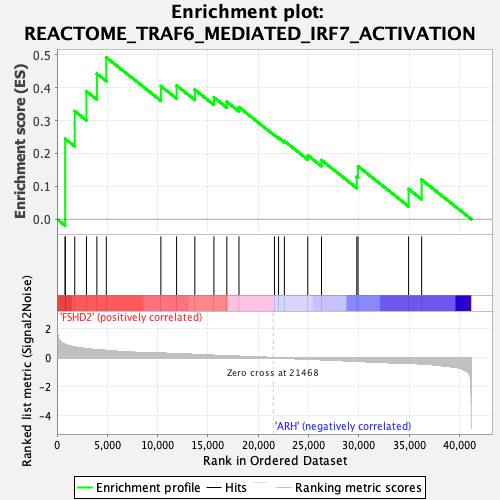
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_TRAF6_MEDIATED_IRF7_ACTIVATION |
| Enrichment Score (ES) | 0.4919849 |
| Normalized Enrichment Score (NES) | 1.5341026 |
| Nominal p-value | 0.040069688 |
| FDR q-value | 0.22432132 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IFNA8 | ENSG00000120242.4 | 811 | 0.910 | 0.1124 | Yes | ||
| 2 | IRF7 | ENSG00000185507.21 | 812 | 0.910 | 0.2445 | Yes | ||
| 3 | IFNA21 | ENSG00000137080.4 | 1763 | 0.738 | 0.3285 | Yes | ||
| 4 | IFNA1 | ENSG00000197919.6 | 2927 | 0.612 | 0.3890 | Yes | ||
| 5 | IFIH1 | ENSG00000115267.8 | 3962 | 0.542 | 0.4426 | Yes | ||
| 6 | IRF3 | ENSG00000126456.16 | 4900 | 0.497 | 0.4920 | Yes | ||
| 7 | IFNA16 | ENSG00000147885.4 | 10318 | 0.305 | 0.4046 | No | ||
| 8 | IFNA5 | ENSG00000147873.5 | 11883 | 0.275 | 0.4066 | No | ||
| 9 | CREBBP | ENSG00000005339.14 | 13694 | 0.216 | 0.3939 | No | ||
| 10 | TRAF2 | ENSG00000127191.18 | 15591 | 0.157 | 0.3706 | No | ||
| 11 | DDX58 | ENSG00000107201.10 | 16873 | 0.121 | 0.3569 | No | ||
| 12 | IKBKE | ENSG00000263528.8 | 18079 | 0.088 | 0.3405 | No | ||
| 13 | EP300 | ENSG00000100393.13 | 21602 | -0.004 | 0.2555 | No | ||
| 14 | MAVS | ENSG00000088888.18 | 22010 | -0.015 | 0.2477 | No | ||
| 15 | TRIM25 | ENSG00000121060.18 | 22590 | -0.028 | 0.2377 | No | ||
| 16 | SIKE1 | ENSG00000052723.11 | 24916 | -0.089 | 0.1941 | No | ||
| 17 | TRAF6 | ENSG00000175104.15 | 26280 | -0.128 | 0.1795 | No | ||
| 18 | RNF135 | ENSG00000181481.14 | 29787 | -0.238 | 0.1288 | No | ||
| 19 | TANK | ENSG00000136560.14 | 29913 | -0.241 | 0.1608 | No | ||
| 20 | IFNA2 | ENSG00000188379.7 | 34933 | -0.363 | 0.0916 | No | ||
| 21 | TBK1 | ENSG00000183735.10 | 36229 | -0.412 | 0.1199 | No |