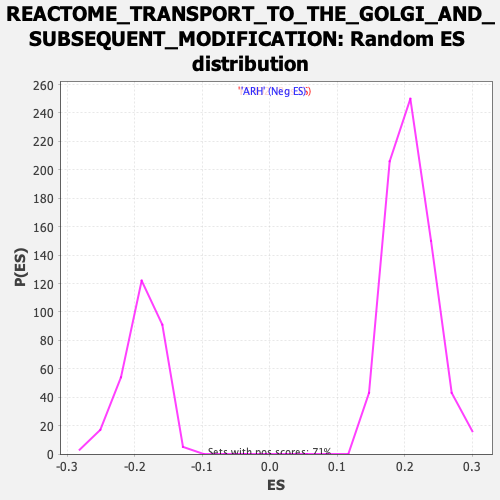
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | REACTOME_TRANSPORT_TO_THE_GOLGI_AND_SUBSEQUENT_MODIFICATION |
| Enrichment Score (ES) | -0.43527246 |
| Normalized Enrichment Score (NES) | -2.296546 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.297462E-4 |
| FWER p-Value | 0.006 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SPTBN2 | ENSG00000173898.13 | 2627 | 0.640 | -0.0508 | No | ||
| 2 | FUCA1 | ENSG00000179163.11 | 4024 | 0.539 | -0.0737 | No | ||
| 3 | LMAN1L | ENSG00000140506.17 | 4099 | 0.538 | -0.0644 | No | ||
| 4 | CHST8 | ENSG00000124302.13 | 5990 | 0.434 | -0.1015 | No | ||
| 5 | DYNC1I1 | ENSG00000158560.14 | 7076 | 0.386 | -0.1200 | No | ||
| 6 | ST6GAL1 | ENSG00000073849.15 | 7610 | 0.366 | -0.1254 | No | ||
| 7 | ARFGAP1 | ENSG00000101199.13 | 7678 | 0.364 | -0.1195 | No | ||
| 8 | TUBB1 | ENSG00000101162.3 | 8127 | 0.356 | -0.1231 | No | ||
| 9 | MGAT4A | ENSG00000071073.13 | 9926 | 0.314 | -0.1605 | No | ||
| 10 | ST8SIA3 | ENSG00000177511.6 | 10620 | 0.305 | -0.1710 | No | ||
| 11 | TUBA3D | ENSG00000075886.11 | 10693 | 0.303 | -0.1665 | No | ||
| 12 | FUT8 | ENSG00000033170.16 | 11841 | 0.277 | -0.1888 | No | ||
| 13 | NAPB | ENSG00000125814.17 | 13487 | 0.222 | -0.2243 | No | ||
| 14 | GRIA1 | ENSG00000155511.18 | 13800 | 0.213 | -0.2275 | No | ||
| 15 | SPTA1 | ENSG00000163554.15 | 14127 | 0.202 | -0.2313 | No | ||
| 16 | MGAT1 | ENSG00000131446.16 | 14591 | 0.187 | -0.2387 | No | ||
| 17 | TMEM115 | ENSG00000126062.4 | 14608 | 0.187 | -0.2353 | No | ||
| 18 | TUBB3 | ENSG00000258947.7 | 15608 | 0.156 | -0.2564 | No | ||
| 19 | CHST10 | ENSG00000115526.11 | 16051 | 0.144 | -0.2642 | No | ||
| 20 | CTSC | ENSG00000109861.16 | 16154 | 0.141 | -0.2638 | No | ||
| 21 | LMAN2 | ENSG00000169223.15 | 16183 | 0.140 | -0.2616 | No | ||
| 22 | MAN1A2 | ENSG00000198162.12 | 16432 | 0.133 | -0.2649 | No | ||
| 23 | COG7 | ENSG00000168434.13 | 16642 | 0.127 | -0.2673 | No | ||
| 24 | STX5 | ENSG00000162236.12 | 16747 | 0.124 | -0.2673 | No | ||
| 25 | COL7A1 | ENSG00000114270.17 | 16820 | 0.122 | -0.2665 | No | ||
| 26 | SEC16A | ENSG00000148396.18 | 17194 | 0.112 | -0.2733 | No | ||
| 27 | TRAPPC2L | ENSG00000167515.10 | 17360 | 0.108 | -0.2751 | No | ||
| 28 | ARFGAP2 | ENSG00000149182.15 | 17482 | 0.104 | -0.2759 | No | ||
| 29 | MAN2A2 | ENSG00000196547.15 | 17736 | 0.098 | -0.2800 | No | ||
| 30 | COG8 | ENSG00000213380.15 | 17846 | 0.095 | -0.2807 | No | ||
| 31 | ANK1 | ENSG00000029534.20 | 17982 | 0.091 | -0.2821 | No | ||
| 32 | TRAPPC9 | ENSG00000167632.16 | 18412 | 0.082 | -0.2909 | No | ||
| 33 | TMED9 | ENSG00000184840.12 | 18711 | 0.073 | -0.2966 | No | ||
| 34 | ST3GAL4 | ENSG00000110080.18 | 18736 | 0.072 | -0.2957 | No | ||
| 35 | AREG | ENSG00000109321.11 | 18751 | 0.072 | -0.2946 | No | ||
| 36 | COG2 | ENSG00000135775.14 | 18893 | 0.068 | -0.2966 | No | ||
| 37 | CSNK1D | ENSG00000141551.14 | 18935 | 0.067 | -0.2962 | No | ||
| 38 | LMAN1 | ENSG00000074695.6 | 19261 | 0.058 | -0.3029 | No | ||
| 39 | TUBA4B | ENSG00000243910.8 | 19289 | 0.057 | -0.3024 | No | ||
| 40 | KDELR1 | ENSG00000105438.9 | 19315 | 0.056 | -0.3019 | No | ||
| 41 | TRAPPC4 | ENSG00000196655.12 | 19317 | 0.056 | -0.3008 | No | ||
| 42 | TRAPPC6A | ENSG00000007255.10 | 19666 | 0.047 | -0.3083 | No | ||
| 43 | CNIH3 | ENSG00000143786.8 | 19901 | 0.041 | -0.3131 | No | ||
| 44 | MIA3 | ENSG00000154305.18 | 20077 | 0.038 | -0.3166 | No | ||
| 45 | BET1L | ENSG00000177951.17 | 20317 | 0.030 | -0.3218 | No | ||
| 46 | STX17 | ENSG00000136874.11 | 20451 | 0.026 | -0.3245 | No | ||
| 47 | GOLGA2 | ENSG00000167110.17 | 20622 | 0.022 | -0.3282 | No | ||
| 48 | TUBA4A | ENSG00000127824.14 | 21284 | 0.004 | -0.3443 | No | ||
| 49 | NAPG | ENSG00000134265.13 | 21412 | 0.001 | -0.3473 | No | ||
| 50 | SPTBN4 | ENSG00000160460.16 | 21556 | -0.003 | -0.3508 | No | ||
| 51 | ARF5 | ENSG00000004059.11 | 21663 | -0.006 | -0.3533 | No | ||
| 52 | CD55 | ENSG00000196352.15 | 21778 | -0.009 | -0.3559 | No | ||
| 53 | MIA2 | ENSG00000150527.17 | 21950 | -0.013 | -0.3598 | No | ||
| 54 | CD59 | ENSG00000085063.16 | 22198 | -0.018 | -0.3654 | No | ||
| 55 | TMED2 | ENSG00000086598.11 | 22331 | -0.022 | -0.3682 | No | ||
| 56 | SEC13 | ENSG00000157020.18 | 22468 | -0.026 | -0.3710 | No | ||
| 57 | ST8SIA6 | ENSG00000148488.17 | 22520 | -0.027 | -0.3717 | No | ||
| 58 | COG4 | ENSG00000103051.19 | 22739 | -0.032 | -0.3763 | No | ||
| 59 | KDELR3 | ENSG00000100196.11 | 22740 | -0.032 | -0.3757 | No | ||
| 60 | GOLGB1 | ENSG00000173230.15 | 22743 | -0.032 | -0.3751 | No | ||
| 61 | DCTN3 | ENSG00000137100.16 | 22796 | -0.033 | -0.3756 | No | ||
| 62 | ARF1 | ENSG00000143761.16 | 23029 | -0.040 | -0.3805 | No | ||
| 63 | B4GALT3 | ENSG00000158850.15 | 23167 | -0.044 | -0.3829 | No | ||
| 64 | MGAT3 | ENSG00000128268.12 | 23231 | -0.045 | -0.3835 | No | ||
| 65 | DYNLL2 | ENSG00000264364.3 | 23336 | -0.048 | -0.3850 | No | ||
| 66 | GOSR1 | ENSG00000108587.15 | 23341 | -0.048 | -0.3841 | No | ||
| 67 | COPE | ENSG00000105669.14 | 23839 | -0.062 | -0.3950 | No | ||
| 68 | SPTBN5 | ENSG00000137877.10 | 23841 | -0.062 | -0.3937 | No | ||
| 69 | COG1 | ENSG00000166685.12 | 23874 | -0.062 | -0.3932 | No | ||
| 70 | YKT6 | ENSG00000106636.8 | 24433 | -0.075 | -0.4053 | No | ||
| 71 | PREB | ENSG00000138073.14 | 24574 | -0.079 | -0.4071 | No | ||
| 72 | COG3 | ENSG00000136152.15 | 25003 | -0.092 | -0.4156 | No | ||
| 73 | TMED3 | ENSG00000166557.13 | 25030 | -0.093 | -0.4143 | No | ||
| 74 | SERPINA1 | ENSG00000197249.13 | 25161 | -0.095 | -0.4155 | No | ||
| 75 | COPA | ENSG00000122218.16 | 25255 | -0.097 | -0.4158 | No | ||
| 76 | TRAPPC5 | ENSG00000181029.9 | 25277 | -0.098 | -0.4142 | No | ||
| 77 | TBC1D20 | ENSG00000125875.14 | 25340 | -0.100 | -0.4137 | No | ||
| 78 | DYNC1LI2 | ENSG00000135720.13 | 25352 | -0.101 | -0.4119 | No | ||
| 79 | GORASP1 | ENSG00000114745.13 | 25461 | -0.104 | -0.4123 | No | ||
| 80 | MGAT4C | ENSG00000182050.13 | 25597 | -0.108 | -0.4134 | No | ||
| 81 | FOLR1 | ENSG00000110195.13 | 26030 | -0.121 | -0.4214 | No | ||
| 82 | MGAT2 | ENSG00000168282.6 | 26125 | -0.124 | -0.4212 | No | ||
| 83 | NAPA | ENSG00000105402.8 | 26346 | -0.131 | -0.4238 | No | ||
| 84 | TUBA8 | ENSG00000183785.15 | 26534 | -0.136 | -0.4256 | No | ||
| 85 | ARF3 | ENSG00000134287.10 | 26805 | -0.144 | -0.4292 | No | ||
| 86 | LMAN2L | ENSG00000114988.11 | 26893 | -0.146 | -0.4283 | No | ||
| 87 | PPP6R1 | ENSG00000105063.19 | 26915 | -0.147 | -0.4257 | No | ||
| 88 | KDELR2 | ENSG00000136240.10 | 26978 | -0.149 | -0.4242 | No | ||
| 89 | SEC24C | ENSG00000176986.16 | 27265 | -0.157 | -0.4279 | No | ||
| 90 | CNIH1 | ENSG00000100528.12 | 27461 | -0.163 | -0.4293 | No | ||
| 91 | GOSR2 | ENSG00000108433.16 | 27707 | -0.171 | -0.4317 | Yes | ||
| 92 | SEC24D | ENSG00000150961.15 | 27788 | -0.173 | -0.4301 | Yes | ||
| 93 | RAB1B | ENSG00000174903.16 | 27815 | -0.174 | -0.4271 | Yes | ||
| 94 | TUBA3C | ENSG00000198033.12 | 27829 | -0.175 | -0.4238 | Yes | ||
| 95 | ANK3 | ENSG00000151150.22 | 28038 | -0.182 | -0.4251 | Yes | ||
| 96 | SAR1B | ENSG00000152700.14 | 28046 | -0.182 | -0.4215 | Yes | ||
| 97 | B4GALT5 | ENSG00000158470.5 | 28090 | -0.184 | -0.4188 | Yes | ||
| 98 | SEC22A | ENSG00000121542.12 | 28141 | -0.185 | -0.4162 | Yes | ||
| 99 | USO1 | ENSG00000138768.14 | 28192 | -0.186 | -0.4135 | Yes | ||
| 100 | COPZ1 | ENSG00000111481.10 | 28332 | -0.190 | -0.4130 | Yes | ||
| 101 | TRAPPC6B | ENSG00000182400.15 | 28340 | -0.191 | -0.4092 | Yes | ||
| 102 | ANK2 | ENSG00000145362.19 | 28436 | -0.194 | -0.4075 | Yes | ||
| 103 | DCTN4 | ENSG00000132912.12 | 28497 | -0.197 | -0.4049 | Yes | ||
| 104 | MGAT5 | ENSG00000152127.9 | 28699 | -0.202 | -0.4056 | Yes | ||
| 105 | BET1 | ENSG00000105829.13 | 28876 | -0.207 | -0.4056 | Yes | ||
| 106 | SPTBN1 | ENSG00000115306.16 | 29149 | -0.216 | -0.4078 | Yes | ||
| 107 | SEC22B | ENSG00000265808.4 | 29171 | -0.217 | -0.4038 | Yes | ||
| 108 | FUT3 | ENSG00000171124.13 | 29226 | -0.218 | -0.4006 | Yes | ||
| 109 | SEC24B | ENSG00000138802.11 | 29309 | -0.222 | -0.3980 | Yes | ||
| 110 | TUBA3E | ENSG00000152086.9 | 29330 | -0.222 | -0.3939 | Yes | ||
| 111 | TRAPPC10 | ENSG00000160218.13 | 29610 | -0.231 | -0.3959 | Yes | ||
| 112 | SPTAN1 | ENSG00000197694.16 | 29628 | -0.232 | -0.3915 | Yes | ||
| 113 | B4GALT4 | ENSG00000121578.13 | 29751 | -0.236 | -0.3896 | Yes | ||
| 114 | TRAPPC3 | ENSG00000054116.12 | 29771 | -0.237 | -0.3852 | Yes | ||
| 115 | RAB1A | ENSG00000138069.18 | 30150 | -0.249 | -0.3893 | Yes | ||
| 116 | F8 | ENSG00000185010.15 | 30159 | -0.249 | -0.3843 | Yes | ||
| 117 | GBF1 | ENSG00000107862.4 | 30343 | -0.255 | -0.3835 | Yes | ||
| 118 | COG6 | ENSG00000133103.17 | 30452 | -0.258 | -0.3808 | Yes | ||
| 119 | TMED10 | ENSG00000170348.9 | 30459 | -0.259 | -0.3755 | Yes | ||
| 120 | TUBB8 | ENSG00000261456.5 | 30717 | -0.267 | -0.3763 | Yes | ||
| 121 | COPG1 | ENSG00000181789.14 | 30735 | -0.267 | -0.3712 | Yes | ||
| 122 | ARFGAP3 | ENSG00000242247.11 | 30822 | -0.270 | -0.3677 | Yes | ||
| 123 | DCTN5 | ENSG00000166847.10 | 31044 | -0.279 | -0.3673 | Yes | ||
| 124 | CTSZ | ENSG00000101160.14 | 31105 | -0.281 | -0.3629 | Yes | ||
| 125 | TUBB4A | ENSG00000104833.12 | 31106 | -0.281 | -0.3571 | Yes | ||
| 126 | PPP6R3 | ENSG00000110075.14 | 31140 | -0.282 | -0.3521 | Yes | ||
| 127 | SEC31A | ENSG00000138674.17 | 31154 | -0.283 | -0.3465 | Yes | ||
| 128 | DYNC1H1 | ENSG00000197102.12 | 31822 | -0.290 | -0.3568 | Yes | ||
| 129 | COPB2 | ENSG00000184432.10 | 32009 | -0.297 | -0.3552 | Yes | ||
| 130 | SEC22C | ENSG00000093183.14 | 32173 | -0.303 | -0.3529 | Yes | ||
| 131 | SCFD1 | ENSG00000092108.21 | 32791 | -0.327 | -0.3612 | Yes | ||
| 132 | SEC16B | ENSG00000120341.18 | 32844 | -0.330 | -0.3556 | Yes | ||
| 133 | DCTN2 | ENSG00000175203.15 | 32923 | -0.333 | -0.3506 | Yes | ||
| 134 | TFG | ENSG00000114354.14 | 32952 | -0.334 | -0.3444 | Yes | ||
| 135 | TUBAL3 | ENSG00000178462.12 | 33185 | -0.336 | -0.3431 | Yes | ||
| 136 | MAN1A1 | ENSG00000111885.7 | 33550 | -0.338 | -0.3450 | Yes | ||
| 137 | B4GALT6 | ENSG00000118276.11 | 34127 | -0.346 | -0.3519 | Yes | ||
| 138 | TMED7 | ENSG00000134970.14 | 35148 | -0.371 | -0.3691 | Yes | ||
| 139 | PPP6C | ENSG00000119414.11 | 35394 | -0.383 | -0.3671 | Yes | ||
| 140 | CAPZB | ENSG00000077549.18 | 35414 | -0.384 | -0.3596 | Yes | ||
| 141 | MGAT4B | ENSG00000161013.17 | 35657 | -0.396 | -0.3573 | Yes | ||
| 142 | DCTN1 | ENSG00000204843.12 | 35697 | -0.397 | -0.3500 | Yes | ||
| 143 | TRAPPC2 | ENSG00000196459.14 | 35732 | -0.399 | -0.3426 | Yes | ||
| 144 | F5 | ENSG00000198734.11 | 35972 | -0.404 | -0.3400 | Yes | ||
| 145 | TRAPPC1 | ENSG00000170043.12 | 36002 | -0.406 | -0.3323 | Yes | ||
| 146 | SEC23IP | ENSG00000107651.13 | 36113 | -0.410 | -0.3265 | Yes | ||
| 147 | DYNC1I2 | ENSG00000077380.15 | 36153 | -0.411 | -0.3190 | Yes | ||
| 148 | DCTN6 | ENSG00000104671.8 | 36330 | -0.417 | -0.3146 | Yes | ||
| 149 | NSF | ENSG00000073969.18 | 36513 | -0.428 | -0.3102 | Yes | ||
| 150 | ARCN1 | ENSG00000095139.14 | 36550 | -0.430 | -0.3022 | Yes | ||
| 151 | SEC24A | ENSG00000113615.13 | 36665 | -0.434 | -0.2960 | Yes | ||
| 152 | COPB1 | ENSG00000129083.12 | 36728 | -0.437 | -0.2884 | Yes | ||
| 153 | ACTR1A | ENSG00000138107.13 | 36970 | -0.450 | -0.2850 | Yes | ||
| 154 | CNIH2 | ENSG00000174871.11 | 37049 | -0.455 | -0.2775 | Yes | ||
| 155 | TUBA1B | ENSG00000123416.15 | 37411 | -0.478 | -0.2764 | Yes | ||
| 156 | ANKRD28 | ENSG00000206560.11 | 37502 | -0.485 | -0.2685 | Yes | ||
| 157 | B4GALT2 | ENSG00000117411.16 | 37504 | -0.485 | -0.2585 | Yes | ||
| 158 | ARF4 | ENSG00000168374.11 | 37528 | -0.487 | -0.2490 | Yes | ||
| 159 | SEC23A | ENSG00000100934.15 | 37819 | -0.514 | -0.2454 | Yes | ||
| 160 | B4GALT1 | ENSG00000086062.12 | 38033 | -0.530 | -0.2396 | Yes | ||
| 161 | CAPZA2 | ENSG00000198898.14 | 38185 | -0.541 | -0.2321 | Yes | ||
| 162 | ST8SIA2 | ENSG00000140557.12 | 38220 | -0.544 | -0.2217 | Yes | ||
| 163 | DYNC1LI1 | ENSG00000144635.9 | 38271 | -0.548 | -0.2115 | Yes | ||
| 164 | DYNLL1 | ENSG00000088986.11 | 38317 | -0.554 | -0.2012 | Yes | ||
| 165 | TUBA1A | ENSG00000167552.14 | 38598 | -0.561 | -0.1964 | Yes | ||
| 166 | MAN2A1 | ENSG00000112893.9 | 39055 | -0.607 | -0.1949 | Yes | ||
| 167 | COPZ2 | ENSG00000005243.10 | 39066 | -0.608 | -0.1826 | Yes | ||
| 168 | TGFA | ENSG00000163235.16 | 39092 | -0.612 | -0.1705 | Yes | ||
| 169 | CAPZA1 | ENSG00000116489.13 | 39314 | -0.624 | -0.1630 | Yes | ||
| 170 | ACTR10 | ENSG00000131966.14 | 39413 | -0.627 | -0.1524 | Yes | ||
| 171 | COG5 | ENSG00000164597.13 | 39556 | -0.642 | -0.1426 | Yes | ||
| 172 | LHB | ENSG00000104826.14 | 39576 | -0.644 | -0.1297 | Yes | ||
| 173 | MCFD2 | ENSG00000180398.12 | 39625 | -0.650 | -0.1174 | Yes | ||
| 174 | MANEA | ENSG00000172469.16 | 39828 | -0.681 | -0.1082 | Yes | ||
| 175 | COPG2 | ENSG00000158623.14 | 40017 | -0.716 | -0.0980 | Yes | ||
| 176 | TUBB4B | ENSG00000188229.6 | 40106 | -0.733 | -0.0850 | Yes | ||
| 177 | TUBB2B | ENSG00000137285.10 | 40109 | -0.733 | -0.0698 | Yes | ||
| 178 | TUBB2A | ENSG00000137267.6 | 40266 | -0.768 | -0.0577 | Yes | ||
| 179 | MAN1C1 | ENSG00000117643.14 | 40471 | -0.830 | -0.0455 | Yes | ||
| 180 | TUBA1C | ENSG00000167553.15 | 40512 | -0.843 | -0.0290 | Yes | ||
| 181 | TUBB6 | ENSG00000176014.13 | 40817 | -0.979 | -0.0162 | Yes | ||
| 182 | SPTB | ENSG00000070182.21 | 41024 | -1.185 | 0.0033 | Yes |