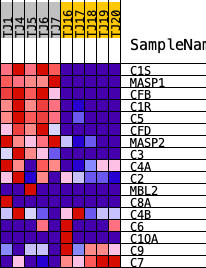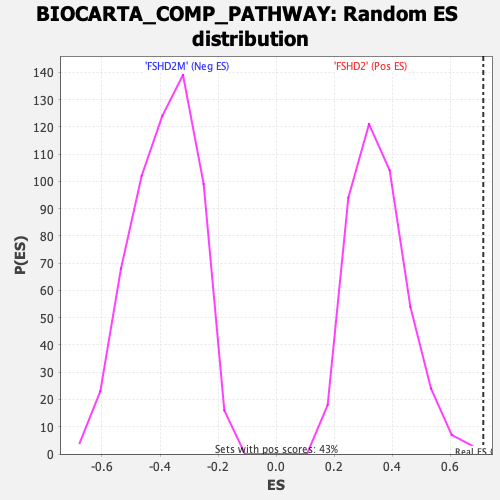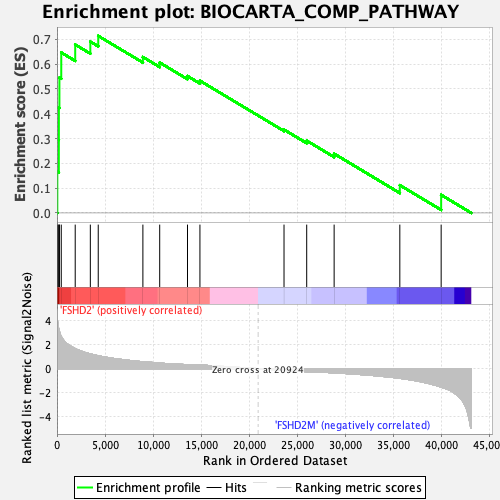
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | BIOCARTA_COMP_PATHWAY |
| Enrichment Score (ES) | 0.7136002 |
| Normalized Enrichment Score (NES) | 2.0230138 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0047104163 |
| FWER p-Value | 0.054 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | C1S | ENSG00000182326.15 | 34 | 4.267 | 0.1669 | Yes | ||
| 2 | MASP1 | ENSG00000127241.16 | 191 | 3.323 | 0.2939 | Yes | ||
| 3 | CFB | ENSG00000243649.8 | 196 | 3.306 | 0.4238 | Yes | ||
| 4 | C1R | ENSG00000159403.18 | 251 | 3.121 | 0.5452 | Yes | ||
| 5 | C5 | ENSG00000106804.7 | 443 | 2.687 | 0.6464 | Yes | ||
| 6 | CFD | ENSG00000197766.8 | 1895 | 1.671 | 0.6784 | Yes | ||
| 7 | MASP2 | ENSG00000009724.17 | 3469 | 1.231 | 0.6903 | Yes | ||
| 8 | C3 | ENSG00000125730.17 | 4292 | 1.078 | 0.7136 | Yes | ||
| 9 | C4A | ENSG00000244731.8 | 8939 | 0.572 | 0.6283 | No | ||
| 10 | C2 | ENSG00000166278.15 | 10689 | 0.466 | 0.6060 | No | ||
| 11 | MBL2 | ENSG00000165471.6 | 13582 | 0.322 | 0.5516 | No | ||
| 12 | C8A | ENSG00000157131.10 | 14876 | 0.287 | 0.5329 | No | ||
| 13 | C4B | ENSG00000224389.9 | 23628 | -0.148 | 0.3357 | No | ||
| 14 | C6 | ENSG00000039537.13 | 25989 | -0.260 | 0.2911 | No | ||
| 15 | C1QA | ENSG00000173372.17 | 28844 | -0.351 | 0.2387 | No | ||
| 16 | C9 | ENSG00000113600.11 | 35681 | -0.807 | 0.1118 | No | ||
| 17 | C7 | ENSG00000112936.19 | 39983 | -1.544 | 0.0728 | No |