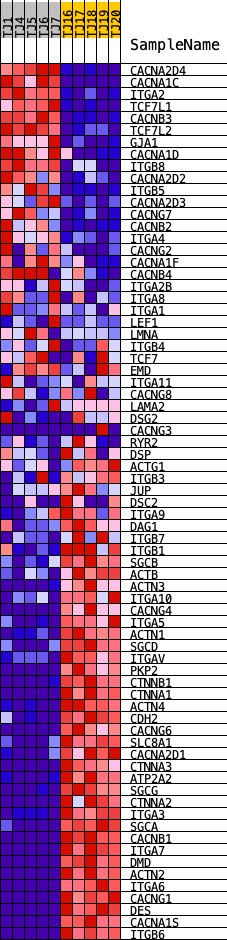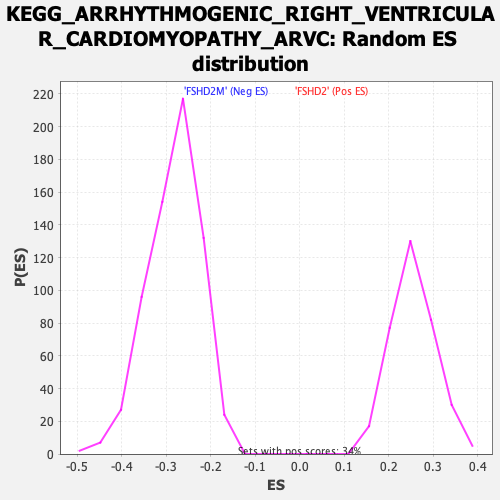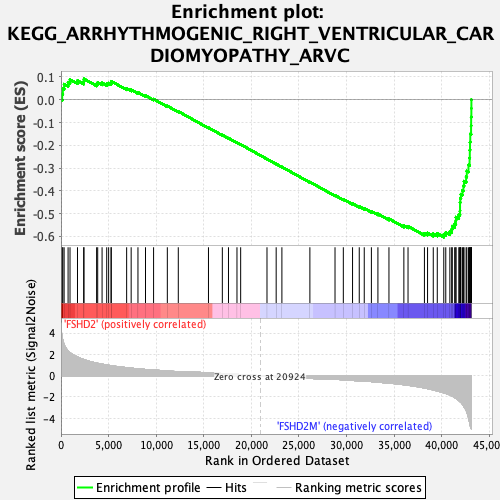
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | KEGG_ARRHYTHMOGENIC_RIGHT_VENTRICULAR_CARDIOMYOPATHY_ARVC |
| Enrichment Score (ES) | -0.6035253 |
| Normalized Enrichment Score (NES) | -2.140604 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.7733868E-4 |
| FWER p-Value | 0.005 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA2D4 | ENSG00000151062.15 | 124 | 3.591 | 0.0250 | No | ||
| 2 | CACNA1C | ENSG00000151067.22 | 193 | 3.311 | 0.0490 | No | ||
| 3 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.0684 | No | ||
| 4 | TCF7L1 | ENSG00000152284.5 | 744 | 2.321 | 0.0768 | No | ||
| 5 | CACNB3 | ENSG00000167535.8 | 947 | 2.135 | 0.0886 | No | ||
| 6 | TCF7L2 | ENSG00000148737.17 | 1732 | 1.743 | 0.0839 | No | ||
| 7 | GJA1 | ENSG00000152661.9 | 2383 | 1.509 | 0.0805 | No | ||
| 8 | CACNA1D | ENSG00000157388.19 | 2414 | 1.501 | 0.0914 | No | ||
| 9 | ITGB8 | ENSG00000105855.10 | 3741 | 1.176 | 0.0698 | No | ||
| 10 | CACNA2D2 | ENSG00000007402.11 | 3855 | 1.154 | 0.0761 | No | ||
| 11 | ITGB5 | ENSG00000082781.12 | 4315 | 1.074 | 0.0737 | No | ||
| 12 | CACNA2D3 | ENSG00000157445.15 | 4802 | 0.997 | 0.0702 | No | ||
| 13 | CACNG7 | ENSG00000105605.7 | 5004 | 0.967 | 0.0730 | No | ||
| 14 | CACNB2 | ENSG00000165995.20 | 5260 | 0.933 | 0.0743 | No | ||
| 15 | ITGA4 | ENSG00000115232.14 | 5278 | 0.930 | 0.0811 | No | ||
| 16 | CACNG2 | ENSG00000166862.6 | 6896 | 0.747 | 0.0494 | No | ||
| 17 | CACNA1F | ENSG00000102001.12 | 7370 | 0.703 | 0.0438 | No | ||
| 18 | CACNB4 | ENSG00000182389.19 | 8084 | 0.640 | 0.0322 | No | ||
| 19 | ITGA2B | ENSG00000005961.19 | 8876 | 0.577 | 0.0183 | No | ||
| 20 | ITGA8 | ENSG00000077943.8 | 9725 | 0.522 | 0.0027 | No | ||
| 21 | ITGA1 | ENSG00000213949.10 | 11169 | 0.434 | -0.0275 | No | ||
| 22 | LEF1 | ENSG00000138795.10 | 12321 | 0.374 | -0.0513 | No | ||
| 23 | LMNA | ENSG00000160789.20 | 15491 | 0.275 | -0.1228 | No | ||
| 24 | ITGB4 | ENSG00000132470.14 | 16947 | 0.199 | -0.1551 | No | ||
| 25 | TCF7 | ENSG00000081059.20 | 17593 | 0.169 | -0.1687 | No | ||
| 26 | EMD | ENSG00000102119.10 | 18485 | 0.125 | -0.1885 | No | ||
| 27 | ITGA11 | ENSG00000137809.17 | 18873 | 0.105 | -0.1966 | No | ||
| 28 | CACNG8 | ENSG00000142408.4 | 21638 | -0.039 | -0.2605 | No | ||
| 29 | LAMA2 | ENSG00000196569.12 | 22603 | -0.091 | -0.2822 | No | ||
| 30 | DSG2 | ENSG00000046604.13 | 23204 | -0.124 | -0.2952 | No | ||
| 31 | CACNG3 | ENSG00000006116.3 | 26144 | -0.263 | -0.3615 | No | ||
| 32 | RYR2 | ENSG00000198626.17 | 28784 | -0.351 | -0.4200 | No | ||
| 33 | DSP | ENSG00000096696.14 | 29657 | -0.385 | -0.4373 | No | ||
| 34 | ACTG1 | ENSG00000184009.12 | 30626 | -0.445 | -0.4563 | No | ||
| 35 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.4689 | No | ||
| 36 | JUP | ENSG00000173801.17 | 31855 | -0.505 | -0.4772 | No | ||
| 37 | DSC2 | ENSG00000134755.17 | 32605 | -0.552 | -0.4903 | No | ||
| 38 | ITGA9 | ENSG00000144668.12 | 33291 | -0.604 | -0.5016 | No | ||
| 39 | DAG1 | ENSG00000173402.11 | 34450 | -0.698 | -0.5230 | No | ||
| 40 | ITGB7 | ENSG00000139626.16 | 36016 | -0.846 | -0.5528 | No | ||
| 41 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.5561 | No | ||
| 42 | SGCB | ENSG00000163069.12 | 38176 | -1.149 | -0.5871 | No | ||
| 43 | ACTB | ENSG00000075624.16 | 38506 | -1.208 | -0.5854 | No | ||
| 44 | ACTN3 | ENSG00000248746.6 | 39105 | -1.332 | -0.5890 | No | ||
| 45 | ITGA10 | ENSG00000143127.13 | 39526 | -1.426 | -0.5877 | No | ||
| 46 | CACNG4 | ENSG00000075461.6 | 40209 | -1.603 | -0.5911 | Yes | ||
| 47 | ITGA5 | ENSG00000161638.11 | 40419 | -1.662 | -0.5831 | Yes | ||
| 48 | ACTN1 | ENSG00000072110.13 | 40848 | -1.801 | -0.5791 | Yes | ||
| 49 | SGCD | ENSG00000170624.13 | 41042 | -1.886 | -0.5689 | Yes | ||
| 50 | ITGAV | ENSG00000138448.12 | 41100 | -1.907 | -0.5555 | Yes | ||
| 51 | PKP2 | ENSG00000057294.14 | 41353 | -2.035 | -0.5455 | Yes | ||
| 52 | CTNNB1 | ENSG00000168036.18 | 41475 | -2.117 | -0.5319 | Yes | ||
| 53 | CTNNA1 | ENSG00000044115.21 | 41488 | -2.125 | -0.5158 | Yes | ||
| 54 | ACTN4 | ENSG00000130402.12 | 41788 | -2.315 | -0.5048 | Yes | ||
| 55 | CDH2 | ENSG00000170558.9 | 41908 | -2.414 | -0.4888 | Yes | ||
| 56 | CACNG6 | ENSG00000130433.7 | 41917 | -2.421 | -0.4702 | Yes | ||
| 57 | SLC8A1 | ENSG00000183023.18 | 41918 | -2.421 | -0.4515 | Yes | ||
| 58 | CACNA2D1 | ENSG00000153956.16 | 41945 | -2.451 | -0.4331 | Yes | ||
| 59 | CTNNA3 | ENSG00000183230.17 | 42006 | -2.494 | -0.4151 | Yes | ||
| 60 | ATP2A2 | ENSG00000174437.17 | 42170 | -2.663 | -0.3983 | Yes | ||
| 61 | SGCG | ENSG00000102683.7 | 42272 | -2.813 | -0.3788 | Yes | ||
| 62 | CTNNA2 | ENSG00000066032.18 | 42337 | -2.897 | -0.3578 | Yes | ||
| 63 | ITGA3 | ENSG00000005884.18 | 42556 | -3.222 | -0.3379 | Yes | ||
| 64 | SGCA | ENSG00000108823.16 | 42629 | -3.407 | -0.3132 | Yes | ||
| 65 | CACNB1 | ENSG00000067191.16 | 42803 | -3.923 | -0.2868 | Yes | ||
| 66 | ITGA7 | ENSG00000135424.16 | 42920 | -4.354 | -0.2557 | Yes | ||
| 67 | DMD | ENSG00000198947.15 | 42943 | -4.488 | -0.2215 | Yes | ||
| 68 | ACTN2 | ENSG00000077522.13 | 42966 | -4.553 | -0.1867 | Yes | ||
| 69 | ITGA6 | ENSG00000091409.15 | 42992 | -4.680 | -0.1510 | Yes | ||
| 70 | CACNG1 | ENSG00000108878.5 | 43077 | -4.944 | -0.1146 | Yes | ||
| 71 | DES | ENSG00000175084.12 | 43086 | -4.954 | -0.0764 | Yes | ||
| 72 | CACNA1S | ENSG00000081248.11 | 43107 | -4.973 | -0.0383 | Yes | ||
| 73 | ITGB6 | ENSG00000115221.12 | 43111 | -4.974 | 0.0002 | Yes |