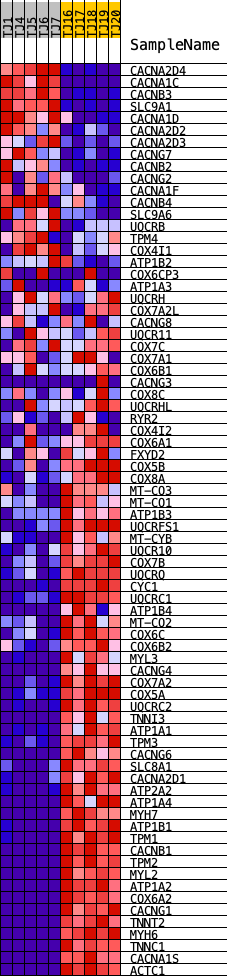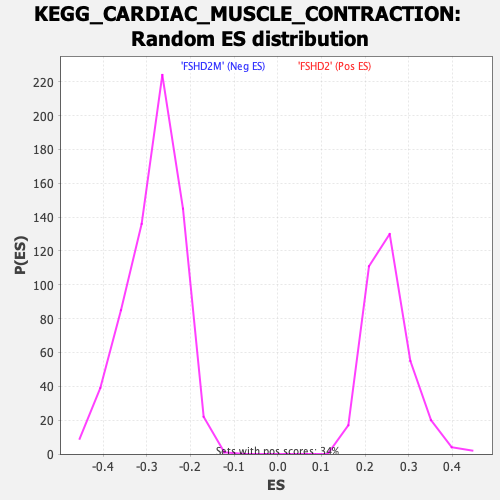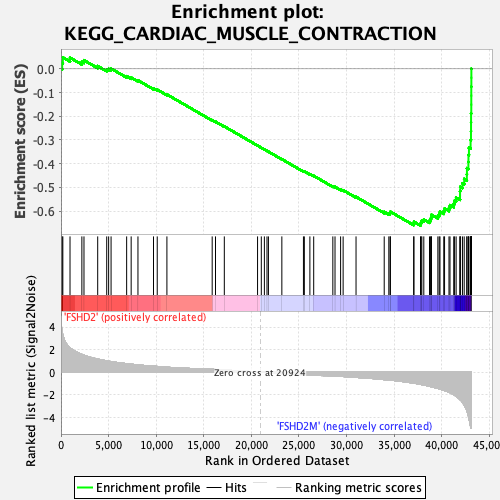
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | KEGG_CARDIAC_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | -0.6612001 |
| Normalized Enrichment Score (NES) | -2.3284388 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA2D4 | ENSG00000151062.15 | 124 | 3.591 | 0.0245 | No | ||
| 2 | CACNA1C | ENSG00000151067.22 | 193 | 3.311 | 0.0481 | No | ||
| 3 | CACNB3 | ENSG00000167535.8 | 947 | 2.135 | 0.0469 | No | ||
| 4 | SLC9A1 | ENSG00000090020.11 | 2192 | 1.568 | 0.0299 | No | ||
| 5 | CACNA1D | ENSG00000157388.19 | 2414 | 1.501 | 0.0362 | No | ||
| 6 | CACNA2D2 | ENSG00000007402.11 | 3855 | 1.154 | 0.0116 | No | ||
| 7 | CACNA2D3 | ENSG00000157445.15 | 4802 | 0.997 | -0.0028 | No | ||
| 8 | CACNG7 | ENSG00000105605.7 | 5004 | 0.967 | -0.0001 | No | ||
| 9 | CACNB2 | ENSG00000165995.20 | 5260 | 0.933 | 0.0011 | No | ||
| 10 | CACNG2 | ENSG00000166862.6 | 6896 | 0.747 | -0.0312 | No | ||
| 11 | CACNA1F | ENSG00000102001.12 | 7370 | 0.703 | -0.0368 | No | ||
| 12 | CACNB4 | ENSG00000182389.19 | 8084 | 0.640 | -0.0485 | No | ||
| 13 | SLC9A6 | ENSG00000198689.11 | 9717 | 0.523 | -0.0825 | No | ||
| 14 | UQCRB | ENSG00000156467.9 | 10109 | 0.503 | -0.0877 | No | ||
| 15 | TPM4 | ENSG00000167460.17 | 11125 | 0.437 | -0.1080 | No | ||
| 16 | COX4I1 | ENSG00000131143.9 | 15881 | 0.255 | -0.2165 | No | ||
| 17 | ATP1B2 | ENSG00000129244.9 | 16230 | 0.234 | -0.2228 | No | ||
| 18 | COX6CP3 | ENSG00000266251.1 | 17155 | 0.190 | -0.2428 | No | ||
| 19 | ATP1A3 | ENSG00000105409.19 | 20648 | 0.014 | -0.3238 | No | ||
| 20 | UQCRH | ENSG00000173660.12 | 21041 | -0.006 | -0.3329 | No | ||
| 21 | COX7A2L | ENSG00000115944.15 | 21364 | -0.024 | -0.3402 | No | ||
| 22 | CACNG8 | ENSG00000142408.4 | 21638 | -0.039 | -0.3462 | No | ||
| 23 | UQCR11 | ENSG00000127540.12 | 21789 | -0.046 | -0.3494 | No | ||
| 24 | COX7C | ENSG00000127184.13 | 23199 | -0.124 | -0.3812 | No | ||
| 25 | COX7A1 | ENSG00000161281.11 | 25467 | -0.225 | -0.4321 | No | ||
| 26 | COX6B1 | ENSG00000126267.11 | 25561 | -0.231 | -0.4325 | No | ||
| 27 | CACNG3 | ENSG00000006116.3 | 26144 | -0.263 | -0.4440 | No | ||
| 28 | COX8C | ENSG00000187581.3 | 26545 | -0.269 | -0.4513 | No | ||
| 29 | UQCRHL | ENSG00000233954.6 | 28555 | -0.340 | -0.4954 | No | ||
| 30 | RYR2 | ENSG00000198626.17 | 28784 | -0.351 | -0.4980 | No | ||
| 31 | COX4I2 | ENSG00000131055.5 | 29371 | -0.368 | -0.5088 | No | ||
| 32 | COX6A1 | ENSG00000111775.3 | 29637 | -0.384 | -0.5120 | No | ||
| 33 | FXYD2 | ENSG00000137731.14 | 30992 | -0.458 | -0.5400 | No | ||
| 34 | COX5B | ENSG00000135940.7 | 33952 | -0.658 | -0.6037 | No | ||
| 35 | COX8A | ENSG00000176340.4 | 34436 | -0.697 | -0.6096 | No | ||
| 36 | MT-CO3 | ENSG00000198938.2 | 34569 | -0.708 | -0.6073 | No | ||
| 37 | MT-CO1 | ENSG00000198804.2 | 34603 | -0.710 | -0.6027 | No | ||
| 38 | ATP1B3 | ENSG00000069849.11 | 37044 | -0.973 | -0.6519 | No | ||
| 39 | UQCRFS1 | ENSG00000169021.6 | 37078 | -0.978 | -0.6453 | No | ||
| 40 | MT-CYB | ENSG00000198727.2 | 37765 | -1.078 | -0.6530 | Yes | ||
| 41 | UQCR10 | ENSG00000184076.13 | 37826 | -1.087 | -0.6461 | Yes | ||
| 42 | COX7B | ENSG00000131174.7 | 37903 | -1.101 | -0.6395 | Yes | ||
| 43 | UQCRQ | ENSG00000164405.11 | 38111 | -1.136 | -0.6356 | Yes | ||
| 44 | CYC1 | ENSG00000179091.5 | 38710 | -1.245 | -0.6400 | Yes | ||
| 45 | UQCRC1 | ENSG00000010256.11 | 38824 | -1.273 | -0.6330 | Yes | ||
| 46 | ATP1B4 | ENSG00000101892.12 | 38881 | -1.285 | -0.6245 | Yes | ||
| 47 | MT-CO2 | ENSG00000198712.1 | 38915 | -1.292 | -0.6154 | Yes | ||
| 48 | COX6C | ENSG00000164919.11 | 39588 | -1.439 | -0.6200 | Yes | ||
| 49 | COX6B2 | ENSG00000160471.13 | 39727 | -1.474 | -0.6120 | Yes | ||
| 50 | MYL3 | ENSG00000160808.10 | 39809 | -1.495 | -0.6025 | Yes | ||
| 51 | CACNG4 | ENSG00000075461.6 | 40209 | -1.603 | -0.5996 | Yes | ||
| 52 | COX7A2 | ENSG00000112695.11 | 40291 | -1.624 | -0.5891 | Yes | ||
| 53 | COX5A | ENSG00000178741.12 | 40756 | -1.769 | -0.5864 | Yes | ||
| 54 | UQCRC2 | ENSG00000140740.11 | 40862 | -1.806 | -0.5751 | Yes | ||
| 55 | TNNI3 | ENSG00000129991.13 | 41255 | -1.980 | -0.5691 | Yes | ||
| 56 | ATP1A1 | ENSG00000163399.16 | 41345 | -2.032 | -0.5557 | Yes | ||
| 57 | TPM3 | ENSG00000143549.21 | 41507 | -2.135 | -0.5432 | Yes | ||
| 58 | CACNG6 | ENSG00000130433.7 | 41917 | -2.421 | -0.5342 | Yes | ||
| 59 | SLC8A1 | ENSG00000183023.18 | 41918 | -2.421 | -0.5158 | Yes | ||
| 60 | CACNA2D1 | ENSG00000153956.16 | 41945 | -2.451 | -0.4977 | Yes | ||
| 61 | ATP2A2 | ENSG00000174437.17 | 42170 | -2.663 | -0.4826 | Yes | ||
| 62 | ATP1A4 | ENSG00000132681.16 | 42353 | -2.919 | -0.4646 | Yes | ||
| 63 | MYH7 | ENSG00000092054.13 | 42632 | -3.422 | -0.4450 | Yes | ||
| 64 | ATP1B1 | ENSG00000143153.13 | 42642 | -3.447 | -0.4190 | Yes | ||
| 65 | TPM1 | ENSG00000140416.21 | 42794 | -3.877 | -0.3929 | Yes | ||
| 66 | CACNB1 | ENSG00000067191.16 | 42803 | -3.923 | -0.3632 | Yes | ||
| 67 | TPM2 | ENSG00000198467.15 | 42854 | -4.136 | -0.3329 | Yes | ||
| 68 | MYL2 | ENSG00000111245.15 | 43025 | -4.822 | -0.3001 | Yes | ||
| 69 | ATP1A2 | ENSG00000018625.15 | 43067 | -4.924 | -0.2635 | Yes | ||
| 70 | COX6A2 | ENSG00000156885.6 | 43073 | -4.938 | -0.2260 | Yes | ||
| 71 | CACNG1 | ENSG00000108878.5 | 43077 | -4.944 | -0.1884 | Yes | ||
| 72 | TNNT2 | ENSG00000118194.20 | 43098 | -4.967 | -0.1511 | Yes | ||
| 73 | MYH6 | ENSG00000197616.12 | 43099 | -4.967 | -0.1132 | Yes | ||
| 74 | TNNC1 | ENSG00000114854.8 | 43100 | -4.968 | -0.0754 | Yes | ||
| 75 | CACNA1S | ENSG00000081248.11 | 43107 | -4.973 | -0.0376 | Yes | ||
| 76 | ACTC1 | ENSG00000159251.8 | 43108 | -4.973 | 0.0003 | Yes |