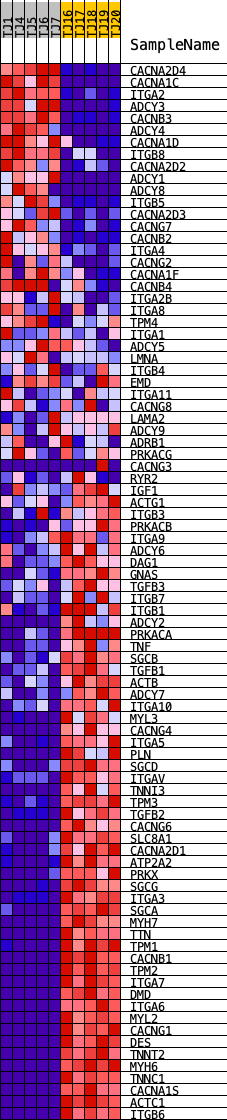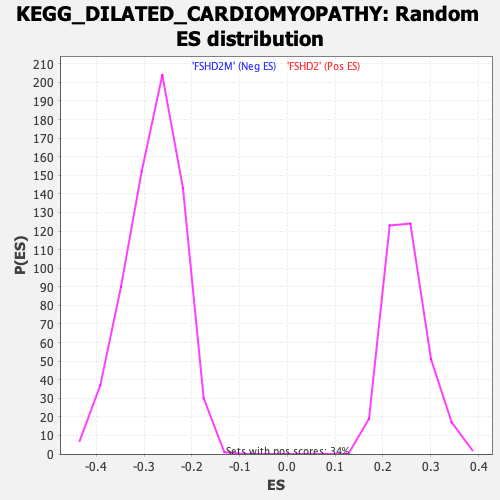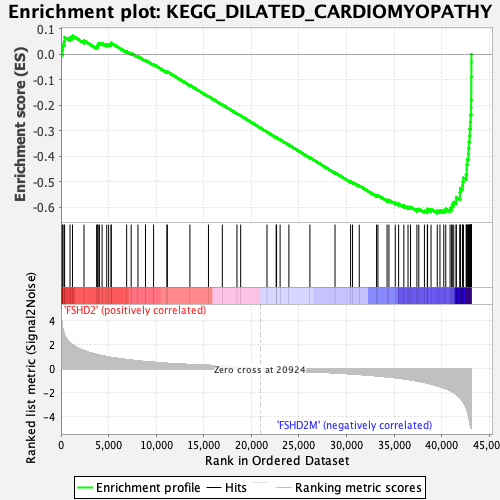
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | KEGG_DILATED_CARDIOMYOPATHY |
| Enrichment Score (ES) | -0.6225343 |
| Normalized Enrichment Score (NES) | -2.2293222 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.7331177E-4 |
| FWER p-Value | 0.001 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CACNA2D4 | ENSG00000151062.15 | 124 | 3.591 | 0.0187 | No | ||
| 2 | CACNA1C | ENSG00000151067.22 | 193 | 3.311 | 0.0371 | No | ||
| 3 | ITGA2 | ENSG00000164171.11 | 327 | 2.899 | 0.0514 | No | ||
| 4 | ADCY3 | ENSG00000138031.14 | 378 | 2.794 | 0.0671 | No | ||
| 5 | CACNB3 | ENSG00000167535.8 | 947 | 2.135 | 0.0668 | No | ||
| 6 | ADCY4 | ENSG00000129467.13 | 1210 | 1.984 | 0.0726 | No | ||
| 7 | CACNA1D | ENSG00000157388.19 | 2414 | 1.501 | 0.0537 | No | ||
| 8 | ITGB8 | ENSG00000105855.10 | 3741 | 1.176 | 0.0300 | No | ||
| 9 | CACNA2D2 | ENSG00000007402.11 | 3855 | 1.154 | 0.0343 | No | ||
| 10 | ADCY1 | ENSG00000164742.16 | 3895 | 1.146 | 0.0403 | No | ||
| 11 | ADCY8 | ENSG00000155897.10 | 4038 | 1.120 | 0.0437 | No | ||
| 12 | ITGB5 | ENSG00000082781.12 | 4315 | 1.074 | 0.0438 | No | ||
| 13 | CACNA2D3 | ENSG00000157445.15 | 4802 | 0.997 | 0.0385 | No | ||
| 14 | CACNG7 | ENSG00000105605.7 | 5004 | 0.967 | 0.0396 | No | ||
| 15 | CACNB2 | ENSG00000165995.20 | 5260 | 0.933 | 0.0393 | No | ||
| 16 | ITGA4 | ENSG00000115232.14 | 5278 | 0.930 | 0.0445 | No | ||
| 17 | CACNG2 | ENSG00000166862.6 | 6896 | 0.747 | 0.0114 | No | ||
| 18 | CACNA1F | ENSG00000102001.12 | 7370 | 0.703 | 0.0047 | No | ||
| 19 | CACNB4 | ENSG00000182389.19 | 8084 | 0.640 | -0.0080 | No | ||
| 20 | ITGA2B | ENSG00000005961.19 | 8876 | 0.577 | -0.0229 | No | ||
| 21 | ITGA8 | ENSG00000077943.8 | 9725 | 0.522 | -0.0395 | No | ||
| 22 | TPM4 | ENSG00000167460.17 | 11125 | 0.437 | -0.0694 | No | ||
| 23 | ITGA1 | ENSG00000213949.10 | 11169 | 0.434 | -0.0678 | No | ||
| 24 | ADCY5 | ENSG00000173175.14 | 13539 | 0.324 | -0.1209 | No | ||
| 25 | LMNA | ENSG00000160789.20 | 15491 | 0.275 | -0.1646 | No | ||
| 26 | ITGB4 | ENSG00000132470.14 | 16947 | 0.199 | -0.1972 | No | ||
| 27 | EMD | ENSG00000102119.10 | 18485 | 0.125 | -0.2321 | No | ||
| 28 | ITGA11 | ENSG00000137809.17 | 18873 | 0.105 | -0.2405 | No | ||
| 29 | CACNG8 | ENSG00000142408.4 | 21638 | -0.039 | -0.3045 | No | ||
| 30 | LAMA2 | ENSG00000196569.12 | 22603 | -0.091 | -0.3263 | No | ||
| 31 | ADCY9 | ENSG00000162104.10 | 22643 | -0.093 | -0.3267 | No | ||
| 32 | ADRB1 | ENSG00000043591.6 | 23016 | -0.113 | -0.3347 | No | ||
| 33 | PRKACG | ENSG00000165059.7 | 23940 | -0.168 | -0.3551 | No | ||
| 34 | CACNG3 | ENSG00000006116.3 | 26144 | -0.263 | -0.4047 | No | ||
| 35 | RYR2 | ENSG00000198626.17 | 28784 | -0.351 | -0.4639 | No | ||
| 36 | IGF1 | ENSG00000017427.16 | 30420 | -0.433 | -0.4993 | No | ||
| 37 | ACTG1 | ENSG00000184009.12 | 30626 | -0.445 | -0.5014 | No | ||
| 38 | ITGB3 | ENSG00000259207.7 | 31330 | -0.481 | -0.5148 | No | ||
| 39 | PRKACB | ENSG00000142875.19 | 33148 | -0.594 | -0.5535 | No | ||
| 40 | ITGA9 | ENSG00000144668.12 | 33291 | -0.604 | -0.5532 | No | ||
| 41 | ADCY6 | ENSG00000174233.11 | 34252 | -0.679 | -0.5714 | No | ||
| 42 | DAG1 | ENSG00000173402.11 | 34450 | -0.698 | -0.5718 | No | ||
| 43 | GNAS | ENSG00000087460.25 | 35113 | -0.751 | -0.5826 | No | ||
| 44 | TGFB3 | ENSG00000119699.7 | 35467 | -0.784 | -0.5861 | No | ||
| 45 | ITGB7 | ENSG00000139626.16 | 36016 | -0.846 | -0.5937 | No | ||
| 46 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.5985 | No | ||
| 47 | ADCY2 | ENSG00000078295.17 | 36714 | -0.931 | -0.5989 | No | ||
| 48 | PRKACA | ENSG00000072062.14 | 37389 | -1.018 | -0.6085 | No | ||
| 49 | TNF | ENSG00000232810.4 | 37586 | -1.048 | -0.6067 | No | ||
| 50 | SGCB | ENSG00000163069.12 | 38176 | -1.149 | -0.6135 | No | ||
| 51 | TGFB1 | ENSG00000105329.10 | 38478 | -1.201 | -0.6132 | No | ||
| 52 | ACTB | ENSG00000075624.16 | 38506 | -1.208 | -0.6066 | No | ||
| 53 | ADCY7 | ENSG00000121281.12 | 38870 | -1.283 | -0.6073 | No | ||
| 54 | ITGA10 | ENSG00000143127.13 | 39526 | -1.426 | -0.6139 | Yes | ||
| 55 | MYL3 | ENSG00000160808.10 | 39809 | -1.495 | -0.6115 | Yes | ||
| 56 | CACNG4 | ENSG00000075461.6 | 40209 | -1.603 | -0.6111 | Yes | ||
| 57 | ITGA5 | ENSG00000161638.11 | 40419 | -1.662 | -0.6060 | Yes | ||
| 58 | PLN | ENSG00000198523.6 | 40873 | -1.812 | -0.6056 | Yes | ||
| 59 | SGCD | ENSG00000170624.13 | 41042 | -1.886 | -0.5981 | Yes | ||
| 60 | ITGAV | ENSG00000138448.12 | 41100 | -1.907 | -0.5880 | Yes | ||
| 61 | TNNI3 | ENSG00000129991.13 | 41255 | -1.980 | -0.5797 | Yes | ||
| 62 | TPM3 | ENSG00000143549.21 | 41507 | -2.135 | -0.5726 | Yes | ||
| 63 | TGFB2 | ENSG00000092969.12 | 41517 | -2.139 | -0.5600 | Yes | ||
| 64 | CACNG6 | ENSG00000130433.7 | 41917 | -2.421 | -0.5547 | Yes | ||
| 65 | SLC8A1 | ENSG00000183023.18 | 41918 | -2.421 | -0.5401 | Yes | ||
| 66 | CACNA2D1 | ENSG00000153956.16 | 41945 | -2.451 | -0.5260 | Yes | ||
| 67 | ATP2A2 | ENSG00000174437.17 | 42170 | -2.663 | -0.5151 | Yes | ||
| 68 | PRKX | ENSG00000183943.6 | 42211 | -2.722 | -0.4997 | Yes | ||
| 69 | SGCG | ENSG00000102683.7 | 42272 | -2.813 | -0.4841 | Yes | ||
| 70 | ITGA3 | ENSG00000005884.18 | 42556 | -3.222 | -0.4713 | Yes | ||
| 71 | SGCA | ENSG00000108823.16 | 42629 | -3.407 | -0.4525 | Yes | ||
| 72 | MYH7 | ENSG00000092054.13 | 42632 | -3.422 | -0.4319 | Yes | ||
| 73 | TTN | ENSG00000155657.26 | 42690 | -3.572 | -0.4118 | Yes | ||
| 74 | TPM1 | ENSG00000140416.21 | 42794 | -3.877 | -0.3908 | Yes | ||
| 75 | CACNB1 | ENSG00000067191.16 | 42803 | -3.923 | -0.3674 | Yes | ||
| 76 | TPM2 | ENSG00000198467.15 | 42854 | -4.136 | -0.3436 | Yes | ||
| 77 | ITGA7 | ENSG00000135424.16 | 42920 | -4.354 | -0.3189 | Yes | ||
| 78 | DMD | ENSG00000198947.15 | 42943 | -4.488 | -0.2924 | Yes | ||
| 79 | ITGA6 | ENSG00000091409.15 | 42992 | -4.680 | -0.2654 | Yes | ||
| 80 | MYL2 | ENSG00000111245.15 | 43025 | -4.822 | -0.2371 | Yes | ||
| 81 | CACNG1 | ENSG00000108878.5 | 43077 | -4.944 | -0.2085 | Yes | ||
| 82 | DES | ENSG00000175084.12 | 43086 | -4.954 | -0.1789 | Yes | ||
| 83 | TNNT2 | ENSG00000118194.20 | 43098 | -4.967 | -0.1492 | Yes | ||
| 84 | MYH6 | ENSG00000197616.12 | 43099 | -4.967 | -0.1193 | Yes | ||
| 85 | TNNC1 | ENSG00000114854.8 | 43100 | -4.968 | -0.0894 | Yes | ||
| 86 | CACNA1S | ENSG00000081248.11 | 43107 | -4.973 | -0.0596 | Yes | ||
| 87 | ACTC1 | ENSG00000159251.8 | 43108 | -4.973 | -0.0297 | Yes | ||
| 88 | ITGB6 | ENSG00000115221.12 | 43111 | -4.974 | 0.0002 | Yes |