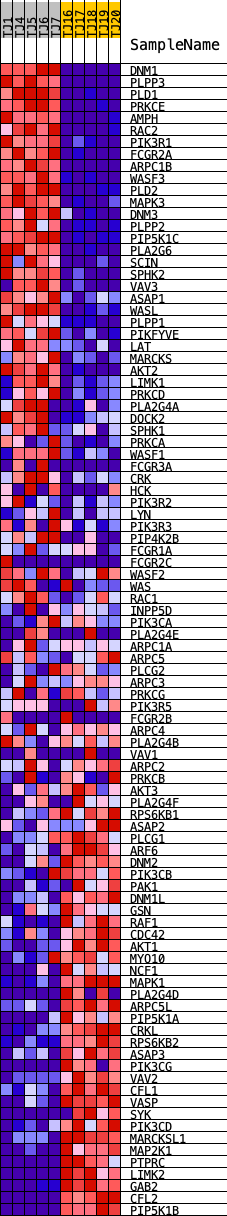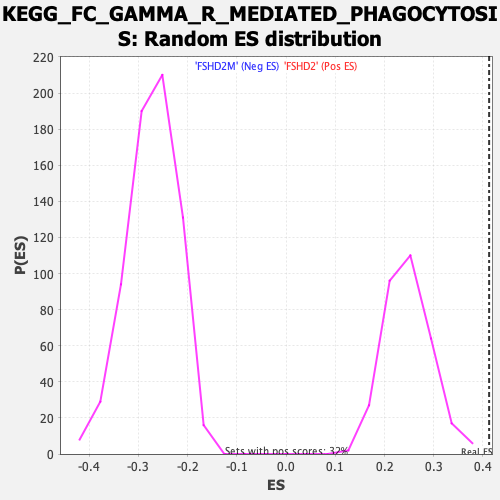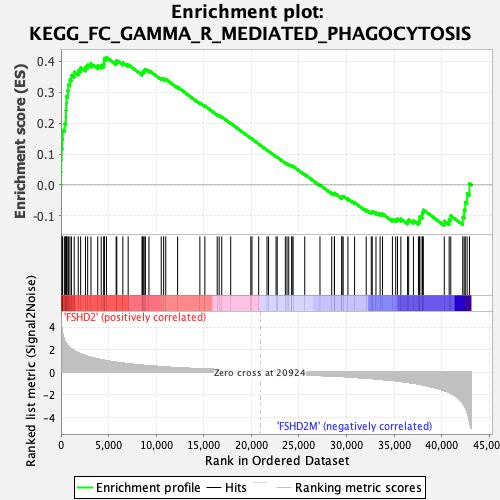
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_FC_GAMMA_R_MEDIATED_PHAGOCYTOSIS |
| Enrichment Score (ES) | 0.4124469 |
| Normalized Enrichment Score (NES) | 1.6732929 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07983056 |
| FWER p-Value | 0.983 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DNM1 | ENSG00000106976.20 | 8 | 4.620 | 0.0419 | Yes | ||
| 2 | PLPP3 | ENSG00000162407.9 | 18 | 4.425 | 0.0820 | Yes | ||
| 3 | PLD1 | ENSG00000075651.16 | 72 | 3.920 | 0.1164 | Yes | ||
| 4 | PRKCE | ENSG00000171132.14 | 128 | 3.568 | 0.1476 | Yes | ||
| 5 | AMPH | ENSG00000078053.17 | 178 | 3.355 | 0.1770 | Yes | ||
| 6 | RAC2 | ENSG00000128340.15 | 379 | 2.785 | 0.1978 | Yes | ||
| 7 | PIK3R1 | ENSG00000145675.15 | 513 | 2.587 | 0.2182 | Yes | ||
| 8 | FCGR2A | ENSG00000143226.13 | 515 | 2.585 | 0.2417 | Yes | ||
| 9 | ARPC1B | ENSG00000130429.15 | 552 | 2.541 | 0.2640 | Yes | ||
| 10 | WASF3 | ENSG00000132970.14 | 571 | 2.508 | 0.2865 | Yes | ||
| 11 | PLD2 | ENSG00000129219.14 | 722 | 2.348 | 0.3043 | Yes | ||
| 12 | MAPK3 | ENSG00000102882.12 | 770 | 2.292 | 0.3241 | Yes | ||
| 13 | DNM3 | ENSG00000197959.14 | 939 | 2.137 | 0.3397 | Yes | ||
| 14 | PLPP2 | ENSG00000141934.10 | 1096 | 2.052 | 0.3547 | Yes | ||
| 15 | PIP5K1C | ENSG00000186111.9 | 1387 | 1.887 | 0.3652 | Yes | ||
| 16 | PLA2G6 | ENSG00000184381.20 | 1810 | 1.710 | 0.3709 | Yes | ||
| 17 | SCIN | ENSG00000006747.15 | 2064 | 1.608 | 0.3797 | Yes | ||
| 18 | SPHK2 | ENSG00000063176.16 | 2570 | 1.450 | 0.3812 | Yes | ||
| 19 | VAV3 | ENSG00000134215.16 | 2801 | 1.377 | 0.3884 | Yes | ||
| 20 | ASAP1 | ENSG00000153317.15 | 3147 | 1.302 | 0.3922 | Yes | ||
| 21 | WASL | ENSG00000106299.8 | 3858 | 1.154 | 0.3862 | Yes | ||
| 22 | PLPP1 | ENSG00000067113.17 | 4224 | 1.088 | 0.3876 | Yes | ||
| 23 | PIKFYVE | ENSG00000115020.17 | 4506 | 1.043 | 0.3906 | Yes | ||
| 24 | LAT | ENSG00000213658.11 | 4530 | 1.040 | 0.3995 | Yes | ||
| 25 | MARCKS | ENSG00000277443.3 | 4535 | 1.038 | 0.4089 | Yes | ||
| 26 | AKT2 | ENSG00000105221.18 | 4775 | 1.001 | 0.4124 | Yes | ||
| 27 | LIMK1 | ENSG00000106683.15 | 5792 | 0.865 | 0.3967 | No | ||
| 28 | PRKCD | ENSG00000163932.15 | 5863 | 0.856 | 0.4029 | No | ||
| 29 | PLA2G4A | ENSG00000116711.10 | 6498 | 0.788 | 0.3953 | No | ||
| 30 | DOCK2 | ENSG00000134516.17 | 7056 | 0.733 | 0.3890 | No | ||
| 31 | SPHK1 | ENSG00000176170.13 | 8512 | 0.605 | 0.3607 | No | ||
| 32 | PRKCA | ENSG00000154229.12 | 8575 | 0.601 | 0.3648 | No | ||
| 33 | WASF1 | ENSG00000112290.13 | 8645 | 0.596 | 0.3686 | No | ||
| 34 | FCGR3A | ENSG00000203747.11 | 8811 | 0.582 | 0.3700 | No | ||
| 35 | CRK | ENSG00000167193.8 | 8864 | 0.578 | 0.3741 | No | ||
| 36 | HCK | ENSG00000101336.14 | 9238 | 0.552 | 0.3705 | No | ||
| 37 | PIK3R2 | ENSG00000105647.18 | 10529 | 0.476 | 0.3448 | No | ||
| 38 | LYN | ENSG00000254087.8 | 10784 | 0.460 | 0.3431 | No | ||
| 39 | PIK3R3 | ENSG00000117461.15 | 11004 | 0.444 | 0.3421 | No | ||
| 40 | PIP4K2B | ENSG00000276293.4 | 12248 | 0.375 | 0.3166 | No | ||
| 41 | FCGR1A | ENSG00000150337.13 | 14587 | 0.305 | 0.2650 | No | ||
| 42 | FCGR2C | ENSG00000244682.7 | 15111 | 0.287 | 0.2555 | No | ||
| 43 | WASF2 | ENSG00000158195.11 | 16408 | 0.225 | 0.2274 | No | ||
| 44 | WAS | ENSG00000015285.10 | 16612 | 0.214 | 0.2246 | No | ||
| 45 | RAC1 | ENSG00000136238.18 | 16886 | 0.203 | 0.2201 | No | ||
| 46 | INPP5D | ENSG00000168918.14 | 17826 | 0.156 | 0.1997 | No | ||
| 47 | PIK3CA | ENSG00000121879.5 | 19940 | 0.051 | 0.1511 | No | ||
| 48 | PLA2G4E | ENSG00000188089.13 | 20059 | 0.045 | 0.1487 | No | ||
| 49 | ARPC1A | ENSG00000241685.10 | 20765 | 0.009 | 0.1324 | No | ||
| 50 | ARPC5 | ENSG00000162704.16 | 21640 | -0.039 | 0.1125 | No | ||
| 51 | PLCG2 | ENSG00000197943.10 | 21804 | -0.047 | 0.1091 | No | ||
| 52 | ARPC3 | ENSG00000111229.16 | 22592 | -0.090 | 0.0916 | No | ||
| 53 | PRKCG | ENSG00000126583.11 | 22711 | -0.097 | 0.0898 | No | ||
| 54 | PIK3R5 | ENSG00000141506.13 | 23567 | -0.145 | 0.0712 | No | ||
| 55 | FCGR2B | ENSG00000072694.20 | 23742 | -0.153 | 0.0686 | No | ||
| 56 | ARPC4 | ENSG00000241553.12 | 23924 | -0.167 | 0.0659 | No | ||
| 57 | PLA2G4B | ENSG00000243708.10 | 24245 | -0.185 | 0.0601 | No | ||
| 58 | VAV1 | ENSG00000141968.8 | 24255 | -0.186 | 0.0616 | No | ||
| 59 | ARPC2 | ENSG00000163466.16 | 24381 | -0.192 | 0.0604 | No | ||
| 60 | PRKCB | ENSG00000166501.14 | 25604 | -0.234 | 0.0342 | No | ||
| 61 | AKT3 | ENSG00000117020.18 | 27205 | -0.288 | -0.0004 | No | ||
| 62 | PLA2G4F | ENSG00000168907.13 | 28437 | -0.334 | -0.0260 | No | ||
| 63 | RPS6KB1 | ENSG00000108443.14 | 28720 | -0.346 | -0.0294 | No | ||
| 64 | ASAP2 | ENSG00000151693.11 | 28722 | -0.346 | -0.0262 | No | ||
| 65 | PLCG1 | ENSG00000124181.14 | 29500 | -0.374 | -0.0409 | No | ||
| 66 | ARF6 | ENSG00000165527.7 | 29502 | -0.374 | -0.0375 | No | ||
| 67 | DNM2 | ENSG00000079805.16 | 29641 | -0.384 | -0.0372 | No | ||
| 68 | PIK3CB | ENSG00000051382.8 | 30141 | -0.417 | -0.0450 | No | ||
| 69 | PAK1 | ENSG00000149269.9 | 30840 | -0.456 | -0.0571 | No | ||
| 70 | DNM1L | ENSG00000087470.18 | 32060 | -0.519 | -0.0807 | No | ||
| 71 | GSN | ENSG00000148180.19 | 32592 | -0.551 | -0.0880 | No | ||
| 72 | RAF1 | ENSG00000132155.11 | 32694 | -0.559 | -0.0853 | No | ||
| 73 | CDC42 | ENSG00000070831.17 | 33086 | -0.589 | -0.0890 | No | ||
| 74 | AKT1 | ENSG00000142208.16 | 33520 | -0.623 | -0.0934 | No | ||
| 75 | MYO10 | ENSG00000145555.15 | 33776 | -0.643 | -0.0935 | No | ||
| 76 | NCF1 | ENSG00000158517.15 | 34813 | -0.725 | -0.1110 | No | ||
| 77 | MAPK1 | ENSG00000100030.14 | 35160 | -0.756 | -0.1121 | No | ||
| 78 | PLA2G4D | ENSG00000159337.7 | 35348 | -0.772 | -0.1094 | No | ||
| 79 | ARPC5L | ENSG00000136950.13 | 35702 | -0.809 | -0.1103 | No | ||
| 80 | PIP5K1A | ENSG00000143398.20 | 36406 | -0.889 | -0.1185 | No | ||
| 81 | CRKL | ENSG00000099942.13 | 36521 | -0.903 | -0.1129 | No | ||
| 82 | RPS6KB2 | ENSG00000175634.15 | 37020 | -0.968 | -0.1157 | No | ||
| 83 | ASAP3 | ENSG00000088280.19 | 37547 | -1.041 | -0.1184 | No | ||
| 84 | PIK3CG | ENSG00000105851.11 | 37680 | -1.063 | -0.1118 | No | ||
| 85 | VAV2 | ENSG00000160293.17 | 37686 | -1.065 | -0.1022 | No | ||
| 86 | CFL1 | ENSG00000172757.13 | 37946 | -1.108 | -0.0982 | No | ||
| 87 | VASP | ENSG00000125753.14 | 37949 | -1.108 | -0.0881 | No | ||
| 88 | SYK | ENSG00000165025.15 | 38062 | -1.127 | -0.0805 | No | ||
| 89 | PIK3CD | ENSG00000171608.16 | 40260 | -1.616 | -0.1168 | No | ||
| 90 | MARCKSL1 | ENSG00000175130.7 | 40775 | -1.776 | -0.1126 | No | ||
| 91 | MAP2K1 | ENSG00000169032.9 | 40944 | -1.840 | -0.0998 | No | ||
| 92 | PTPRC | ENSG00000081237.20 | 42203 | -2.706 | -0.1043 | No | ||
| 93 | LIMK2 | ENSG00000182541.18 | 42346 | -2.914 | -0.0811 | No | ||
| 94 | GAB2 | ENSG00000033327.13 | 42468 | -3.087 | -0.0558 | No | ||
| 95 | CFL2 | ENSG00000165410.15 | 42663 | -3.483 | -0.0286 | No | ||
| 96 | PIP5K1B | ENSG00000107242.19 | 42907 | -4.305 | 0.0049 | No |