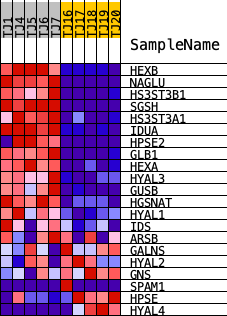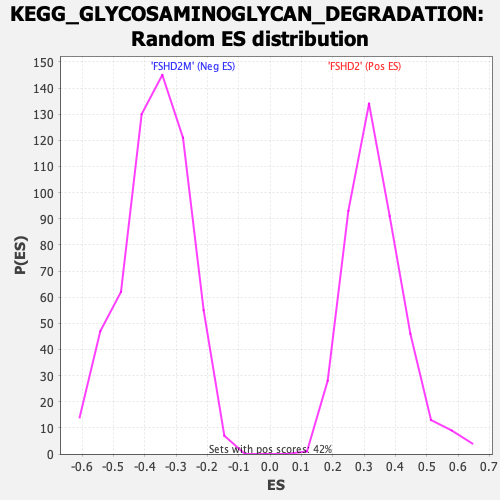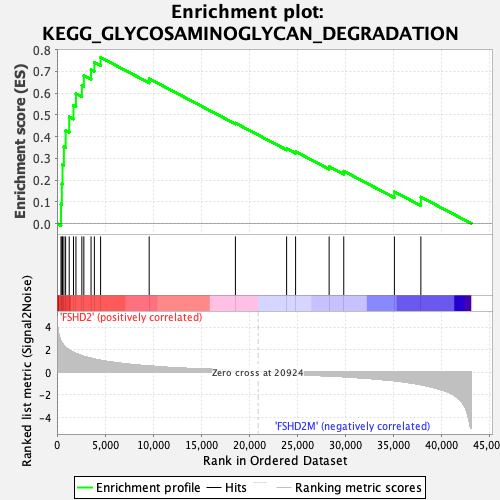
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_GLYCOSAMINOGLYCAN_DEGRADATION |
| Enrichment Score (ES) | 0.76408863 |
| Normalized Enrichment Score (NES) | 2.2700171 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HEXB | ENSG00000049860.14 | 415 | 2.734 | 0.0897 | Yes | ||
| 2 | NAGLU | ENSG00000108784.10 | 504 | 2.596 | 0.1819 | Yes | ||
| 3 | HS3ST3B1 | ENSG00000125430.9 | 574 | 2.503 | 0.2712 | Yes | ||
| 4 | SGSH | ENSG00000181523.13 | 719 | 2.352 | 0.3532 | Yes | ||
| 5 | HS3ST3A1 | ENSG00000153976.3 | 898 | 2.175 | 0.4281 | Yes | ||
| 6 | IDUA | ENSG00000127415.13 | 1270 | 1.956 | 0.4905 | Yes | ||
| 7 | HPSE2 | ENSG00000172987.12 | 1715 | 1.749 | 0.5437 | Yes | ||
| 8 | GLB1 | ENSG00000170266.16 | 1972 | 1.642 | 0.5974 | Yes | ||
| 9 | HEXA | ENSG00000213614.9 | 2587 | 1.446 | 0.6356 | Yes | ||
| 10 | HYAL3 | ENSG00000186792.17 | 2793 | 1.380 | 0.6810 | Yes | ||
| 11 | GUSB | ENSG00000169919.17 | 3543 | 1.217 | 0.7078 | Yes | ||
| 12 | HGSNAT | ENSG00000165102.15 | 3889 | 1.147 | 0.7414 | Yes | ||
| 13 | HYAL1 | ENSG00000114378.17 | 4537 | 1.037 | 0.7641 | Yes | ||
| 14 | IDS | ENSG00000010404.18 | 9593 | 0.530 | 0.6661 | No | ||
| 15 | ARSB | ENSG00000113273.17 | 18569 | 0.120 | 0.4622 | No | ||
| 16 | GALNS | ENSG00000141012.13 | 23898 | -0.165 | 0.3445 | No | ||
| 17 | HYAL2 | ENSG00000068001.14 | 24832 | -0.220 | 0.3309 | No | ||
| 18 | GNS | ENSG00000135677.11 | 28324 | -0.331 | 0.2619 | No | ||
| 19 | SPAM1 | ENSG00000106304.15 | 29845 | -0.394 | 0.2409 | No | ||
| 20 | HPSE | ENSG00000173083.15 | 35116 | -0.752 | 0.1459 | No | ||
| 21 | HYAL4 | ENSG00000106302.10 | 37871 | -1.095 | 0.1218 | No |