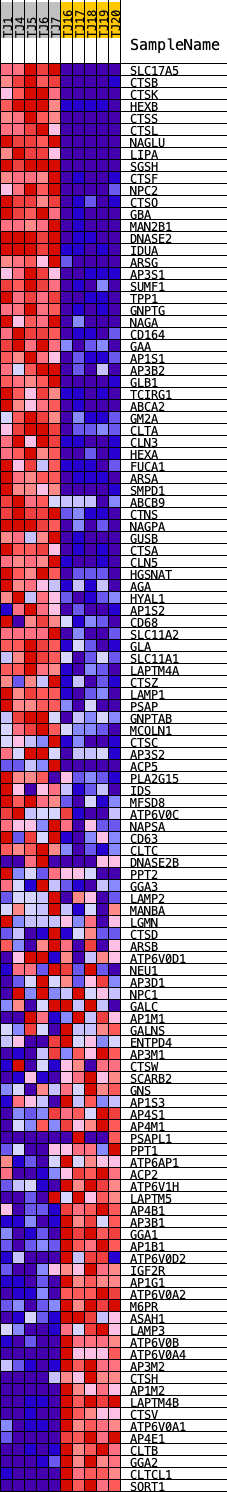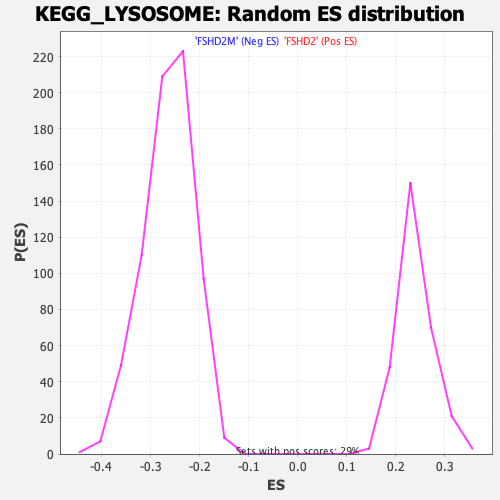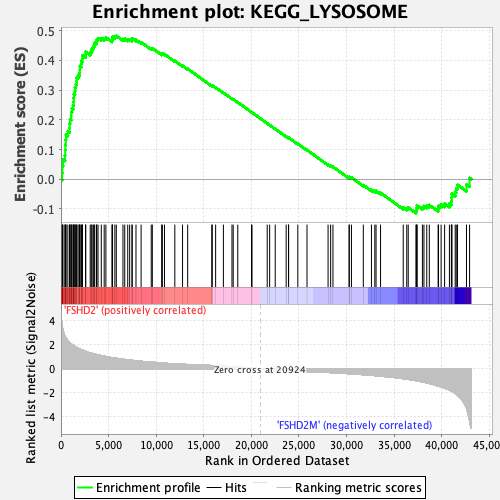
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_LYSOSOME |
| Enrichment Score (ES) | 0.48280397 |
| Normalized Enrichment Score (NES) | 2.0169766 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.004864547 |
| FWER p-Value | 0.061 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLC17A5 | ENSG00000119899.13 | 119 | 3.608 | 0.0220 | Yes | ||
| 2 | CTSB | ENSG00000164733.21 | 145 | 3.504 | 0.0454 | Yes | ||
| 3 | CTSK | ENSG00000143387.13 | 210 | 3.273 | 0.0664 | Yes | ||
| 4 | HEXB | ENSG00000049860.14 | 415 | 2.734 | 0.0804 | Yes | ||
| 5 | CTSS | ENSG00000163131.11 | 442 | 2.687 | 0.0982 | Yes | ||
| 6 | CTSL | ENSG00000135047.15 | 458 | 2.665 | 0.1162 | Yes | ||
| 7 | NAGLU | ENSG00000108784.10 | 504 | 2.596 | 0.1329 | Yes | ||
| 8 | LIPA | ENSG00000107798.18 | 528 | 2.571 | 0.1500 | Yes | ||
| 9 | SGSH | ENSG00000181523.13 | 719 | 2.352 | 0.1617 | Yes | ||
| 10 | CTSF | ENSG00000174080.11 | 899 | 2.175 | 0.1725 | Yes | ||
| 11 | NPC2 | ENSG00000119655.10 | 904 | 2.166 | 0.1873 | Yes | ||
| 12 | CTSO | ENSG00000256043.3 | 953 | 2.131 | 0.2008 | Yes | ||
| 13 | GBA | ENSG00000177628.16 | 1072 | 2.067 | 0.2122 | Yes | ||
| 14 | MAN2B1 | ENSG00000104774.13 | 1074 | 2.066 | 0.2263 | Yes | ||
| 15 | DNASE2 | ENSG00000105612.9 | 1147 | 2.018 | 0.2385 | Yes | ||
| 16 | IDUA | ENSG00000127415.13 | 1270 | 1.956 | 0.2491 | Yes | ||
| 17 | ARSG | ENSG00000141337.12 | 1326 | 1.924 | 0.2610 | Yes | ||
| 18 | AP3S1 | ENSG00000177879.16 | 1336 | 1.916 | 0.2739 | Yes | ||
| 19 | SUMF1 | ENSG00000144455.14 | 1354 | 1.904 | 0.2866 | Yes | ||
| 20 | TPP1 | ENSG00000166340.17 | 1461 | 1.860 | 0.2969 | Yes | ||
| 21 | GNPTG | ENSG00000090581.10 | 1478 | 1.853 | 0.3092 | Yes | ||
| 22 | NAGA | ENSG00000198951.12 | 1559 | 1.815 | 0.3198 | Yes | ||
| 23 | CD164 | ENSG00000135535.17 | 1647 | 1.782 | 0.3300 | Yes | ||
| 24 | GAA | ENSG00000171298.13 | 1658 | 1.772 | 0.3419 | Yes | ||
| 25 | AP1S1 | ENSG00000106367.15 | 1803 | 1.716 | 0.3504 | Yes | ||
| 26 | AP3B2 | ENSG00000103723.15 | 1945 | 1.652 | 0.3584 | Yes | ||
| 27 | GLB1 | ENSG00000170266.16 | 1972 | 1.642 | 0.3691 | Yes | ||
| 28 | TCIRG1 | ENSG00000110719.10 | 1974 | 1.641 | 0.3803 | Yes | ||
| 29 | ABCA2 | ENSG00000107331.17 | 2112 | 1.595 | 0.3881 | Yes | ||
| 30 | GM2A | ENSG00000196743.9 | 2138 | 1.587 | 0.3984 | Yes | ||
| 31 | CLTA | ENSG00000122705.17 | 2253 | 1.549 | 0.4063 | Yes | ||
| 32 | CLN3 | ENSG00000188603.19 | 2258 | 1.546 | 0.4169 | Yes | ||
| 33 | HEXA | ENSG00000213614.9 | 2587 | 1.446 | 0.4191 | Yes | ||
| 34 | FUCA1 | ENSG00000179163.11 | 2594 | 1.443 | 0.4289 | Yes | ||
| 35 | ARSA | ENSG00000100299.18 | 3068 | 1.317 | 0.4269 | Yes | ||
| 36 | SMPD1 | ENSG00000166311.10 | 3178 | 1.292 | 0.4333 | Yes | ||
| 37 | ABCB9 | ENSG00000150967.18 | 3261 | 1.275 | 0.4401 | Yes | ||
| 38 | CTNS | ENSG00000040531.14 | 3394 | 1.246 | 0.4456 | Yes | ||
| 39 | NAGPA | ENSG00000103174.13 | 3475 | 1.229 | 0.4522 | Yes | ||
| 40 | GUSB | ENSG00000169919.17 | 3543 | 1.217 | 0.4589 | Yes | ||
| 41 | CTSA | ENSG00000064601.18 | 3731 | 1.177 | 0.4627 | Yes | ||
| 42 | CLN5 | ENSG00000102805.15 | 3756 | 1.173 | 0.4702 | Yes | ||
| 43 | HGSNAT | ENSG00000165102.15 | 3889 | 1.147 | 0.4750 | Yes | ||
| 44 | AGA | ENSG00000038002.9 | 4232 | 1.087 | 0.4745 | Yes | ||
| 45 | HYAL1 | ENSG00000114378.17 | 4537 | 1.037 | 0.4745 | Yes | ||
| 46 | AP1S2 | ENSG00000182287.15 | 4715 | 1.011 | 0.4773 | Yes | ||
| 47 | CD68 | ENSG00000129226.14 | 5359 | 0.919 | 0.4687 | Yes | ||
| 48 | SLC11A2 | ENSG00000110911.16 | 5394 | 0.914 | 0.4742 | Yes | ||
| 49 | GLA | ENSG00000102393.10 | 5414 | 0.912 | 0.4800 | Yes | ||
| 50 | SLC11A1 | ENSG00000018280.17 | 5657 | 0.881 | 0.4804 | Yes | ||
| 51 | LAPTM4A | ENSG00000068697.7 | 5809 | 0.863 | 0.4828 | Yes | ||
| 52 | CTSZ | ENSG00000101160.14 | 6508 | 0.788 | 0.4720 | No | ||
| 53 | LAMP1 | ENSG00000185896.11 | 6695 | 0.768 | 0.4729 | No | ||
| 54 | PSAP | ENSG00000197746.14 | 7009 | 0.737 | 0.4707 | No | ||
| 55 | GNPTAB | ENSG00000111670.16 | 7212 | 0.718 | 0.4709 | No | ||
| 56 | MCOLN1 | ENSG00000090674.16 | 7458 | 0.694 | 0.4700 | No | ||
| 57 | CTSC | ENSG00000109861.16 | 7476 | 0.693 | 0.4743 | No | ||
| 58 | AP3S2 | ENSG00000157823.17 | 7877 | 0.658 | 0.4695 | No | ||
| 59 | ACP5 | ENSG00000102575.13 | 8414 | 0.611 | 0.4613 | No | ||
| 60 | PLA2G15 | ENSG00000103066.13 | 9480 | 0.540 | 0.4402 | No | ||
| 61 | IDS | ENSG00000010404.18 | 9593 | 0.530 | 0.4412 | No | ||
| 62 | MFSD8 | ENSG00000164073.10 | 10574 | 0.473 | 0.4217 | No | ||
| 63 | ATP6V0C | ENSG00000185883.12 | 10646 | 0.469 | 0.4233 | No | ||
| 64 | NAPSA | ENSG00000131400.8 | 10869 | 0.454 | 0.4212 | No | ||
| 65 | CD63 | ENSG00000135404.11 | 11956 | 0.390 | 0.3986 | No | ||
| 66 | CLTC | ENSG00000141367.12 | 12768 | 0.357 | 0.3822 | No | ||
| 67 | DNASE2B | ENSG00000137976.7 | 13308 | 0.339 | 0.3720 | No | ||
| 68 | PPT2 | ENSG00000221988.13 | 15840 | 0.258 | 0.3149 | No | ||
| 69 | GGA3 | ENSG00000125447.17 | 15887 | 0.254 | 0.3156 | No | ||
| 70 | LAMP2 | ENSG00000005893.15 | 16244 | 0.233 | 0.3089 | No | ||
| 71 | MANBA | ENSG00000109323.11 | 17058 | 0.195 | 0.2913 | No | ||
| 72 | LGMN | ENSG00000100600.15 | 17960 | 0.148 | 0.2714 | No | ||
| 73 | CTSD | ENSG00000117984.14 | 18089 | 0.141 | 0.2694 | No | ||
| 74 | ARSB | ENSG00000113273.17 | 18569 | 0.120 | 0.2591 | No | ||
| 75 | ATP6V0D1 | ENSG00000159720.12 | 20013 | 0.048 | 0.2259 | No | ||
| 76 | NEU1 | ENSG00000204386.11 | 20064 | 0.045 | 0.2250 | No | ||
| 77 | AP3D1 | ENSG00000065000.18 | 21662 | -0.040 | 0.1881 | No | ||
| 78 | NPC1 | ENSG00000141458.13 | 21917 | -0.052 | 0.1826 | No | ||
| 79 | GALC | ENSG00000054983.17 | 22503 | -0.084 | 0.1696 | No | ||
| 80 | AP1M1 | ENSG00000072958.8 | 23653 | -0.149 | 0.1439 | No | ||
| 81 | GALNS | ENSG00000141012.13 | 23898 | -0.165 | 0.1393 | No | ||
| 82 | ENTPD4 | ENSG00000197217.13 | 23911 | -0.166 | 0.1402 | No | ||
| 83 | AP3M1 | ENSG00000185009.12 | 24877 | -0.223 | 0.1193 | No | ||
| 84 | CTSW | ENSG00000172543.8 | 25841 | -0.250 | 0.0986 | No | ||
| 85 | SCARB2 | ENSG00000138760.10 | 28055 | -0.314 | 0.0493 | No | ||
| 86 | GNS | ENSG00000135677.11 | 28324 | -0.331 | 0.0453 | No | ||
| 87 | AP1S3 | ENSG00000152056.17 | 28567 | -0.340 | 0.0420 | No | ||
| 88 | AP4S1 | ENSG00000100478.15 | 30265 | -0.426 | 0.0055 | No | ||
| 89 | AP4M1 | ENSG00000221838.10 | 30280 | -0.427 | 0.0081 | No | ||
| 90 | PSAPL1 | ENSG00000178597.7 | 30511 | -0.436 | 0.0057 | No | ||
| 91 | PPT1 | ENSG00000131238.17 | 31757 | -0.501 | -0.0198 | No | ||
| 92 | ATP6AP1 | ENSG00000071553.17 | 32612 | -0.553 | -0.0359 | No | ||
| 93 | ACP2 | ENSG00000134575.12 | 32966 | -0.580 | -0.0401 | No | ||
| 94 | ATP6V1H | ENSG00000047249.18 | 33094 | -0.590 | -0.0390 | No | ||
| 95 | LAPTM5 | ENSG00000162511.8 | 33566 | -0.626 | -0.0457 | No | ||
| 96 | AP4B1 | ENSG00000134262.13 | 35950 | -0.838 | -0.0953 | No | ||
| 97 | AP3B1 | ENSG00000132842.13 | 36321 | -0.880 | -0.0979 | No | ||
| 98 | GGA1 | ENSG00000100083.19 | 36475 | -0.897 | -0.0953 | No | ||
| 99 | AP1B1 | ENSG00000100280.16 | 37284 | -1.004 | -0.1072 | No | ||
| 100 | ATP6V0D2 | ENSG00000147614.4 | 37357 | -1.014 | -0.1019 | No | ||
| 101 | IGF2R | ENSG00000197081.14 | 37384 | -1.018 | -0.0955 | No | ||
| 102 | AP1G1 | ENSG00000166747.12 | 37396 | -1.019 | -0.0888 | No | ||
| 103 | ATP6V0A2 | ENSG00000185344.14 | 37964 | -1.111 | -0.0944 | No | ||
| 104 | M6PR | ENSG00000003056.8 | 38125 | -1.138 | -0.0903 | No | ||
| 105 | ASAH1 | ENSG00000104763.19 | 38422 | -1.192 | -0.0890 | No | ||
| 106 | LAMP3 | ENSG00000078081.8 | 38690 | -1.241 | -0.0867 | No | ||
| 107 | ATP6V0B | ENSG00000117410.14 | 39615 | -1.446 | -0.0983 | No | ||
| 108 | ATP6V0A4 | ENSG00000105929.16 | 39655 | -1.455 | -0.0892 | No | ||
| 109 | AP3M2 | ENSG00000070718.12 | 39930 | -1.530 | -0.0851 | No | ||
| 110 | CTSH | ENSG00000103811.16 | 40296 | -1.626 | -0.0824 | No | ||
| 111 | AP1M2 | ENSG00000129354.11 | 40801 | -1.786 | -0.0819 | No | ||
| 112 | LAPTM4B | ENSG00000104341.17 | 41005 | -1.871 | -0.0738 | No | ||
| 113 | CTSV | ENSG00000136943.11 | 41043 | -1.887 | -0.0617 | No | ||
| 114 | ATP6V0A1 | ENSG00000033627.16 | 41061 | -1.892 | -0.0491 | No | ||
| 115 | AP4E1 | ENSG00000081014.11 | 41408 | -2.068 | -0.0430 | No | ||
| 116 | CLTB | ENSG00000175416.13 | 41519 | -2.140 | -0.0308 | No | ||
| 117 | GGA2 | ENSG00000103365.15 | 41653 | -2.222 | -0.0187 | No | ||
| 118 | CLTCL1 | ENSG00000070371.16 | 42597 | -3.327 | -0.0178 | No | ||
| 119 | SORT1 | ENSG00000134243.12 | 42922 | -4.361 | 0.0046 | No |