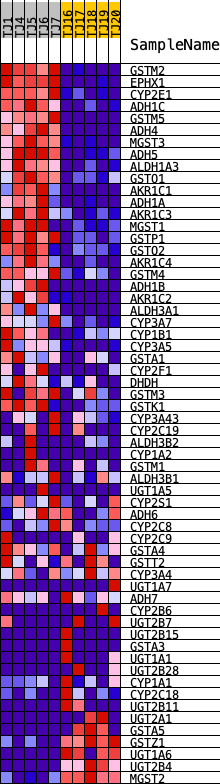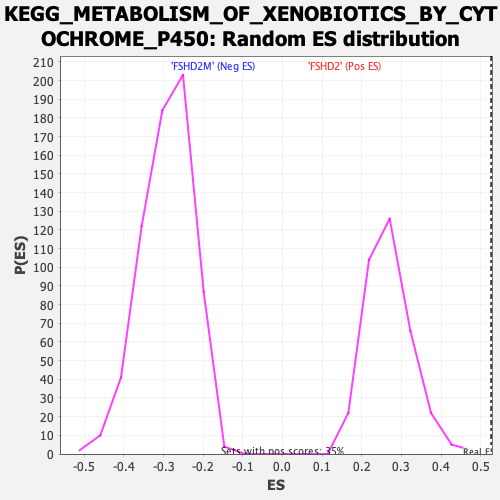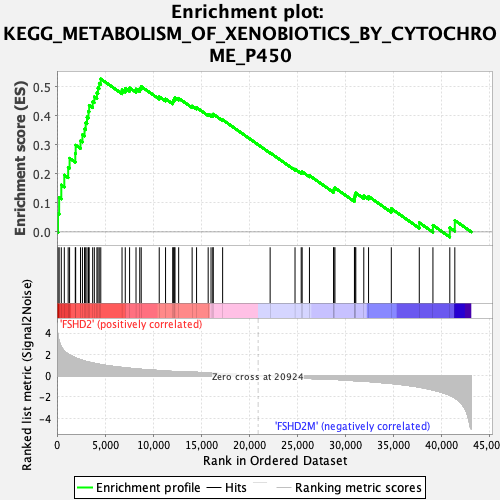
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | KEGG_METABOLISM_OF_XENOBIOTICS_BY_CYTOCHROME_P450 |
| Enrichment Score (ES) | 0.5265708 |
| Normalized Enrichment Score (NES) | 1.9559581 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.011280382 |
| FWER p-Value | 0.168 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GSTM2 | ENSG00000213366.13 | 105 | 3.658 | 0.0632 | Yes | ||
| 2 | EPHX1 | ENSG00000143819.12 | 223 | 3.237 | 0.1186 | Yes | ||
| 3 | CYP2E1 | ENSG00000130649.10 | 447 | 2.683 | 0.1616 | Yes | ||
| 4 | ADH1C | ENSG00000248144.6 | 760 | 2.300 | 0.1957 | Yes | ||
| 5 | GSTM5 | ENSG00000134201.12 | 1163 | 2.010 | 0.2224 | Yes | ||
| 6 | ADH4 | ENSG00000198099.8 | 1310 | 1.932 | 0.2537 | Yes | ||
| 7 | MGST3 | ENSG00000143198.13 | 1903 | 1.667 | 0.2699 | Yes | ||
| 8 | ADH5 | ENSG00000197894.11 | 1943 | 1.653 | 0.2986 | Yes | ||
| 9 | ALDH1A3 | ENSG00000184254.17 | 2445 | 1.491 | 0.3138 | Yes | ||
| 10 | GSTO1 | ENSG00000148834.13 | 2646 | 1.427 | 0.3347 | Yes | ||
| 11 | AKR1C1 | ENSG00000187134.14 | 2865 | 1.362 | 0.3541 | Yes | ||
| 12 | ADH1A | ENSG00000187758.8 | 2986 | 1.336 | 0.3753 | Yes | ||
| 13 | AKR1C3 | ENSG00000196139.14 | 3123 | 1.306 | 0.3956 | Yes | ||
| 14 | MGST1 | ENSG00000008394.13 | 3277 | 1.273 | 0.4149 | Yes | ||
| 15 | GSTP1 | ENSG00000084207.17 | 3349 | 1.255 | 0.4358 | Yes | ||
| 16 | GSTO2 | ENSG00000065621.15 | 3713 | 1.180 | 0.4486 | Yes | ||
| 17 | AKR1C4 | ENSG00000198610.11 | 3885 | 1.148 | 0.4652 | Yes | ||
| 18 | GSTM4 | ENSG00000168765.17 | 4156 | 1.103 | 0.4787 | Yes | ||
| 19 | ADH1B | ENSG00000196616.14 | 4264 | 1.081 | 0.4957 | Yes | ||
| 20 | AKR1C2 | ENSG00000151632.17 | 4394 | 1.062 | 0.5117 | Yes | ||
| 21 | ALDH3A1 | ENSG00000108602.18 | 4556 | 1.035 | 0.5266 | Yes | ||
| 22 | CYP3A7 | ENSG00000160870.14 | 6768 | 0.761 | 0.4889 | No | ||
| 23 | CYP1B1 | ENSG00000138061.12 | 7105 | 0.727 | 0.4941 | No | ||
| 24 | CYP3A5 | ENSG00000106258.15 | 7555 | 0.685 | 0.4960 | No | ||
| 25 | GSTA1 | ENSG00000243955.6 | 8224 | 0.627 | 0.4917 | No | ||
| 26 | CYP2F1 | ENSG00000197446.9 | 8613 | 0.598 | 0.4935 | No | ||
| 27 | DHDH | ENSG00000104808.8 | 8765 | 0.586 | 0.5005 | No | ||
| 28 | GSTM3 | ENSG00000134202.11 | 10640 | 0.469 | 0.4654 | No | ||
| 29 | GSTK1 | ENSG00000197448.14 | 11298 | 0.427 | 0.4578 | No | ||
| 30 | CYP3A43 | ENSG00000021461.16 | 12045 | 0.386 | 0.4474 | No | ||
| 31 | CYP2C19 | ENSG00000165841.11 | 12110 | 0.382 | 0.4528 | No | ||
| 32 | ALDH3B2 | ENSG00000132746.14 | 12210 | 0.377 | 0.4572 | No | ||
| 33 | CYP1A2 | ENSG00000140505.7 | 12281 | 0.375 | 0.4623 | No | ||
| 34 | GSTM1 | ENSG00000134184.13 | 12663 | 0.363 | 0.4600 | No | ||
| 35 | ALDH3B1 | ENSG00000006534.16 | 14064 | 0.314 | 0.4331 | No | ||
| 36 | UGT1A5 | ENSG00000240224.1 | 14526 | 0.308 | 0.4279 | No | ||
| 37 | CYP2S1 | ENSG00000167600.14 | 15734 | 0.264 | 0.4046 | No | ||
| 38 | ADH6 | ENSG00000172955.17 | 16031 | 0.247 | 0.4022 | No | ||
| 39 | CYP2C8 | ENSG00000138115.14 | 16222 | 0.235 | 0.4020 | No | ||
| 40 | CYP2C9 | ENSG00000138109.11 | 16255 | 0.232 | 0.4054 | No | ||
| 41 | GSTA4 | ENSG00000170899.11 | 17239 | 0.185 | 0.3859 | No | ||
| 42 | GSTT2 | ENSG00000099984.11 | 22183 | -0.067 | 0.2723 | No | ||
| 43 | CYP3A4 | ENSG00000160868.15 | 24773 | -0.217 | 0.2161 | No | ||
| 44 | UGT1A7 | ENSG00000244122.2 | 25416 | -0.223 | 0.2052 | No | ||
| 45 | ADH7 | ENSG00000196344.11 | 25515 | -0.228 | 0.2070 | No | ||
| 46 | CYP2B6 | ENSG00000197408.10 | 26275 | -0.263 | 0.1941 | No | ||
| 47 | UGT2B7 | ENSG00000171234.14 | 28785 | -0.351 | 0.1421 | No | ||
| 48 | UGT2B15 | ENSG00000196620.10 | 28824 | -0.351 | 0.1475 | No | ||
| 49 | GSTA3 | ENSG00000174156.15 | 28928 | -0.351 | 0.1515 | No | ||
| 50 | UGT1A1 | ENSG00000241635.7 | 30957 | -0.458 | 0.1126 | No | ||
| 51 | UGT2B28 | ENSG00000135226.18 | 30987 | -0.458 | 0.1201 | No | ||
| 52 | CYP1A1 | ENSG00000140465.14 | 31019 | -0.461 | 0.1277 | No | ||
| 53 | CYP2C18 | ENSG00000108242.13 | 31100 | -0.467 | 0.1342 | No | ||
| 54 | UGT2B11 | ENSG00000213759.10 | 31930 | -0.510 | 0.1241 | No | ||
| 55 | UGT2A1 | ENSG00000173610.12 | 32427 | -0.540 | 0.1223 | No | ||
| 56 | GSTA5 | ENSG00000182793.11 | 34795 | -0.724 | 0.0803 | No | ||
| 57 | GSTZ1 | ENSG00000100577.19 | 37706 | -1.068 | 0.0319 | No | ||
| 58 | UGT1A6 | ENSG00000167165.19 | 39127 | -1.336 | 0.0229 | No | ||
| 59 | UGT2B4 | ENSG00000156096.14 | 40883 | -1.816 | 0.0148 | No | ||
| 60 | MGST2 | ENSG00000085871.9 | 41405 | -2.068 | 0.0398 | No |