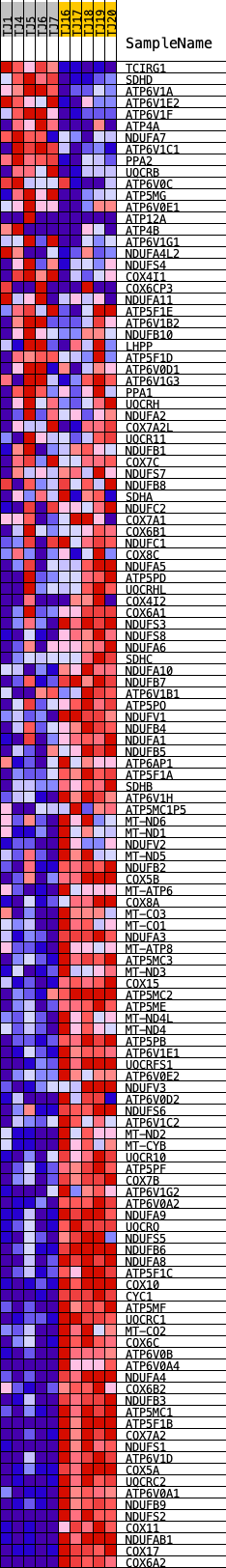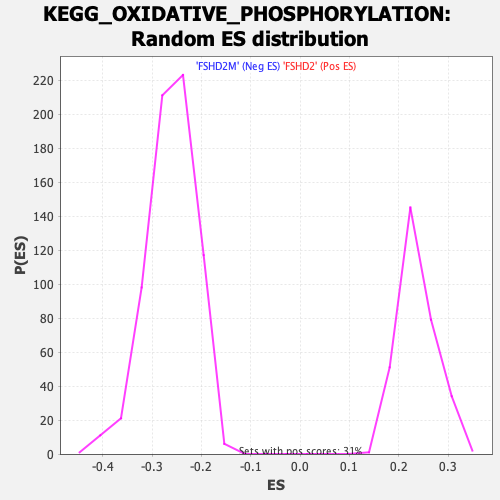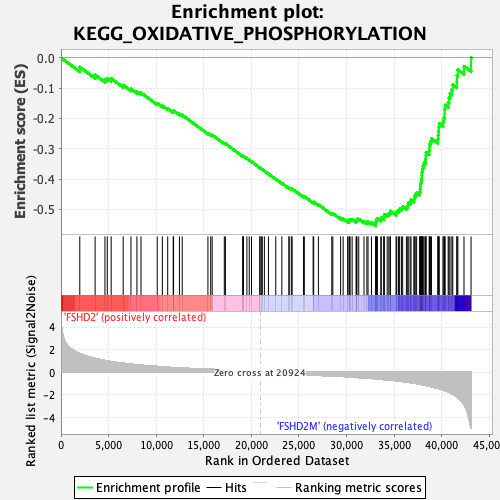
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | KEGG_OXIDATIVE_PHOSPHORYLATION |
| Enrichment Score (ES) | -0.55360335 |
| Normalized Enrichment Score (NES) | -2.1163445 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.9372736E-4 |
| FWER p-Value | 0.006 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TCIRG1 | ENSG00000110719.10 | 1974 | 1.641 | -0.0306 | No | ||
| 2 | SDHD | ENSG00000204370.12 | 3584 | 1.209 | -0.0567 | No | ||
| 3 | ATP6V1A | ENSG00000114573.10 | 4617 | 1.026 | -0.0712 | No | ||
| 4 | ATP6V1E2 | ENSG00000250565.7 | 4861 | 0.988 | -0.0676 | No | ||
| 5 | ATP6V1F | ENSG00000128524.5 | 5271 | 0.931 | -0.0684 | No | ||
| 6 | ATP4A | ENSG00000105675.9 | 6532 | 0.785 | -0.0904 | No | ||
| 7 | NDUFA7 | ENSG00000267855.6 | 7349 | 0.706 | -0.1028 | No | ||
| 8 | ATP6V1C1 | ENSG00000155097.12 | 7976 | 0.649 | -0.1113 | No | ||
| 9 | PPA2 | ENSG00000138777.20 | 8404 | 0.612 | -0.1155 | No | ||
| 10 | UQCRB | ENSG00000156467.9 | 10109 | 0.503 | -0.1504 | No | ||
| 11 | ATP6V0C | ENSG00000185883.12 | 10646 | 0.469 | -0.1585 | No | ||
| 12 | ATP5MG | ENSG00000167283.8 | 11209 | 0.432 | -0.1676 | No | ||
| 13 | ATP6V0E1 | ENSG00000113732.9 | 11789 | 0.397 | -0.1773 | No | ||
| 14 | ATP12A | ENSG00000075673.11 | 11826 | 0.396 | -0.1745 | No | ||
| 15 | ATP4B | ENSG00000186009.5 | 12441 | 0.365 | -0.1853 | No | ||
| 16 | ATP6V1G1 | ENSG00000136888.7 | 12733 | 0.359 | -0.1887 | No | ||
| 17 | NDUFA4L2 | ENSG00000185633.10 | 15433 | 0.279 | -0.2489 | No | ||
| 18 | NDUFS4 | ENSG00000164258.12 | 15726 | 0.264 | -0.2533 | No | ||
| 19 | COX4I1 | ENSG00000131143.9 | 15881 | 0.255 | -0.2545 | No | ||
| 20 | COX6CP3 | ENSG00000266251.1 | 17155 | 0.190 | -0.2823 | No | ||
| 21 | NDUFA11 | ENSG00000174886.13 | 17272 | 0.185 | -0.2833 | No | ||
| 22 | ATP5F1E | ENSG00000124172.10 | 19090 | 0.095 | -0.3246 | No | ||
| 23 | ATP6V1B2 | ENSG00000147416.11 | 19117 | 0.094 | -0.3244 | No | ||
| 24 | NDUFB10 | ENSG00000140990.15 | 19165 | 0.090 | -0.3246 | No | ||
| 25 | LHPP | ENSG00000107902.14 | 19528 | 0.070 | -0.3324 | No | ||
| 26 | ATP5F1D | ENSG00000099624.8 | 19774 | 0.057 | -0.3375 | No | ||
| 27 | ATP6V0D1 | ENSG00000159720.12 | 20013 | 0.048 | -0.3426 | No | ||
| 28 | ATP6V1G3 | ENSG00000151418.11 | 20888 | 0.003 | -0.3629 | No | ||
| 29 | PPA1 | ENSG00000180817.12 | 20900 | 0.002 | -0.3632 | No | ||
| 30 | UQCRH | ENSG00000173660.12 | 21041 | -0.006 | -0.3664 | No | ||
| 31 | NDUFA2 | ENSG00000131495.8 | 21146 | -0.013 | -0.3687 | No | ||
| 32 | COX7A2L | ENSG00000115944.15 | 21364 | -0.024 | -0.3735 | No | ||
| 33 | UQCR11 | ENSG00000127540.12 | 21789 | -0.046 | -0.3829 | No | ||
| 34 | NDUFB1 | ENSG00000183648.10 | 22559 | -0.088 | -0.4000 | No | ||
| 35 | COX7C | ENSG00000127184.13 | 23199 | -0.124 | -0.4137 | No | ||
| 36 | NDUFS7 | ENSG00000115286.20 | 23943 | -0.168 | -0.4294 | No | ||
| 37 | NDUFB8 | ENSG00000166136.16 | 23962 | -0.169 | -0.4283 | No | ||
| 38 | SDHA | ENSG00000073578.17 | 24192 | -0.182 | -0.4319 | No | ||
| 39 | NDUFC2 | ENSG00000151366.13 | 24279 | -0.187 | -0.4321 | No | ||
| 40 | COX7A1 | ENSG00000161281.11 | 25467 | -0.225 | -0.4577 | No | ||
| 41 | COX6B1 | ENSG00000126267.11 | 25561 | -0.231 | -0.4577 | No | ||
| 42 | NDUFC1 | ENSG00000109390.12 | 26484 | -0.265 | -0.4766 | No | ||
| 43 | COX8C | ENSG00000187581.3 | 26545 | -0.269 | -0.4755 | No | ||
| 44 | NDUFA5 | ENSG00000128609.15 | 27038 | -0.276 | -0.4844 | No | ||
| 45 | ATP5PD | ENSG00000167863.12 | 28443 | -0.334 | -0.5139 | No | ||
| 46 | UQCRHL | ENSG00000233954.6 | 28555 | -0.340 | -0.5133 | No | ||
| 47 | COX4I2 | ENSG00000131055.5 | 29371 | -0.368 | -0.5288 | No | ||
| 48 | COX6A1 | ENSG00000111775.3 | 29637 | -0.384 | -0.5314 | No | ||
| 49 | NDUFS3 | ENSG00000213619.10 | 30108 | -0.414 | -0.5385 | No | ||
| 50 | NDUFS8 | ENSG00000110717.13 | 30259 | -0.426 | -0.5380 | No | ||
| 51 | NDUFA6 | ENSG00000184983.11 | 30283 | -0.428 | -0.5346 | No | ||
| 52 | SDHC | ENSG00000143252.14 | 30372 | -0.429 | -0.5326 | No | ||
| 53 | NDUFA10 | ENSG00000130414.12 | 30583 | -0.440 | -0.5334 | No | ||
| 54 | NDUFB7 | ENSG00000099795.7 | 30995 | -0.459 | -0.5386 | No | ||
| 55 | ATP6V1B1 | ENSG00000116039.13 | 31042 | -0.463 | -0.5354 | No | ||
| 56 | ATP5PO | ENSG00000241837.7 | 31091 | -0.467 | -0.5321 | No | ||
| 57 | NDUFV1 | ENSG00000167792.13 | 31274 | -0.476 | -0.5319 | No | ||
| 58 | NDUFB4 | ENSG00000065518.8 | 31830 | -0.503 | -0.5401 | No | ||
| 59 | NDUFA1 | ENSG00000125356.7 | 32133 | -0.521 | -0.5423 | No | ||
| 60 | NDUFB5 | ENSG00000136521.13 | 32252 | -0.531 | -0.5401 | No | ||
| 61 | ATP6AP1 | ENSG00000071553.17 | 32612 | -0.553 | -0.5433 | No | ||
| 62 | ATP5F1A | ENSG00000152234.16 | 33057 | -0.587 | -0.5481 | Yes | ||
| 63 | SDHB | ENSG00000117118.9 | 33081 | -0.589 | -0.5432 | Yes | ||
| 64 | ATP6V1H | ENSG00000047249.18 | 33094 | -0.590 | -0.5379 | Yes | ||
| 65 | ATP5MC1P5 | ENSG00000227590.2 | 33130 | -0.593 | -0.5332 | Yes | ||
| 66 | MT-ND6 | ENSG00000198695.2 | 33237 | -0.601 | -0.5301 | Yes | ||
| 67 | MT-ND1 | ENSG00000198888.2 | 33602 | -0.630 | -0.5326 | Yes | ||
| 68 | NDUFV2 | ENSG00000178127.13 | 33641 | -0.633 | -0.5276 | Yes | ||
| 69 | MT-ND5 | ENSG00000198786.2 | 33909 | -0.654 | -0.5277 | Yes | ||
| 70 | NDUFB2 | ENSG00000090266.13 | 33932 | -0.656 | -0.5221 | Yes | ||
| 71 | COX5B | ENSG00000135940.7 | 33952 | -0.658 | -0.5164 | Yes | ||
| 72 | MT-ATP6 | ENSG00000198899.2 | 34264 | -0.680 | -0.5173 | Yes | ||
| 73 | COX8A | ENSG00000176340.4 | 34436 | -0.697 | -0.5148 | Yes | ||
| 74 | MT-CO3 | ENSG00000198938.2 | 34569 | -0.708 | -0.5112 | Yes | ||
| 75 | MT-CO1 | ENSG00000198804.2 | 34603 | -0.710 | -0.5054 | Yes | ||
| 76 | NDUFA3 | ENSG00000170906.16 | 35193 | -0.759 | -0.5120 | Yes | ||
| 77 | MT-ATP8 | ENSG00000228253.1 | 35268 | -0.765 | -0.5066 | Yes | ||
| 78 | ATP5MC3 | ENSG00000154518.9 | 35448 | -0.783 | -0.5034 | Yes | ||
| 79 | MT-ND3 | ENSG00000198840.2 | 35569 | -0.794 | -0.4988 | Yes | ||
| 80 | COX15 | ENSG00000014919.12 | 35759 | -0.814 | -0.4956 | Yes | ||
| 81 | ATP5MC2 | ENSG00000135390.20 | 35892 | -0.830 | -0.4909 | Yes | ||
| 82 | ATP5ME | ENSG00000169020.10 | 36282 | -0.876 | -0.4918 | Yes | ||
| 83 | MT-ND4L | ENSG00000212907.2 | 36434 | -0.893 | -0.4869 | Yes | ||
| 84 | MT-ND4 | ENSG00000198886.2 | 36465 | -0.896 | -0.4793 | Yes | ||
| 85 | ATP5PB | ENSG00000116459.11 | 36670 | -0.924 | -0.4754 | Yes | ||
| 86 | ATP6V1E1 | ENSG00000131100.13 | 36765 | -0.936 | -0.4688 | Yes | ||
| 87 | UQCRFS1 | ENSG00000169021.6 | 37078 | -0.978 | -0.4669 | Yes | ||
| 88 | ATP6V0E2 | ENSG00000171130.18 | 37106 | -0.981 | -0.4584 | Yes | ||
| 89 | NDUFV3 | ENSG00000160194.18 | 37212 | -0.994 | -0.4516 | Yes | ||
| 90 | ATP6V0D2 | ENSG00000147614.4 | 37357 | -1.014 | -0.4455 | Yes | ||
| 91 | NDUFS6 | ENSG00000145494.11 | 37676 | -1.062 | -0.4429 | Yes | ||
| 92 | ATP6V1C2 | ENSG00000143882.12 | 37716 | -1.070 | -0.4338 | Yes | ||
| 93 | MT-ND2 | ENSG00000198763.3 | 37747 | -1.076 | -0.4245 | Yes | ||
| 94 | MT-CYB | ENSG00000198727.2 | 37765 | -1.078 | -0.4148 | Yes | ||
| 95 | UQCR10 | ENSG00000184076.13 | 37826 | -1.087 | -0.4061 | Yes | ||
| 96 | ATP5PF | ENSG00000154723.12 | 37895 | -1.100 | -0.3974 | Yes | ||
| 97 | COX7B | ENSG00000131174.7 | 37903 | -1.101 | -0.3873 | Yes | ||
| 98 | ATP6V1G2 | ENSG00000213760.10 | 37912 | -1.102 | -0.3771 | Yes | ||
| 99 | ATP6V0A2 | ENSG00000185344.14 | 37964 | -1.111 | -0.3680 | Yes | ||
| 100 | NDUFA9 | ENSG00000139180.11 | 38015 | -1.119 | -0.3587 | Yes | ||
| 101 | UQCRQ | ENSG00000164405.11 | 38111 | -1.136 | -0.3503 | Yes | ||
| 102 | NDUFS5 | ENSG00000168653.11 | 38239 | -1.160 | -0.3424 | Yes | ||
| 103 | NDUFB6 | ENSG00000165264.11 | 38305 | -1.173 | -0.3329 | Yes | ||
| 104 | NDUFA8 | ENSG00000119421.7 | 38322 | -1.176 | -0.3223 | Yes | ||
| 105 | ATP5F1C | ENSG00000165629.20 | 38343 | -1.178 | -0.3118 | Yes | ||
| 106 | COX10 | ENSG00000006695.12 | 38672 | -1.238 | -0.3079 | Yes | ||
| 107 | CYC1 | ENSG00000179091.5 | 38710 | -1.245 | -0.2971 | Yes | ||
| 108 | ATP5MF | ENSG00000241468.7 | 38712 | -1.245 | -0.2855 | Yes | ||
| 109 | UQCRC1 | ENSG00000010256.11 | 38824 | -1.273 | -0.2762 | Yes | ||
| 110 | MT-CO2 | ENSG00000198712.1 | 38915 | -1.292 | -0.2662 | Yes | ||
| 111 | COX6C | ENSG00000164919.11 | 39588 | -1.439 | -0.2684 | Yes | ||
| 112 | ATP6V0B | ENSG00000117410.14 | 39615 | -1.446 | -0.2555 | Yes | ||
| 113 | ATP6V0A4 | ENSG00000105929.16 | 39655 | -1.455 | -0.2428 | Yes | ||
| 114 | NDUFA4 | ENSG00000189043.10 | 39683 | -1.463 | -0.2298 | Yes | ||
| 115 | COX6B2 | ENSG00000160471.13 | 39727 | -1.474 | -0.2170 | Yes | ||
| 116 | NDUFB3 | ENSG00000119013.9 | 40121 | -1.579 | -0.2114 | Yes | ||
| 117 | ATP5MC1 | ENSG00000159199.14 | 40212 | -1.603 | -0.1986 | Yes | ||
| 118 | ATP5F1B | ENSG00000110955.9 | 40290 | -1.624 | -0.1852 | Yes | ||
| 119 | COX7A2 | ENSG00000112695.11 | 40291 | -1.624 | -0.1700 | Yes | ||
| 120 | NDUFS1 | ENSG00000023228.14 | 40350 | -1.642 | -0.1560 | Yes | ||
| 121 | ATP6V1D | ENSG00000100554.12 | 40671 | -1.741 | -0.1472 | Yes | ||
| 122 | COX5A | ENSG00000178741.12 | 40756 | -1.769 | -0.1327 | Yes | ||
| 123 | UQCRC2 | ENSG00000140740.11 | 40862 | -1.806 | -0.1182 | Yes | ||
| 124 | ATP6V0A1 | ENSG00000033627.16 | 41061 | -1.892 | -0.1052 | Yes | ||
| 125 | NDUFB9 | ENSG00000147684.9 | 41155 | -1.935 | -0.0893 | Yes | ||
| 126 | NDUFS2 | ENSG00000158864.12 | 41582 | -2.182 | -0.0788 | Yes | ||
| 127 | COX11 | ENSG00000166260.11 | 41593 | -2.189 | -0.0586 | Yes | ||
| 128 | NDUFAB1 | ENSG00000004779.10 | 41690 | -2.242 | -0.0399 | Yes | ||
| 129 | COX17 | ENSG00000138495.6 | 42332 | -2.886 | -0.0278 | Yes | ||
| 130 | COX6A2 | ENSG00000156885.6 | 43073 | -4.938 | 0.0011 | Yes |