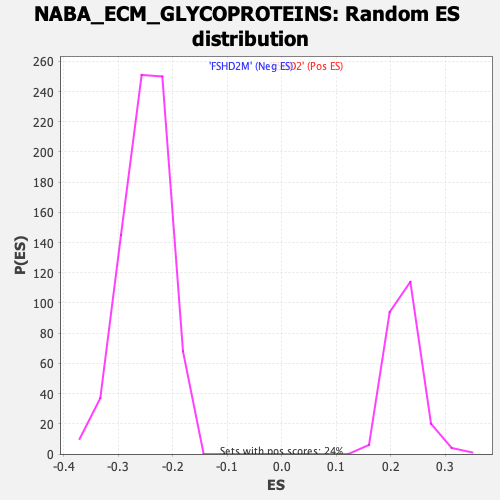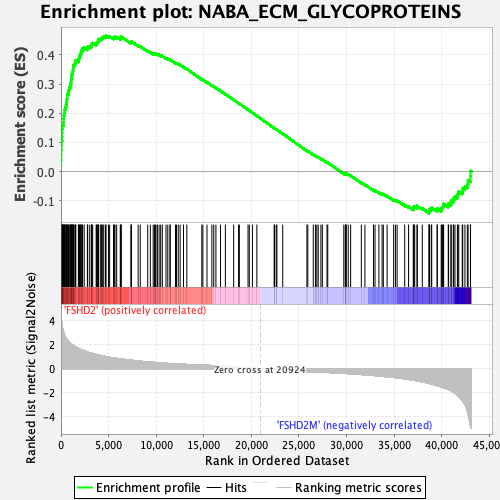
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | NABA_ECM_GLYCOPROTEINS |
| Enrichment Score (ES) | 0.46542585 |
| Normalized Enrichment Score (NES) | 2.0771675 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0042593204 |
| FWER p-Value | 0.027 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SLIT3 | ENSG00000184347.14 | 0 | 4.923 | 0.0195 | Yes | ||
| 2 | MFAP4 | ENSG00000166482.11 | 2 | 4.870 | 0.0387 | Yes | ||
| 3 | FBLN5 | ENSG00000140092.14 | 11 | 4.579 | 0.0566 | Yes | ||
| 4 | BMPER | ENSG00000164619.10 | 17 | 4.433 | 0.0740 | Yes | ||
| 5 | PCOLCE | ENSG00000106333.13 | 33 | 4.267 | 0.0906 | Yes | ||
| 6 | FBLN1 | ENSG00000077942.19 | 63 | 3.993 | 0.1057 | Yes | ||
| 7 | VWCE | ENSG00000167992.13 | 118 | 3.609 | 0.1187 | Yes | ||
| 8 | CTHRC1 | ENSG00000164932.13 | 123 | 3.593 | 0.1328 | Yes | ||
| 9 | LAMA4 | ENSG00000112769.20 | 127 | 3.582 | 0.1469 | Yes | ||
| 10 | IGFBP4 | ENSG00000141753.7 | 151 | 3.478 | 0.1601 | Yes | ||
| 11 | TNXB | ENSG00000168477.19 | 279 | 3.050 | 0.1692 | Yes | ||
| 12 | DPT | ENSG00000143196.5 | 284 | 3.034 | 0.1811 | Yes | ||
| 13 | PCOLCE2 | ENSG00000163710.9 | 294 | 3.004 | 0.1928 | Yes | ||
| 14 | LAMB1 | ENSG00000091136.14 | 367 | 2.820 | 0.2023 | Yes | ||
| 15 | HMCN1 | ENSG00000143341.12 | 385 | 2.781 | 0.2129 | Yes | ||
| 16 | SLIT2 | ENSG00000145147.20 | 459 | 2.664 | 0.2217 | Yes | ||
| 17 | NID1 | ENSG00000116962.15 | 541 | 2.561 | 0.2300 | Yes | ||
| 18 | MATN2 | ENSG00000132561.14 | 579 | 2.497 | 0.2390 | Yes | ||
| 19 | NTN1 | ENSG00000065320.9 | 605 | 2.467 | 0.2481 | Yes | ||
| 20 | AEBP1 | ENSG00000106624.11 | 659 | 2.404 | 0.2564 | Yes | ||
| 21 | SRPX | ENSG00000101955.15 | 660 | 2.402 | 0.2659 | Yes | ||
| 22 | VIT | ENSG00000205221.12 | 783 | 2.278 | 0.2721 | Yes | ||
| 23 | EMILIN2 | ENSG00000132205.11 | 806 | 2.257 | 0.2805 | Yes | ||
| 24 | EMILIN1 | ENSG00000138080.14 | 864 | 2.201 | 0.2879 | Yes | ||
| 25 | FBLN2 | ENSG00000163520.14 | 977 | 2.116 | 0.2936 | Yes | ||
| 26 | SVEP1 | ENSG00000165124.18 | 984 | 2.111 | 0.3018 | Yes | ||
| 27 | RSPO1 | ENSG00000169218.14 | 1063 | 2.074 | 0.3082 | Yes | ||
| 28 | CCN3 | ENSG00000136999.5 | 1071 | 2.068 | 0.3162 | Yes | ||
| 29 | ECM1 | ENSG00000143369.15 | 1104 | 2.046 | 0.3236 | Yes | ||
| 30 | SNED1 | ENSG00000162804.14 | 1129 | 2.032 | 0.3311 | Yes | ||
| 31 | SPON2 | ENSG00000159674.12 | 1174 | 2.003 | 0.3380 | Yes | ||
| 32 | IGFBP6 | ENSG00000167779.9 | 1241 | 1.971 | 0.3442 | Yes | ||
| 33 | LAMB4 | ENSG00000091128.13 | 1267 | 1.956 | 0.3514 | Yes | ||
| 34 | MFAP2 | ENSG00000117122.14 | 1290 | 1.945 | 0.3586 | Yes | ||
| 35 | NTNG1 | ENSG00000162631.18 | 1308 | 1.933 | 0.3658 | Yes | ||
| 36 | THBS2 | ENSG00000186340.15 | 1447 | 1.867 | 0.3700 | Yes | ||
| 37 | LTBP4 | ENSG00000090006.17 | 1533 | 1.830 | 0.3752 | Yes | ||
| 38 | MATN1 | ENSG00000162510.6 | 1553 | 1.817 | 0.3820 | Yes | ||
| 39 | IGFBP3 | ENSG00000146674.15 | 1828 | 1.702 | 0.3823 | Yes | ||
| 40 | NDNF | ENSG00000173376.14 | 1863 | 1.685 | 0.3882 | Yes | ||
| 41 | TSKU | ENSG00000182704.8 | 1939 | 1.654 | 0.3930 | Yes | ||
| 42 | IGFBP5 | ENSG00000115461.5 | 1987 | 1.637 | 0.3984 | Yes | ||
| 43 | LAMA1 | ENSG00000101680.15 | 2061 | 1.609 | 0.4030 | Yes | ||
| 44 | SPON1 | ENSG00000262655.4 | 2107 | 1.596 | 0.4083 | Yes | ||
| 45 | RSPO3 | ENSG00000146374.14 | 2124 | 1.591 | 0.4142 | Yes | ||
| 46 | CCN5 | ENSG00000064205.10 | 2260 | 1.545 | 0.4172 | Yes | ||
| 47 | GAS6 | ENSG00000183087.15 | 2269 | 1.542 | 0.4231 | Yes | ||
| 48 | MXRA5 | ENSG00000101825.8 | 2436 | 1.494 | 0.4251 | Yes | ||
| 49 | IGFBP2 | ENSG00000115457.10 | 2774 | 1.385 | 0.4228 | Yes | ||
| 50 | CCN4 | ENSG00000104415.14 | 2781 | 1.383 | 0.4281 | Yes | ||
| 51 | EFEMP2 | ENSG00000172638.13 | 2973 | 1.339 | 0.4289 | Yes | ||
| 52 | ECM2 | ENSG00000106823.12 | 3170 | 1.296 | 0.4295 | Yes | ||
| 53 | LTBP1 | ENSG00000049323.16 | 3213 | 1.284 | 0.4336 | Yes | ||
| 54 | NTN3 | ENSG00000162068.2 | 3239 | 1.278 | 0.4381 | Yes | ||
| 55 | MATN4 | ENSG00000124159.15 | 3329 | 1.259 | 0.4410 | Yes | ||
| 56 | VWF | ENSG00000110799.13 | 3693 | 1.186 | 0.4372 | Yes | ||
| 57 | MGP | ENSG00000111341.10 | 3725 | 1.178 | 0.4412 | Yes | ||
| 58 | GLDN | ENSG00000186417.14 | 3816 | 1.162 | 0.4437 | Yes | ||
| 59 | FBLN7 | ENSG00000144152.13 | 3897 | 1.146 | 0.4463 | Yes | ||
| 60 | VWA2 | ENSG00000165816.13 | 3906 | 1.145 | 0.4507 | Yes | ||
| 61 | SMOC2 | ENSG00000112562.18 | 3944 | 1.138 | 0.4543 | Yes | ||
| 62 | THBS1 | ENSG00000137801.11 | 4127 | 1.107 | 0.4544 | Yes | ||
| 63 | IGSF10 | ENSG00000152580.8 | 4244 | 1.084 | 0.4560 | Yes | ||
| 64 | KCP | ENSG00000135253.13 | 4307 | 1.075 | 0.4588 | Yes | ||
| 65 | TNC | ENSG00000041982.16 | 4396 | 1.062 | 0.4610 | Yes | ||
| 66 | NELL2 | ENSG00000184613.10 | 4492 | 1.045 | 0.4629 | Yes | ||
| 67 | NTN5 | ENSG00000142233.11 | 4679 | 1.017 | 0.4626 | Yes | ||
| 68 | LGI2 | ENSG00000153012.12 | 4730 | 1.009 | 0.4654 | Yes | ||
| 69 | VWA3A | ENSG00000175267.15 | 4988 | 0.970 | 0.4633 | No | ||
| 70 | VWA7 | ENSG00000204396.11 | 5110 | 0.953 | 0.4642 | No | ||
| 71 | LGI1 | ENSG00000108231.13 | 5517 | 0.899 | 0.4583 | No | ||
| 72 | EFEMP1 | ENSG00000115380.20 | 5601 | 0.888 | 0.4599 | No | ||
| 73 | MMRN2 | ENSG00000173269.14 | 5672 | 0.880 | 0.4617 | No | ||
| 74 | FBN1 | ENSG00000166147.13 | 5850 | 0.858 | 0.4610 | No | ||
| 75 | COMP | ENSG00000105664.11 | 6225 | 0.814 | 0.4555 | No | ||
| 76 | THBS3 | ENSG00000169231.13 | 6284 | 0.808 | 0.4574 | No | ||
| 77 | THSD4 | ENSG00000187720.14 | 6296 | 0.807 | 0.4603 | No | ||
| 78 | IGFALS | ENSG00000099769.5 | 6320 | 0.805 | 0.4629 | No | ||
| 79 | VTN | ENSG00000109072.14 | 7359 | 0.704 | 0.4416 | No | ||
| 80 | FNDC8 | ENSG00000073598.6 | 7366 | 0.704 | 0.4442 | No | ||
| 81 | THBS4 | ENSG00000113296.14 | 7404 | 0.700 | 0.4461 | No | ||
| 82 | LAMC3 | ENSG00000050555.18 | 8105 | 0.637 | 0.4323 | No | ||
| 83 | RELN | ENSG00000189056.14 | 8324 | 0.618 | 0.4297 | No | ||
| 84 | RSPO2 | ENSG00000147655.12 | 9109 | 0.562 | 0.4136 | No | ||
| 85 | EMILIN3 | ENSG00000183798.5 | 9386 | 0.545 | 0.4094 | No | ||
| 86 | FGA | ENSG00000171560.16 | 9683 | 0.524 | 0.4046 | No | ||
| 87 | MFGE8 | ENSG00000140545.15 | 9817 | 0.521 | 0.4035 | No | ||
| 88 | FN1 | ENSG00000115414.19 | 9913 | 0.516 | 0.4033 | No | ||
| 89 | LAMB2 | ENSG00000172037.14 | 9976 | 0.514 | 0.4039 | No | ||
| 90 | CRISPLD1 | ENSG00000121005.9 | 10130 | 0.502 | 0.4023 | No | ||
| 91 | PAPLN | ENSG00000100767.16 | 10307 | 0.491 | 0.4002 | No | ||
| 92 | EYS | ENSG00000188107.15 | 10470 | 0.480 | 0.3983 | No | ||
| 93 | CILP | ENSG00000138615.6 | 10643 | 0.469 | 0.3962 | No | ||
| 94 | CILP2 | ENSG00000160161.9 | 11030 | 0.442 | 0.3889 | No | ||
| 95 | FBN2 | ENSG00000138829.12 | 11219 | 0.432 | 0.3862 | No | ||
| 96 | INTS14 | ENSG00000138614.15 | 11455 | 0.416 | 0.3824 | No | ||
| 97 | CCN6 | ENSG00000112761.20 | 11461 | 0.416 | 0.3839 | No | ||
| 98 | ZP3 | ENSG00000188372.15 | 12028 | 0.387 | 0.3723 | No | ||
| 99 | FNDC7 | ENSG00000143107.10 | 12130 | 0.381 | 0.3714 | No | ||
| 100 | ZP1 | ENSG00000149506.11 | 12316 | 0.374 | 0.3686 | No | ||
| 101 | ADIPOQ | ENSG00000181092.10 | 12514 | 0.365 | 0.3655 | No | ||
| 102 | ZP2 | ENSG00000103310.10 | 12868 | 0.355 | 0.3587 | No | ||
| 103 | LTBP3 | ENSG00000168056.16 | 13216 | 0.345 | 0.3519 | No | ||
| 104 | FGL2 | ENSG00000127951.7 | 14791 | 0.292 | 0.3164 | No | ||
| 105 | DMP1 | ENSG00000152592.14 | 14902 | 0.287 | 0.3150 | No | ||
| 106 | SPARCL1 | ENSG00000152583.12 | 15335 | 0.283 | 0.3061 | No | ||
| 107 | SSPO | ENSG00000197558.12 | 15851 | 0.257 | 0.2951 | No | ||
| 108 | PXDNL | ENSG00000147485.13 | 16046 | 0.246 | 0.2915 | No | ||
| 109 | ANOS1 | ENSG00000011201.12 | 16273 | 0.231 | 0.2872 | No | ||
| 110 | AMELX | ENSG00000125363.14 | 16753 | 0.208 | 0.2768 | No | ||
| 111 | EMID1 | ENSG00000186998.16 | 17267 | 0.185 | 0.2656 | No | ||
| 112 | INTS6L | ENSG00000165359.15 | 18138 | 0.139 | 0.2459 | No | ||
| 113 | SLIT1 | ENSG00000187122.17 | 18654 | 0.116 | 0.2344 | No | ||
| 114 | RSPO4 | ENSG00000101282.9 | 18734 | 0.111 | 0.2330 | No | ||
| 115 | LGI3 | ENSG00000168481.9 | 19647 | 0.064 | 0.2120 | No | ||
| 116 | CRISPLD2 | ENSG00000103196.12 | 19779 | 0.057 | 0.2092 | No | ||
| 117 | LGI4 | ENSG00000153902.14 | 20108 | 0.042 | 0.2017 | No | ||
| 118 | SPP1 | ENSG00000118785.14 | 20575 | 0.018 | 0.1909 | No | ||
| 119 | CCN1 | ENSG00000142871.17 | 22390 | -0.079 | 0.1490 | No | ||
| 120 | LAMC1 | ENSG00000135862.6 | 22432 | -0.081 | 0.1483 | No | ||
| 121 | LAMA2 | ENSG00000196569.12 | 22603 | -0.091 | 0.1447 | No | ||
| 122 | FGG | ENSG00000171557.17 | 22680 | -0.095 | 0.1433 | No | ||
| 123 | FNDC1 | ENSG00000164694.17 | 23291 | -0.129 | 0.1296 | No | ||
| 124 | MEPE | ENSG00000152595.16 | 25833 | -0.250 | 0.0714 | No | ||
| 125 | AGRN | ENSG00000188157.15 | 25926 | -0.255 | 0.0703 | No | ||
| 126 | TSPEAR | ENSG00000175894.18 | 26499 | -0.266 | 0.0580 | No | ||
| 127 | MMRN1 | ENSG00000138722.10 | 26727 | -0.275 | 0.0538 | No | ||
| 128 | OTOL1 | ENSG00000182447.4 | 26871 | -0.275 | 0.0516 | No | ||
| 129 | CRIM1 | ENSG00000150938.10 | 27020 | -0.275 | 0.0492 | No | ||
| 130 | VWA5B2 | ENSG00000145198.14 | 27302 | -0.295 | 0.0438 | No | ||
| 131 | AMBN | ENSG00000178522.14 | 27454 | -0.297 | 0.0415 | No | ||
| 132 | VWA3B | ENSG00000168658.19 | 27943 | -0.305 | 0.0313 | No | ||
| 133 | DSPP | ENSG00000152591.15 | 28022 | -0.311 | 0.0308 | No | ||
| 134 | PXDN | ENSG00000130508.11 | 29709 | -0.387 | -0.0070 | No | ||
| 135 | TINAGL1 | ENSG00000142910.16 | 29866 | -0.395 | -0.0091 | No | ||
| 136 | TNR | ENSG00000116147.17 | 29931 | -0.401 | -0.0090 | No | ||
| 137 | VWA1 | ENSG00000179403.12 | 29936 | -0.402 | -0.0075 | No | ||
| 138 | SBSPON | ENSG00000164764.11 | 29941 | -0.402 | -0.0060 | No | ||
| 139 | NTNG2 | ENSG00000196358.11 | 30172 | -0.419 | -0.0097 | No | ||
| 140 | POMZP3 | ENSG00000146707.14 | 30425 | -0.433 | -0.0138 | No | ||
| 141 | COCH | ENSG00000100473.18 | 31549 | -0.493 | -0.0380 | No | ||
| 142 | LTBP2 | ENSG00000119681.12 | 31926 | -0.510 | -0.0448 | No | ||
| 143 | CCN2 | ENSG00000118523.6 | 32823 | -0.570 | -0.0634 | No | ||
| 144 | LAMA3 | ENSG00000053747.16 | 32988 | -0.582 | -0.0649 | No | ||
| 145 | POSTN | ENSG00000133110.15 | 33395 | -0.612 | -0.0719 | No | ||
| 146 | EDIL3 | ENSG00000164176.13 | 33729 | -0.640 | -0.0772 | No | ||
| 147 | VWA5B1 | ENSG00000158816.15 | 33860 | -0.650 | -0.0776 | No | ||
| 148 | FBN3 | ENSG00000142449.13 | 34259 | -0.679 | -0.0842 | No | ||
| 149 | VWDE | ENSG00000146530.14 | 34944 | -0.738 | -0.0972 | No | ||
| 150 | USH2A | ENSG00000042781.13 | 35129 | -0.754 | -0.0985 | No | ||
| 151 | ABI3BP | ENSG00000154175.17 | 35307 | -0.769 | -0.0996 | No | ||
| 152 | MFAP1 | ENSG00000140259.7 | 36107 | -0.857 | -0.1148 | No | ||
| 153 | CRELD2 | ENSG00000184164.14 | 36510 | -0.902 | -0.1206 | No | ||
| 154 | ELN | ENSG00000049540.17 | 37004 | -0.966 | -0.1283 | No | ||
| 155 | SPARC | ENSG00000113140.11 | 37052 | -0.974 | -0.1255 | No | ||
| 156 | NID2 | ENSG00000087303.18 | 37054 | -0.975 | -0.1217 | No | ||
| 157 | FRAS1 | ENSG00000138759.19 | 37156 | -0.987 | -0.1202 | No | ||
| 158 | TNFAIP6 | ENSG00000123610.5 | 37407 | -1.021 | -0.1219 | No | ||
| 159 | LAMC2 | ENSG00000058085.15 | 37414 | -1.022 | -0.1180 | No | ||
| 160 | SMOC1 | ENSG00000198732.10 | 37952 | -1.109 | -0.1262 | No | ||
| 161 | COLQ | ENSG00000206561.13 | 38666 | -1.236 | -0.1379 | No | ||
| 162 | MFAP5 | ENSG00000197614.11 | 38668 | -1.237 | -0.1330 | No | ||
| 163 | BGLAP | ENSG00000242252.2 | 38699 | -1.243 | -0.1288 | No | ||
| 164 | IBSP | ENSG00000029559.7 | 38893 | -1.288 | -0.1282 | No | ||
| 165 | TECTB | ENSG00000119913.6 | 38943 | -1.297 | -0.1242 | No | ||
| 166 | NELL1 | ENSG00000165973.19 | 39509 | -1.424 | -0.1318 | No | ||
| 167 | OIT3 | ENSG00000138315.13 | 39527 | -1.426 | -0.1265 | No | ||
| 168 | IGFBP7 | ENSG00000163453.11 | 39937 | -1.531 | -0.1300 | No | ||
| 169 | SRPX2 | ENSG00000102359.7 | 39993 | -1.546 | -0.1251 | No | ||
| 170 | MATN3 | ENSG00000132031.13 | 40083 | -1.566 | -0.1210 | No | ||
| 171 | LAMB3 | ENSG00000196878.15 | 40157 | -1.589 | -0.1164 | No | ||
| 172 | IGFBP1 | ENSG00000146678.10 | 40197 | -1.599 | -0.1110 | No | ||
| 173 | HMCN2 | ENSG00000148357.16 | 40680 | -1.743 | -0.1154 | No | ||
| 174 | TGFBI | ENSG00000120708.17 | 40715 | -1.752 | -0.1092 | No | ||
| 175 | NTN4 | ENSG00000074527.12 | 40972 | -1.850 | -0.1079 | No | ||
| 176 | CRELD1 | ENSG00000163703.17 | 40973 | -1.851 | -0.1005 | No | ||
| 177 | MFAP3 | ENSG00000037749.12 | 41206 | -1.961 | -0.0982 | No | ||
| 178 | TECTA | ENSG00000109927.10 | 41250 | -1.977 | -0.0914 | No | ||
| 179 | VWA5A | ENSG00000110002.16 | 41384 | -2.054 | -0.0864 | No | ||
| 180 | TNN | ENSG00000120332.16 | 41639 | -2.216 | -0.0835 | No | ||
| 181 | ZPLD1 | ENSG00000170044.8 | 41688 | -2.241 | -0.0758 | No | ||
| 182 | TINAG | ENSG00000137251.16 | 41783 | -2.314 | -0.0688 | No | ||
| 183 | LRG1 | ENSG00000171236.10 | 42146 | -2.634 | -0.0668 | No | ||
| 184 | DMBT1 | ENSG00000187908.19 | 42199 | -2.691 | -0.0574 | No | ||
| 185 | LAMA5 | ENSG00000130702.15 | 42431 | -3.030 | -0.0508 | No | ||
| 186 | NPNT | ENSG00000168743.12 | 42684 | -3.542 | -0.0426 | No | ||
| 187 | EGFLAM | ENSG00000164318.18 | 42765 | -3.770 | -0.0296 | No | ||
| 188 | OTOG | ENSG00000188162.11 | 43012 | -4.761 | -0.0165 | No | ||
| 189 | IGFBPL1 | ENSG00000137142.5 | 43020 | -4.797 | 0.0023 | No |