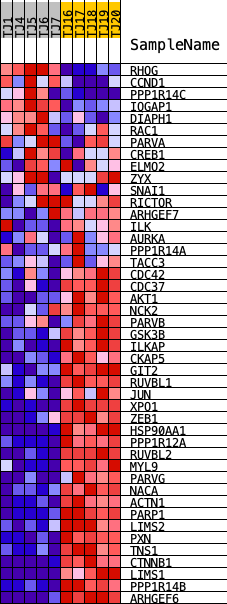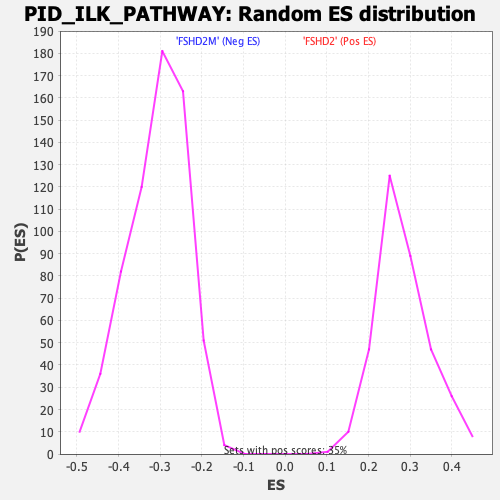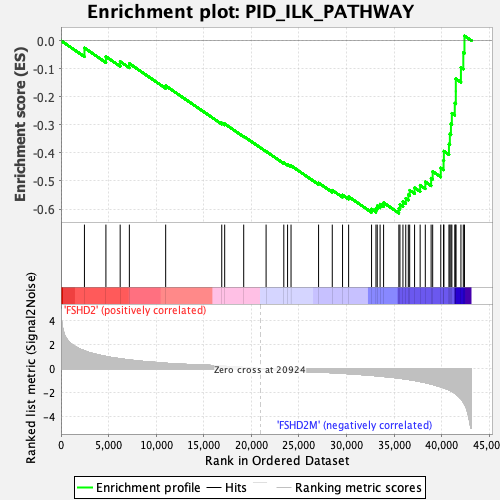
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | PID_ILK_PATHWAY |
| Enrichment Score (ES) | -0.61456674 |
| Normalized Enrichment Score (NES) | -1.9987456 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0011709168 |
| FWER p-Value | 0.041 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RHOG | ENSG00000177105.10 | 2462 | 1.485 | -0.0258 | No | ||
| 2 | CCND1 | ENSG00000110092.3 | 4717 | 1.011 | -0.0568 | No | ||
| 3 | PPP1R14C | ENSG00000198729.5 | 6216 | 0.815 | -0.0744 | No | ||
| 4 | IQGAP1 | ENSG00000140575.13 | 7181 | 0.721 | -0.0815 | No | ||
| 5 | DIAPH1 | ENSG00000131504.17 | 10995 | 0.445 | -0.1607 | No | ||
| 6 | RAC1 | ENSG00000136238.18 | 16886 | 0.203 | -0.2931 | No | ||
| 7 | PARVA | ENSG00000197702.14 | 17194 | 0.188 | -0.2963 | No | ||
| 8 | CREB1 | ENSG00000118260.15 | 19192 | 0.089 | -0.3408 | No | ||
| 9 | ELMO2 | ENSG00000062598.18 | 21544 | -0.034 | -0.3946 | No | ||
| 10 | ZYX | ENSG00000159840.16 | 23404 | -0.135 | -0.4349 | No | ||
| 11 | SNAI1 | ENSG00000124216.4 | 23800 | -0.158 | -0.4408 | No | ||
| 12 | RICTOR | ENSG00000164327.13 | 24164 | -0.180 | -0.4454 | No | ||
| 13 | ARHGEF7 | ENSG00000102606.18 | 27057 | -0.277 | -0.5067 | No | ||
| 14 | ILK | ENSG00000166333.13 | 28496 | -0.336 | -0.5330 | No | ||
| 15 | AURKA | ENSG00000087586.17 | 29575 | -0.379 | -0.5500 | No | ||
| 16 | PPP1R14A | ENSG00000167641.11 | 30214 | -0.422 | -0.5559 | No | ||
| 17 | TACC3 | ENSG00000013810.19 | 32622 | -0.553 | -0.6001 | Yes | ||
| 18 | CDC42 | ENSG00000070831.17 | 33086 | -0.589 | -0.5984 | Yes | ||
| 19 | CDC37 | ENSG00000105401.9 | 33219 | -0.600 | -0.5888 | Yes | ||
| 20 | AKT1 | ENSG00000142208.16 | 33520 | -0.623 | -0.5826 | Yes | ||
| 21 | NCK2 | ENSG00000071051.14 | 33887 | -0.653 | -0.5774 | Yes | ||
| 22 | PARVB | ENSG00000188677.14 | 35491 | -0.786 | -0.5980 | Yes | ||
| 23 | GSK3B | ENSG00000082701.15 | 35603 | -0.798 | -0.5837 | Yes | ||
| 24 | ILKAP | ENSG00000132323.9 | 35920 | -0.834 | -0.5734 | Yes | ||
| 25 | CKAP5 | ENSG00000175216.15 | 36214 | -0.869 | -0.5619 | Yes | ||
| 26 | GIT2 | ENSG00000139436.21 | 36484 | -0.899 | -0.5492 | Yes | ||
| 27 | RUVBL1 | ENSG00000175792.12 | 36627 | -0.919 | -0.5331 | Yes | ||
| 28 | JUN | ENSG00000177606.7 | 37141 | -0.985 | -0.5242 | Yes | ||
| 29 | XPO1 | ENSG00000082898.17 | 37727 | -1.072 | -0.5151 | Yes | ||
| 30 | ZEB1 | ENSG00000148516.21 | 38269 | -1.166 | -0.5031 | Yes | ||
| 31 | HSP90AA1 | ENSG00000080824.18 | 38879 | -1.285 | -0.4901 | Yes | ||
| 32 | PPP1R12A | ENSG00000058272.19 | 39043 | -1.320 | -0.4660 | Yes | ||
| 33 | RUVBL2 | ENSG00000183207.14 | 39894 | -1.521 | -0.4536 | Yes | ||
| 34 | MYL9 | ENSG00000101335.10 | 40187 | -1.596 | -0.4267 | Yes | ||
| 35 | PARVG | ENSG00000138964.17 | 40220 | -1.605 | -0.3936 | Yes | ||
| 36 | NACA | ENSG00000196531.11 | 40753 | -1.768 | -0.3686 | Yes | ||
| 37 | ACTN1 | ENSG00000072110.13 | 40848 | -1.801 | -0.3328 | Yes | ||
| 38 | PARP1 | ENSG00000143799.13 | 40962 | -1.848 | -0.2964 | Yes | ||
| 39 | LIMS2 | ENSG00000072163.19 | 41058 | -1.892 | -0.2586 | Yes | ||
| 40 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.2226 | Yes | ||
| 41 | TNS1 | ENSG00000079308.19 | 41474 | -2.116 | -0.1805 | Yes | ||
| 42 | CTNNB1 | ENSG00000168036.18 | 41475 | -2.117 | -0.1358 | Yes | ||
| 43 | LIMS1 | ENSG00000169756.16 | 42011 | -2.498 | -0.0954 | Yes | ||
| 44 | PPP1R14B | ENSG00000173457.11 | 42261 | -2.779 | -0.0426 | Yes | ||
| 45 | ARHGEF6 | ENSG00000129675.16 | 42382 | -2.958 | 0.0171 | Yes |