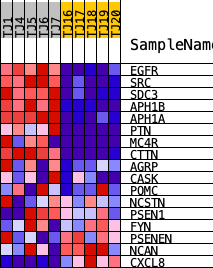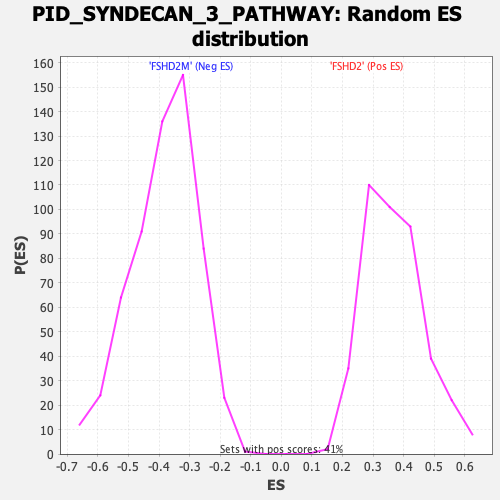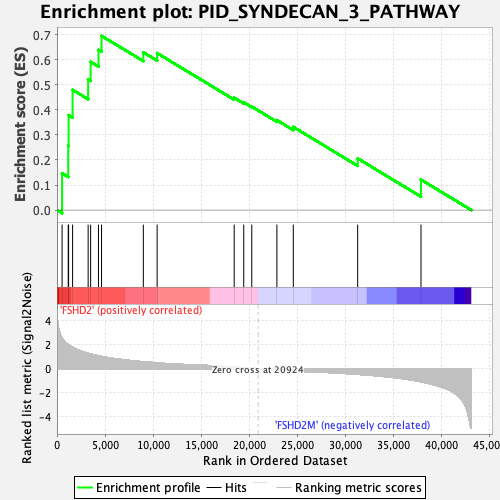
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | PID_SYNDECAN_3_PATHWAY |
| Enrichment Score (ES) | 0.6957495 |
| Normalized Enrichment Score (NES) | 1.8935255 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.019945312 |
| FWER p-Value | 0.333 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EGFR | ENSG00000146648.18 | 529 | 2.570 | 0.1470 | Yes | ||
| 2 | SRC | ENSG00000197122.11 | 1168 | 2.008 | 0.2567 | Yes | ||
| 3 | SDC3 | ENSG00000162512.16 | 1208 | 1.985 | 0.3789 | Yes | ||
| 4 | APH1B | ENSG00000138613.14 | 1623 | 1.791 | 0.4803 | Yes | ||
| 5 | APH1A | ENSG00000117362.13 | 3241 | 1.278 | 0.5221 | Yes | ||
| 6 | PTN | ENSG00000105894.12 | 3501 | 1.225 | 0.5920 | Yes | ||
| 7 | MC4R | ENSG00000166603.5 | 4316 | 1.074 | 0.6397 | Yes | ||
| 8 | CTTN | ENSG00000085733.16 | 4636 | 1.024 | 0.6957 | Yes | ||
| 9 | AGRP | ENSG00000159723.4 | 8985 | 0.569 | 0.6301 | No | ||
| 10 | CASK | ENSG00000147044.21 | 10423 | 0.483 | 0.6268 | No | ||
| 11 | POMC | ENSG00000115138.11 | 18442 | 0.128 | 0.4487 | No | ||
| 12 | NCSTN | ENSG00000162736.17 | 19436 | 0.074 | 0.4303 | No | ||
| 13 | PSEN1 | ENSG00000080815.19 | 20272 | 0.034 | 0.4130 | No | ||
| 14 | FYN | ENSG00000010810.17 | 22887 | -0.107 | 0.3590 | No | ||
| 15 | PSENEN | ENSG00000205155.8 | 24593 | -0.204 | 0.3320 | No | ||
| 16 | NCAN | ENSG00000130287.14 | 31294 | -0.478 | 0.2062 | No | ||
| 17 | CXCL8 | ENSG00000169429.11 | 37884 | -1.098 | 0.1215 | No |