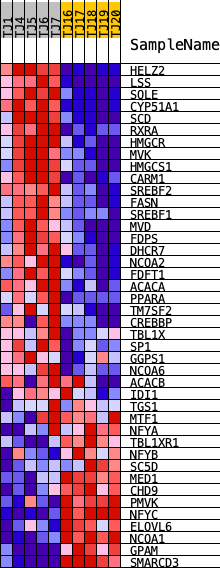
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_ACTIVATION_OF_GENE_EXPRESSION_BY_SREBF_SREBP |
| Enrichment Score (ES) | 0.57437086 |
| Normalized Enrichment Score (NES) | 1.9876584 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0075565595 |
| FWER p-Value | 0.109 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HELZ2 | ENSG00000130589.16 | 462 | 2.659 | 0.0446 | Yes | ||
| 2 | LSS | ENSG00000160285.15 | 553 | 2.538 | 0.0954 | Yes | ||
| 3 | SQLE | ENSG00000104549.12 | 823 | 2.241 | 0.1358 | Yes | ||
| 4 | CYP51A1 | ENSG00000001630.17 | 829 | 2.232 | 0.1822 | Yes | ||
| 5 | SCD | ENSG00000099194.6 | 1288 | 1.946 | 0.2121 | Yes | ||
| 6 | RXRA | ENSG00000186350.12 | 1332 | 1.918 | 0.2510 | Yes | ||
| 7 | HMGCR | ENSG00000113161.16 | 1456 | 1.862 | 0.2869 | Yes | ||
| 8 | MVK | ENSG00000110921.14 | 1775 | 1.727 | 0.3155 | Yes | ||
| 9 | HMGCS1 | ENSG00000112972.15 | 2715 | 1.404 | 0.3229 | Yes | ||
| 10 | CARM1 | ENSG00000142453.11 | 2879 | 1.361 | 0.3475 | Yes | ||
| 11 | SREBF2 | ENSG00000198911.12 | 3043 | 1.323 | 0.3712 | Yes | ||
| 12 | FASN | ENSG00000169710.9 | 3072 | 1.316 | 0.3980 | Yes | ||
| 13 | SREBF1 | ENSG00000072310.17 | 3181 | 1.292 | 0.4224 | Yes | ||
| 14 | MVD | ENSG00000167508.12 | 3276 | 1.273 | 0.4467 | Yes | ||
| 15 | FDPS | ENSG00000160752.14 | 3502 | 1.225 | 0.4670 | Yes | ||
| 16 | DHCR7 | ENSG00000172893.15 | 3663 | 1.193 | 0.4881 | Yes | ||
| 17 | NCOA2 | ENSG00000140396.13 | 3691 | 1.187 | 0.5122 | Yes | ||
| 18 | FDFT1 | ENSG00000079459.13 | 3715 | 1.180 | 0.5362 | Yes | ||
| 19 | ACACA | ENSG00000278540.5 | 4007 | 1.126 | 0.5529 | Yes | ||
| 20 | PPARA | ENSG00000186951.16 | 4083 | 1.114 | 0.5744 | Yes | ||
| 21 | TM7SF2 | ENSG00000149809.15 | 6196 | 0.817 | 0.5424 | No | ||
| 22 | CREBBP | ENSG00000005339.14 | 7714 | 0.671 | 0.5211 | No | ||
| 23 | TBL1X | ENSG00000101849.17 | 8055 | 0.642 | 0.5266 | No | ||
| 24 | SP1 | ENSG00000185591.10 | 9660 | 0.525 | 0.5003 | No | ||
| 25 | GGPS1 | ENSG00000152904.11 | 10416 | 0.484 | 0.4928 | No | ||
| 26 | NCOA6 | ENSG00000198646.14 | 10986 | 0.446 | 0.4889 | No | ||
| 27 | ACACB | ENSG00000076555.15 | 16077 | 0.244 | 0.3758 | No | ||
| 28 | IDI1 | ENSG00000067064.11 | 20014 | 0.048 | 0.2854 | No | ||
| 29 | TGS1 | ENSG00000137574.11 | 20524 | 0.021 | 0.2741 | No | ||
| 30 | MTF1 | ENSG00000188786.10 | 23796 | -0.157 | 0.2014 | No | ||
| 31 | NFYA | ENSG00000001167.14 | 28408 | -0.331 | 0.1013 | No | ||
| 32 | TBL1XR1 | ENSG00000177565.16 | 30634 | -0.446 | 0.0589 | No | ||
| 33 | NFYB | ENSG00000120837.8 | 32930 | -0.578 | 0.0176 | No | ||
| 34 | SC5D | ENSG00000109929.10 | 34365 | -0.689 | -0.0013 | No | ||
| 35 | MED1 | ENSG00000125686.12 | 34391 | -0.691 | 0.0125 | No | ||
| 36 | CHD9 | ENSG00000177200.17 | 34510 | -0.703 | 0.0244 | No | ||
| 37 | PMVK | ENSG00000163344.6 | 36154 | -0.862 | 0.0042 | No | ||
| 38 | NFYC | ENSG00000066136.20 | 37025 | -0.969 | 0.0042 | No | ||
| 39 | ELOVL6 | ENSG00000170522.9 | 38394 | -1.186 | -0.0028 | No | ||
| 40 | NCOA1 | ENSG00000084676.15 | 39735 | -1.475 | -0.0032 | No | ||
| 41 | GPAM | ENSG00000119927.14 | 40785 | -1.781 | 0.0095 | No | ||
| 42 | SMARCD3 | ENSG00000082014.16 | 41524 | -2.144 | 0.0370 | No |