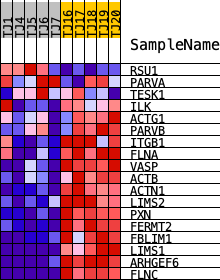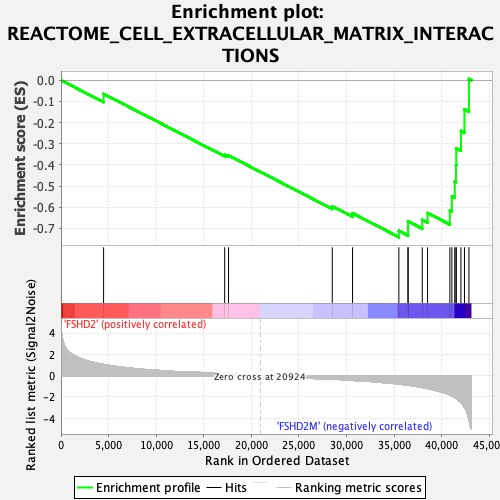
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
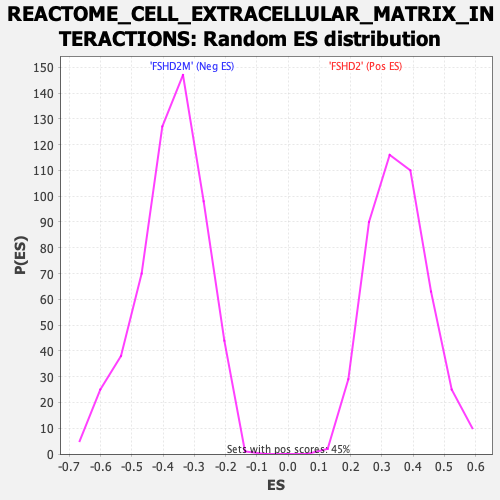
| GeneSet | REACTOME_CELL_EXTRACELLULAR_MATRIX_INTERACTIONS |
| Enrichment Score (ES) | -0.7412965 |
| Normalized Enrichment Score (NES) | -1.9851278 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0012679844 |
| FWER p-Value | 0.052 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | RSU1 | ENSG00000148484.18 | 4475 | 1.048 | -0.0645 | No | ||
| 2 | PARVA | ENSG00000197702.14 | 17194 | 0.188 | -0.3525 | No | ||
| 3 | TESK1 | ENSG00000107140.16 | 17596 | 0.169 | -0.3555 | No | ||
| 4 | ILK | ENSG00000166333.13 | 28496 | -0.336 | -0.5957 | No | ||
| 5 | ACTG1 | ENSG00000184009.12 | 30626 | -0.445 | -0.6284 | No | ||
| 6 | PARVB | ENSG00000188677.14 | 35491 | -0.786 | -0.7118 | Yes | ||
| 7 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.7005 | Yes | ||
| 8 | FLNA | ENSG00000196924.17 | 36458 | -0.895 | -0.6670 | Yes | ||
| 9 | VASP | ENSG00000125753.14 | 37949 | -1.108 | -0.6600 | Yes | ||
| 10 | ACTB | ENSG00000075624.16 | 38506 | -1.208 | -0.6276 | Yes | ||
| 11 | ACTN1 | ENSG00000072110.13 | 40848 | -1.801 | -0.6143 | Yes | ||
| 12 | LIMS2 | ENSG00000072163.19 | 41058 | -1.892 | -0.5482 | Yes | ||
| 13 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.4787 | Yes | ||
| 14 | FERMT2 | ENSG00000073712.15 | 41491 | -2.127 | -0.4018 | Yes | ||
| 15 | FBLIM1 | ENSG00000162458.13 | 41520 | -2.143 | -0.3220 | Yes | ||
| 16 | LIMS1 | ENSG00000169756.16 | 42011 | -2.498 | -0.2396 | Yes | ||
| 17 | ARHGEF6 | ENSG00000129675.16 | 42382 | -2.958 | -0.1372 | Yes | ||
| 18 | FLNC | ENSG00000128591.15 | 42852 | -4.113 | 0.0062 | Yes |