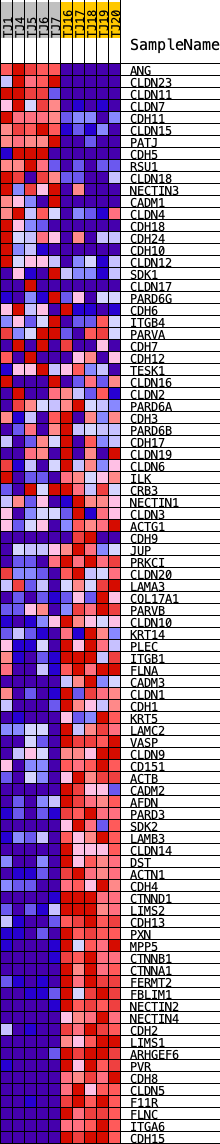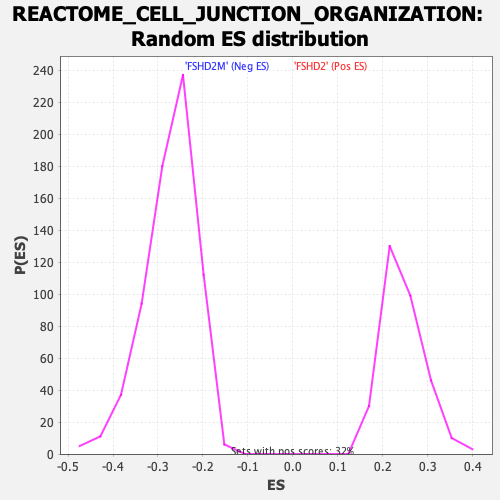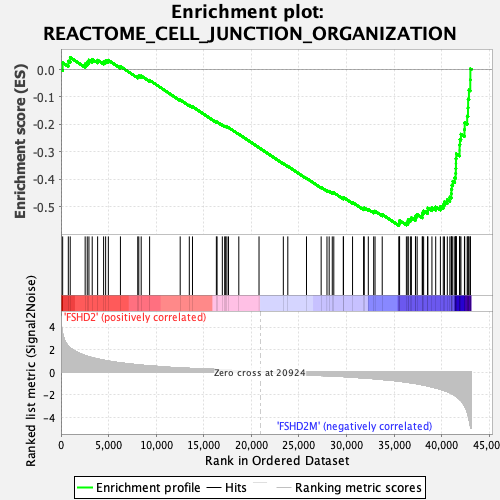
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_CELL_JUNCTION_ORGANIZATION |
| Enrichment Score (ES) | -0.5680938 |
| Normalized Enrichment Score (NES) | -2.075948 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.482205E-4 |
| FWER p-Value | 0.008 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ANG | ENSG00000214274.9 | 174 | 3.361 | 0.0247 | No | ||
| 2 | CLDN23 | ENSG00000253958.2 | 768 | 2.293 | 0.0306 | No | ||
| 3 | CLDN11 | ENSG00000013297.11 | 969 | 2.122 | 0.0441 | No | ||
| 4 | CLDN7 | ENSG00000181885.18 | 2537 | 1.461 | 0.0201 | No | ||
| 5 | CDH11 | ENSG00000140937.14 | 2757 | 1.391 | 0.0270 | No | ||
| 6 | CLDN15 | ENSG00000106404.13 | 2934 | 1.348 | 0.0344 | No | ||
| 7 | PATJ | ENSG00000132849.20 | 3279 | 1.273 | 0.0373 | No | ||
| 8 | CDH5 | ENSG00000179776.19 | 3847 | 1.155 | 0.0340 | No | ||
| 9 | RSU1 | ENSG00000148484.18 | 4475 | 1.048 | 0.0284 | No | ||
| 10 | CLDN18 | ENSG00000066405.13 | 4688 | 1.016 | 0.0322 | No | ||
| 11 | NECTIN3 | ENSG00000177707.11 | 4974 | 0.972 | 0.0339 | No | ||
| 12 | CADM1 | ENSG00000182985.17 | 6243 | 0.812 | 0.0113 | No | ||
| 13 | CLDN4 | ENSG00000189143.9 | 8062 | 0.642 | -0.0254 | No | ||
| 14 | CDH18 | ENSG00000145526.12 | 8172 | 0.631 | -0.0225 | No | ||
| 15 | CDH24 | ENSG00000139880.19 | 8424 | 0.611 | -0.0232 | No | ||
| 16 | CDH10 | ENSG00000040731.10 | 9308 | 0.550 | -0.0390 | No | ||
| 17 | CLDN12 | ENSG00000157224.16 | 12521 | 0.365 | -0.1105 | No | ||
| 18 | SDK1 | ENSG00000146555.19 | 13478 | 0.329 | -0.1299 | No | ||
| 19 | CLDN17 | ENSG00000156282.5 | 13812 | 0.322 | -0.1349 | No | ||
| 20 | PARD6G | ENSG00000178184.16 | 16320 | 0.229 | -0.1912 | No | ||
| 21 | CDH6 | ENSG00000113361.13 | 16393 | 0.226 | -0.1909 | No | ||
| 22 | ITGB4 | ENSG00000132470.14 | 16947 | 0.199 | -0.2021 | No | ||
| 23 | PARVA | ENSG00000197702.14 | 17194 | 0.188 | -0.2062 | No | ||
| 24 | CDH7 | ENSG00000081138.14 | 17296 | 0.183 | -0.2070 | No | ||
| 25 | CDH12 | ENSG00000154162.15 | 17428 | 0.176 | -0.2085 | No | ||
| 26 | TESK1 | ENSG00000107140.16 | 17596 | 0.169 | -0.2109 | No | ||
| 27 | CLDN16 | ENSG00000113946.3 | 18678 | 0.114 | -0.2351 | No | ||
| 28 | CLDN2 | ENSG00000165376.10 | 20801 | 0.007 | -0.2843 | No | ||
| 29 | PARD6A | ENSG00000102981.9 | 23355 | -0.133 | -0.3425 | No | ||
| 30 | CDH3 | ENSG00000062038.14 | 23825 | -0.160 | -0.3521 | No | ||
| 31 | PARD6B | ENSG00000124171.9 | 25788 | -0.246 | -0.3956 | No | ||
| 32 | CDH17 | ENSG00000079112.10 | 27328 | -0.297 | -0.4288 | No | ||
| 33 | CLDN19 | ENSG00000164007.11 | 27939 | -0.305 | -0.4404 | No | ||
| 34 | CLDN6 | ENSG00000184697.7 | 28170 | -0.321 | -0.4430 | No | ||
| 35 | ILK | ENSG00000166333.13 | 28496 | -0.336 | -0.4476 | No | ||
| 36 | CRB3 | ENSG00000130545.16 | 28656 | -0.341 | -0.4484 | No | ||
| 37 | NECTIN1 | ENSG00000110400.11 | 29652 | -0.385 | -0.4682 | No | ||
| 38 | CLDN3 | ENSG00000165215.6 | 29677 | -0.385 | -0.4655 | No | ||
| 39 | ACTG1 | ENSG00000184009.12 | 30626 | -0.445 | -0.4837 | No | ||
| 40 | CDH9 | ENSG00000113100.10 | 31783 | -0.501 | -0.5063 | No | ||
| 41 | JUP | ENSG00000173801.17 | 31855 | -0.505 | -0.5036 | No | ||
| 42 | PRKCI | ENSG00000163558.13 | 32282 | -0.533 | -0.5090 | No | ||
| 43 | CLDN20 | ENSG00000171217.5 | 32845 | -0.572 | -0.5171 | No | ||
| 44 | LAMA3 | ENSG00000053747.16 | 32988 | -0.582 | -0.5155 | No | ||
| 45 | COL17A1 | ENSG00000065618.21 | 33743 | -0.641 | -0.5275 | No | ||
| 46 | PARVB | ENSG00000188677.14 | 35491 | -0.786 | -0.5614 | Yes | ||
| 47 | CLDN10 | ENSG00000134873.10 | 35539 | -0.791 | -0.5557 | Yes | ||
| 48 | KRT14 | ENSG00000186847.6 | 35568 | -0.794 | -0.5496 | Yes | ||
| 49 | PLEC | ENSG00000178209.15 | 36280 | -0.875 | -0.5586 | Yes | ||
| 50 | ITGB1 | ENSG00000150093.19 | 36454 | -0.895 | -0.5549 | Yes | ||
| 51 | FLNA | ENSG00000196924.17 | 36458 | -0.895 | -0.5474 | Yes | ||
| 52 | CADM3 | ENSG00000162706.13 | 36703 | -0.930 | -0.5451 | Yes | ||
| 53 | CLDN1 | ENSG00000163347.6 | 36807 | -0.941 | -0.5394 | Yes | ||
| 54 | CDH1 | ENSG00000039068.19 | 37235 | -0.997 | -0.5408 | Yes | ||
| 55 | KRT5 | ENSG00000186081.12 | 37249 | -0.999 | -0.5326 | Yes | ||
| 56 | LAMC2 | ENSG00000058085.15 | 37414 | -1.022 | -0.5276 | Yes | ||
| 57 | VASP | ENSG00000125753.14 | 37949 | -1.108 | -0.5306 | Yes | ||
| 58 | CLDN9 | ENSG00000213937.4 | 37970 | -1.113 | -0.5215 | Yes | ||
| 59 | CD151 | ENSG00000177697.19 | 38093 | -1.134 | -0.5146 | Yes | ||
| 60 | ACTB | ENSG00000075624.16 | 38506 | -1.208 | -0.5139 | Yes | ||
| 61 | CADM2 | ENSG00000175161.13 | 38535 | -1.213 | -0.5041 | Yes | ||
| 62 | AFDN | ENSG00000130396.20 | 38965 | -1.301 | -0.5030 | Yes | ||
| 63 | PARD3 | ENSG00000148498.16 | 39361 | -1.390 | -0.5003 | Yes | ||
| 64 | SDK2 | ENSG00000069188.17 | 39854 | -1.507 | -0.4988 | Yes | ||
| 65 | LAMB3 | ENSG00000196878.15 | 40157 | -1.589 | -0.4922 | Yes | ||
| 66 | CLDN14 | ENSG00000159261.12 | 40281 | -1.620 | -0.4812 | Yes | ||
| 67 | DST | ENSG00000151914.20 | 40581 | -1.710 | -0.4735 | Yes | ||
| 68 | ACTN1 | ENSG00000072110.13 | 40848 | -1.801 | -0.4643 | Yes | ||
| 69 | CDH4 | ENSG00000179242.16 | 40997 | -1.866 | -0.4518 | Yes | ||
| 70 | CTNND1 | ENSG00000198561.13 | 41012 | -1.873 | -0.4361 | Yes | ||
| 71 | LIMS2 | ENSG00000072163.19 | 41058 | -1.892 | -0.4209 | Yes | ||
| 72 | CDH13 | ENSG00000140945.17 | 41157 | -1.936 | -0.4067 | Yes | ||
| 73 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.3940 | Yes | ||
| 74 | MPP5 | ENSG00000072415.9 | 41456 | -2.104 | -0.3781 | Yes | ||
| 75 | CTNNB1 | ENSG00000168036.18 | 41475 | -2.117 | -0.3604 | Yes | ||
| 76 | CTNNA1 | ENSG00000044115.21 | 41488 | -2.125 | -0.3425 | Yes | ||
| 77 | FERMT2 | ENSG00000073712.15 | 41491 | -2.127 | -0.3243 | Yes | ||
| 78 | FBLIM1 | ENSG00000162458.13 | 41520 | -2.143 | -0.3066 | Yes | ||
| 79 | NECTIN2 | ENSG00000130202.10 | 41868 | -2.379 | -0.2944 | Yes | ||
| 80 | NECTIN4 | ENSG00000143217.9 | 41869 | -2.381 | -0.2740 | Yes | ||
| 81 | CDH2 | ENSG00000170558.9 | 41908 | -2.414 | -0.2542 | Yes | ||
| 82 | LIMS1 | ENSG00000169756.16 | 42011 | -2.498 | -0.2352 | Yes | ||
| 83 | ARHGEF6 | ENSG00000129675.16 | 42382 | -2.958 | -0.2185 | Yes | ||
| 84 | PVR | ENSG00000073008.15 | 42407 | -2.989 | -0.1935 | Yes | ||
| 85 | CDH8 | ENSG00000150394.14 | 42668 | -3.494 | -0.1696 | Yes | ||
| 86 | CLDN5 | ENSG00000184113.9 | 42752 | -3.728 | -0.1397 | Yes | ||
| 87 | F11R | ENSG00000158769.18 | 42767 | -3.776 | -0.1077 | Yes | ||
| 88 | FLNC | ENSG00000128591.15 | 42852 | -4.113 | -0.0744 | Yes | ||
| 89 | ITGA6 | ENSG00000091409.15 | 42992 | -4.680 | -0.0376 | Yes | ||
| 90 | CDH15 | ENSG00000129910.8 | 43003 | -4.739 | 0.0027 | Yes |