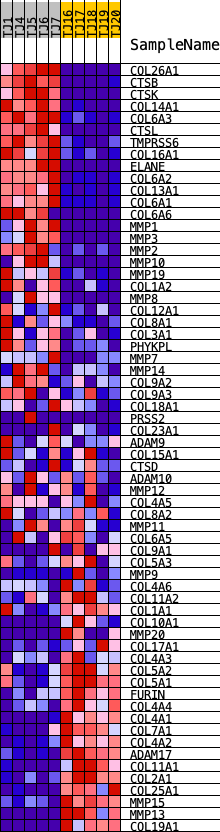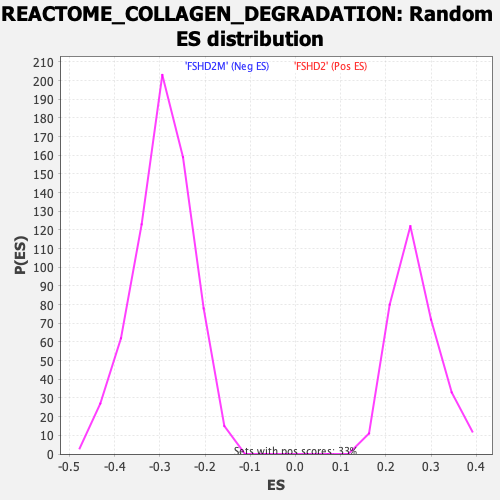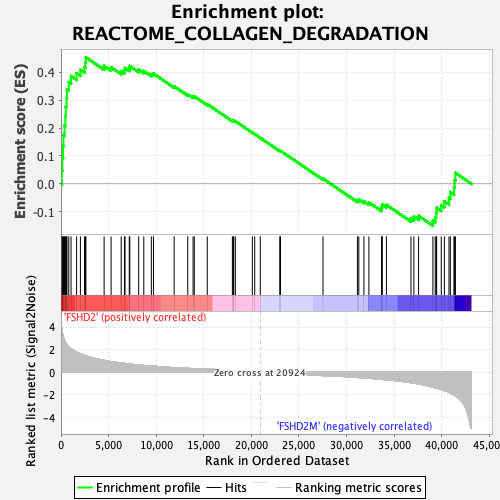
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_COLLAGEN_DEGRADATION |
| Enrichment Score (ES) | 0.4529119 |
| Normalized Enrichment Score (NES) | 1.7154838 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06491897 |
| FWER p-Value | 0.954 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | COL26A1 | ENSG00000160963.14 | 88 | 3.782 | 0.0480 | Yes | ||
| 2 | CTSB | ENSG00000164733.21 | 145 | 3.504 | 0.0931 | Yes | ||
| 3 | CTSK | ENSG00000143387.13 | 210 | 3.273 | 0.1350 | Yes | ||
| 4 | COL14A1 | ENSG00000187955.12 | 257 | 3.107 | 0.1750 | Yes | ||
| 5 | COL6A3 | ENSG00000163359.15 | 371 | 2.812 | 0.2096 | Yes | ||
| 6 | CTSL | ENSG00000135047.15 | 458 | 2.665 | 0.2429 | Yes | ||
| 7 | TMPRSS6 | ENSG00000187045.18 | 499 | 2.600 | 0.2764 | Yes | ||
| 8 | COL16A1 | ENSG00000084636.18 | 589 | 2.483 | 0.3072 | Yes | ||
| 9 | ELANE | ENSG00000197561.7 | 623 | 2.444 | 0.3388 | Yes | ||
| 10 | COL6A2 | ENSG00000142173.15 | 816 | 2.246 | 0.3641 | Yes | ||
| 11 | COL13A1 | ENSG00000197467.14 | 1056 | 2.080 | 0.3861 | Yes | ||
| 12 | COL6A1 | ENSG00000142156.14 | 1639 | 1.786 | 0.3962 | Yes | ||
| 13 | COL6A6 | ENSG00000206384.10 | 2037 | 1.617 | 0.4084 | Yes | ||
| 14 | MMP1 | ENSG00000196611.5 | 2473 | 1.481 | 0.4179 | Yes | ||
| 15 | MMP3 | ENSG00000149968.12 | 2562 | 1.453 | 0.4351 | Yes | ||
| 16 | MMP2 | ENSG00000087245.13 | 2616 | 1.437 | 0.4529 | Yes | ||
| 17 | MMP10 | ENSG00000166670.10 | 4529 | 1.040 | 0.4223 | No | ||
| 18 | MMP19 | ENSG00000123342.16 | 5262 | 0.933 | 0.4176 | No | ||
| 19 | COL1A2 | ENSG00000164692.17 | 6324 | 0.804 | 0.4036 | No | ||
| 20 | MMP8 | ENSG00000118113.12 | 6665 | 0.771 | 0.4060 | No | ||
| 21 | COL12A1 | ENSG00000111799.21 | 6723 | 0.765 | 0.4148 | No | ||
| 22 | COL8A1 | ENSG00000144810.16 | 7170 | 0.722 | 0.4140 | No | ||
| 23 | COL3A1 | ENSG00000168542.14 | 7229 | 0.715 | 0.4221 | No | ||
| 24 | PHYKPL | ENSG00000175309.15 | 8161 | 0.632 | 0.4088 | No | ||
| 25 | MMP7 | ENSG00000137673.9 | 8697 | 0.591 | 0.4042 | No | ||
| 26 | MMP14 | ENSG00000157227.13 | 9489 | 0.539 | 0.3930 | No | ||
| 27 | COL9A2 | ENSG00000049089.15 | 9709 | 0.523 | 0.3948 | No | ||
| 28 | COL9A3 | ENSG00000092758.18 | 11887 | 0.394 | 0.3495 | No | ||
| 29 | COL18A1 | ENSG00000182871.16 | 13315 | 0.339 | 0.3208 | No | ||
| 30 | PRSS2 | ENSG00000275896.5 | 13875 | 0.322 | 0.3121 | No | ||
| 31 | COL23A1 | ENSG00000050767.17 | 13999 | 0.317 | 0.3134 | No | ||
| 32 | ADAM9 | ENSG00000168615.12 | 15364 | 0.282 | 0.2855 | No | ||
| 33 | COL15A1 | ENSG00000204291.11 | 18002 | 0.146 | 0.2262 | No | ||
| 34 | CTSD | ENSG00000117984.14 | 18089 | 0.141 | 0.2261 | No | ||
| 35 | ADAM10 | ENSG00000137845.15 | 18147 | 0.139 | 0.2266 | No | ||
| 36 | MMP12 | ENSG00000262406.3 | 18315 | 0.129 | 0.2244 | No | ||
| 37 | COL4A5 | ENSG00000188153.13 | 20097 | 0.043 | 0.1836 | No | ||
| 38 | COL8A2 | ENSG00000171812.13 | 20347 | 0.029 | 0.1782 | No | ||
| 39 | MMP11 | ENSG00000099953.10 | 20933 | -0.001 | 0.1646 | No | ||
| 40 | COL6A5 | ENSG00000172752.14 | 22980 | -0.111 | 0.1186 | No | ||
| 41 | COL9A1 | ENSG00000112280.16 | 23057 | -0.115 | 0.1183 | No | ||
| 42 | COL5A3 | ENSG00000080573.7 | 27521 | -0.299 | 0.0187 | No | ||
| 43 | MMP9 | ENSG00000100985.7 | 31139 | -0.471 | -0.0591 | No | ||
| 44 | COL4A6 | ENSG00000197565.15 | 31285 | -0.477 | -0.0562 | No | ||
| 45 | COL11A2 | ENSG00000204248.10 | 31819 | -0.502 | -0.0619 | No | ||
| 46 | COL1A1 | ENSG00000108821.13 | 32343 | -0.536 | -0.0669 | No | ||
| 47 | COL10A1 | ENSG00000123500.10 | 33676 | -0.637 | -0.0895 | No | ||
| 48 | MMP20 | ENSG00000137674.4 | 33705 | -0.639 | -0.0816 | No | ||
| 49 | COL17A1 | ENSG00000065618.21 | 33743 | -0.641 | -0.0740 | No | ||
| 50 | COL4A3 | ENSG00000169031.19 | 34183 | -0.675 | -0.0753 | No | ||
| 51 | COL5A2 | ENSG00000204262.14 | 36764 | -0.936 | -0.1228 | No | ||
| 52 | COL5A1 | ENSG00000130635.15 | 37069 | -0.977 | -0.1169 | No | ||
| 53 | FURIN | ENSG00000140564.12 | 37563 | -1.044 | -0.1146 | No | ||
| 54 | COL4A4 | ENSG00000081052.12 | 39066 | -1.323 | -0.1319 | No | ||
| 55 | COL4A1 | ENSG00000187498.16 | 39310 | -1.380 | -0.1193 | No | ||
| 56 | COL7A1 | ENSG00000114270.17 | 39421 | -1.404 | -0.1033 | No | ||
| 57 | COL4A2 | ENSG00000134871.19 | 39460 | -1.413 | -0.0854 | No | ||
| 58 | ADAM17 | ENSG00000151694.14 | 39944 | -1.533 | -0.0764 | No | ||
| 59 | COL11A1 | ENSG00000060718.22 | 40275 | -1.618 | -0.0626 | No | ||
| 60 | COL2A1 | ENSG00000139219.19 | 40754 | -1.768 | -0.0503 | No | ||
| 61 | COL25A1 | ENSG00000188517.16 | 40900 | -1.824 | -0.0295 | No | ||
| 62 | MMP15 | ENSG00000102996.5 | 41275 | -1.992 | -0.0118 | No | ||
| 63 | MMP13 | ENSG00000137745.12 | 41348 | -2.033 | 0.0134 | No | ||
| 64 | COL19A1 | ENSG00000082293.13 | 41431 | -2.090 | 0.0392 | No |