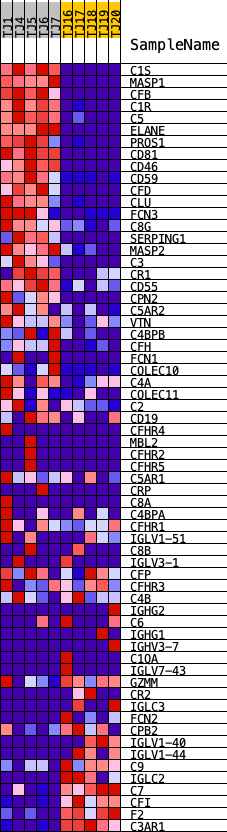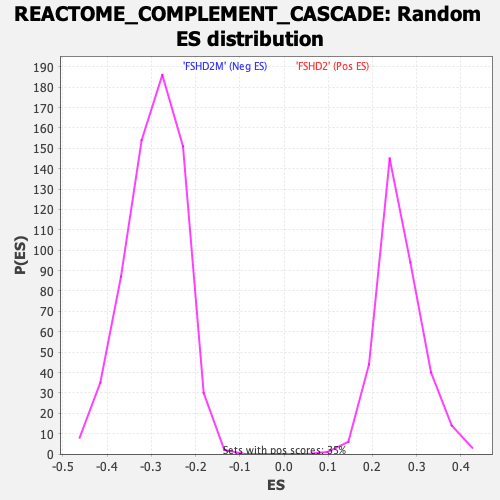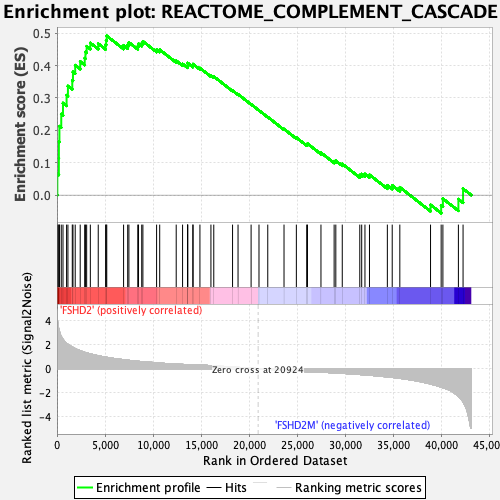
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_COMPLEMENT_CASCADE |
| Enrichment Score (ES) | 0.49139106 |
| Normalized Enrichment Score (NES) | 1.871737 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.019802848 |
| FWER p-Value | 0.417 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | C1S | ENSG00000182326.15 | 34 | 4.267 | 0.0657 | Yes | ||
| 2 | MASP1 | ENSG00000127241.16 | 191 | 3.323 | 0.1139 | Yes | ||
| 3 | CFB | ENSG00000243649.8 | 196 | 3.306 | 0.1654 | Yes | ||
| 4 | C1R | ENSG00000159403.18 | 251 | 3.121 | 0.2128 | Yes | ||
| 5 | C5 | ENSG00000106804.7 | 443 | 2.687 | 0.2502 | Yes | ||
| 6 | ELANE | ENSG00000197561.7 | 623 | 2.444 | 0.2842 | Yes | ||
| 7 | PROS1 | ENSG00000184500.15 | 997 | 2.107 | 0.3083 | Yes | ||
| 8 | CD81 | ENSG00000110651.11 | 1137 | 2.025 | 0.3367 | Yes | ||
| 9 | CD46 | ENSG00000117335.19 | 1590 | 1.805 | 0.3543 | Yes | ||
| 10 | CD59 | ENSG00000085063.16 | 1662 | 1.770 | 0.3803 | Yes | ||
| 11 | CFD | ENSG00000197766.8 | 1895 | 1.671 | 0.4009 | Yes | ||
| 12 | CLU | ENSG00000120885.22 | 2415 | 1.500 | 0.4123 | Yes | ||
| 13 | FCN3 | ENSG00000142748.12 | 2884 | 1.360 | 0.4226 | Yes | ||
| 14 | C8G | ENSG00000176919.13 | 2959 | 1.343 | 0.4418 | Yes | ||
| 15 | SERPING1 | ENSG00000149131.15 | 3087 | 1.313 | 0.4593 | Yes | ||
| 16 | MASP2 | ENSG00000009724.17 | 3469 | 1.231 | 0.4697 | Yes | ||
| 17 | C3 | ENSG00000125730.17 | 4292 | 1.078 | 0.4674 | Yes | ||
| 18 | CR1 | ENSG00000203710.11 | 5061 | 0.959 | 0.4645 | Yes | ||
| 19 | CD55 | ENSG00000196352.15 | 5115 | 0.952 | 0.4781 | Yes | ||
| 20 | CPN2 | ENSG00000178772.7 | 5178 | 0.943 | 0.4914 | Yes | ||
| 21 | C5AR2 | ENSG00000134830.6 | 6930 | 0.744 | 0.4623 | No | ||
| 22 | VTN | ENSG00000109072.14 | 7359 | 0.704 | 0.4634 | No | ||
| 23 | C4BPB | ENSG00000123843.13 | 7483 | 0.692 | 0.4713 | No | ||
| 24 | CFH | ENSG00000000971.15 | 8408 | 0.612 | 0.4594 | No | ||
| 25 | FCN1 | ENSG00000085265.11 | 8484 | 0.608 | 0.4671 | No | ||
| 26 | COLEC10 | ENSG00000184374.3 | 8818 | 0.582 | 0.4684 | No | ||
| 27 | C4A | ENSG00000244731.8 | 8939 | 0.572 | 0.4746 | No | ||
| 28 | COLEC11 | ENSG00000118004.17 | 10366 | 0.487 | 0.4490 | No | ||
| 29 | C2 | ENSG00000166278.15 | 10689 | 0.466 | 0.4488 | No | ||
| 30 | CD19 | ENSG00000177455.14 | 12407 | 0.369 | 0.4147 | No | ||
| 31 | CFHR4 | ENSG00000134365.13 | 13070 | 0.350 | 0.4048 | No | ||
| 32 | MBL2 | ENSG00000165471.6 | 13582 | 0.322 | 0.3979 | No | ||
| 33 | CFHR2 | ENSG00000080910.13 | 13602 | 0.322 | 0.4025 | No | ||
| 34 | CFHR5 | ENSG00000134389.9 | 13603 | 0.322 | 0.4075 | No | ||
| 35 | C5AR1 | ENSG00000197405.8 | 14141 | 0.308 | 0.3999 | No | ||
| 36 | CRP | ENSG00000132693.12 | 14159 | 0.308 | 0.4043 | No | ||
| 37 | C8A | ENSG00000157131.10 | 14876 | 0.287 | 0.3921 | No | ||
| 38 | C4BPA | ENSG00000123838.11 | 16021 | 0.248 | 0.3694 | No | ||
| 39 | CFHR1 | ENSG00000244414.6 | 16317 | 0.229 | 0.3661 | No | ||
| 40 | IGLV1-51 | ENSG00000211644.3 | 18272 | 0.132 | 0.3228 | No | ||
| 41 | C8B | ENSG00000021852.13 | 18843 | 0.105 | 0.3112 | No | ||
| 42 | IGLV3-1 | ENSG00000211673.2 | 20202 | 0.037 | 0.2803 | No | ||
| 43 | CFP | ENSG00000126759.13 | 21023 | -0.005 | 0.2613 | No | ||
| 44 | CFHR3 | ENSG00000116785.13 | 21936 | -0.053 | 0.2409 | No | ||
| 45 | C4B | ENSG00000224389.9 | 23628 | -0.148 | 0.2040 | No | ||
| 46 | IGHG2 | ENSG00000211893.4 | 24912 | -0.223 | 0.1776 | No | ||
| 47 | C6 | ENSG00000039537.13 | 25989 | -0.260 | 0.1567 | No | ||
| 48 | IGHG1 | ENSG00000211896.7 | 26070 | -0.263 | 0.1589 | No | ||
| 49 | IGHV3-7 | ENSG00000211938.2 | 27473 | -0.297 | 0.1310 | No | ||
| 50 | C1QA | ENSG00000173372.17 | 28844 | -0.351 | 0.1047 | No | ||
| 51 | IGLV7-43 | ENSG00000211652.2 | 29019 | -0.351 | 0.1061 | No | ||
| 52 | GZMM | ENSG00000197540.8 | 29690 | -0.386 | 0.0966 | No | ||
| 53 | CR2 | ENSG00000117322.17 | 31514 | -0.489 | 0.0619 | No | ||
| 54 | IGLC3 | ENSG00000211679.2 | 31713 | -0.498 | 0.0650 | No | ||
| 55 | FCN2 | ENSG00000160339.16 | 32052 | -0.518 | 0.0652 | No | ||
| 56 | CPB2 | ENSG00000080618.16 | 32522 | -0.544 | 0.0628 | No | ||
| 57 | IGLV1-40 | ENSG00000211653.2 | 34383 | -0.691 | 0.0304 | No | ||
| 58 | IGLV1-44 | ENSG00000211651.3 | 34889 | -0.732 | 0.0301 | No | ||
| 59 | C9 | ENSG00000113600.11 | 35681 | -0.807 | 0.0243 | No | ||
| 60 | IGLC2 | ENSG00000211677.2 | 38878 | -1.284 | -0.0299 | No | ||
| 61 | C7 | ENSG00000112936.19 | 39983 | -1.544 | -0.0315 | No | ||
| 62 | CFI | ENSG00000205403.13 | 40164 | -1.591 | -0.0109 | No | ||
| 63 | F2 | ENSG00000180210.14 | 41777 | -2.308 | -0.0123 | No | ||
| 64 | C3AR1 | ENSG00000171860.5 | 42263 | -2.787 | 0.0199 | No |