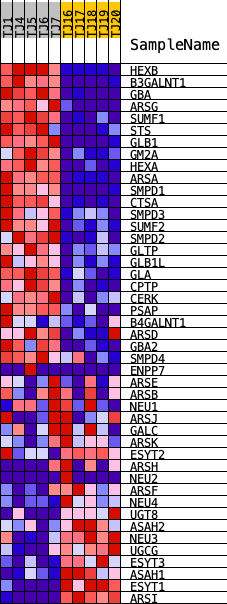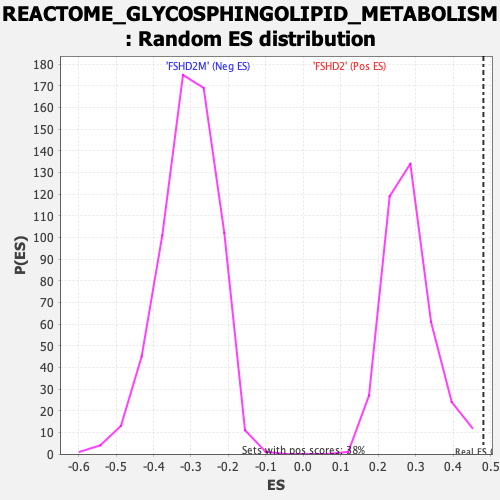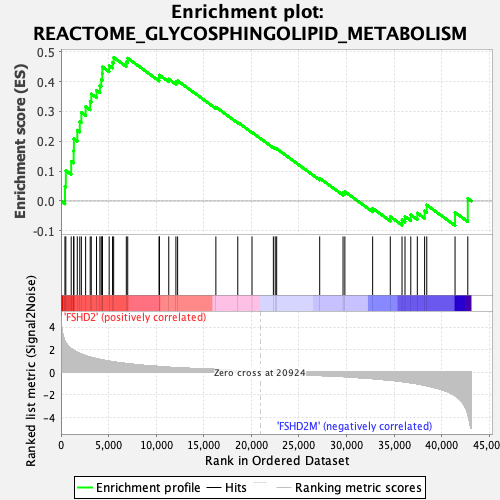
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_GLYCOSPHINGOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.4805664 |
| Normalized Enrichment Score (NES) | 1.7062752 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.068300046 |
| FWER p-Value | 0.962 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HEXB | ENSG00000049860.14 | 415 | 2.734 | 0.0487 | Yes | ||
| 2 | B3GALNT1 | ENSG00000169255.15 | 507 | 2.592 | 0.1018 | Yes | ||
| 3 | GBA | ENSG00000177628.16 | 1072 | 2.067 | 0.1328 | Yes | ||
| 4 | ARSG | ENSG00000141337.12 | 1326 | 1.924 | 0.1680 | Yes | ||
| 5 | SUMF1 | ENSG00000144455.14 | 1354 | 1.904 | 0.2079 | Yes | ||
| 6 | STS | ENSG00000101846.7 | 1712 | 1.750 | 0.2370 | Yes | ||
| 7 | GLB1 | ENSG00000170266.16 | 1972 | 1.642 | 0.2659 | Yes | ||
| 8 | GM2A | ENSG00000196743.9 | 2138 | 1.587 | 0.2960 | Yes | ||
| 9 | HEXA | ENSG00000213614.9 | 2587 | 1.446 | 0.3164 | Yes | ||
| 10 | ARSA | ENSG00000100299.18 | 3068 | 1.317 | 0.3333 | Yes | ||
| 11 | SMPD1 | ENSG00000166311.10 | 3178 | 1.292 | 0.3583 | Yes | ||
| 12 | CTSA | ENSG00000064601.18 | 3731 | 1.177 | 0.3706 | Yes | ||
| 13 | SMPD3 | ENSG00000103056.12 | 4087 | 1.114 | 0.3861 | Yes | ||
| 14 | SUMF2 | ENSG00000129103.18 | 4226 | 1.088 | 0.4061 | Yes | ||
| 15 | SMPD2 | ENSG00000135587.9 | 4336 | 1.071 | 0.4264 | Yes | ||
| 16 | GLTP | ENSG00000139433.10 | 4348 | 1.069 | 0.4490 | Yes | ||
| 17 | GLB1L | ENSG00000163521.16 | 5059 | 0.960 | 0.4529 | Yes | ||
| 18 | GLA | ENSG00000102393.10 | 5414 | 0.912 | 0.4642 | Yes | ||
| 19 | CPTP | ENSG00000224051.7 | 5532 | 0.896 | 0.4806 | Yes | ||
| 20 | CERK | ENSG00000100422.14 | 6861 | 0.751 | 0.4658 | No | ||
| 21 | PSAP | ENSG00000197746.14 | 7009 | 0.737 | 0.4781 | No | ||
| 22 | B4GALNT1 | ENSG00000135454.14 | 10301 | 0.491 | 0.4121 | No | ||
| 23 | ARSD | ENSG00000006756.16 | 10345 | 0.488 | 0.4215 | No | ||
| 24 | GBA2 | ENSG00000070610.14 | 11319 | 0.426 | 0.4080 | No | ||
| 25 | SMPD4 | ENSG00000136699.19 | 12070 | 0.384 | 0.3988 | No | ||
| 26 | ENPP7 | ENSG00000182156.10 | 12251 | 0.375 | 0.4026 | No | ||
| 27 | ARSE | ENSG00000157399.15 | 16268 | 0.232 | 0.3143 | No | ||
| 28 | ARSB | ENSG00000113273.17 | 18569 | 0.120 | 0.2635 | No | ||
| 29 | NEU1 | ENSG00000204386.11 | 20064 | 0.045 | 0.2297 | No | ||
| 30 | ARSJ | ENSG00000180801.14 | 22309 | -0.075 | 0.1792 | No | ||
| 31 | GALC | ENSG00000054983.17 | 22503 | -0.084 | 0.1766 | No | ||
| 32 | ARSK | ENSG00000164291.17 | 22655 | -0.093 | 0.1750 | No | ||
| 33 | ESYT2 | ENSG00000117868.17 | 27175 | -0.286 | 0.0762 | No | ||
| 34 | ARSH | ENSG00000205667.2 | 29627 | -0.383 | 0.0275 | No | ||
| 35 | NEU2 | ENSG00000115488.3 | 29819 | -0.394 | 0.0314 | No | ||
| 36 | ARSF | ENSG00000062096.15 | 32737 | -0.564 | -0.0243 | No | ||
| 37 | NEU4 | ENSG00000204099.11 | 34590 | -0.709 | -0.0521 | No | ||
| 38 | UGT8 | ENSG00000174607.11 | 35823 | -0.820 | -0.0632 | No | ||
| 39 | ASAH2 | ENSG00000188611.15 | 36136 | -0.860 | -0.0521 | No | ||
| 40 | NEU3 | ENSG00000162139.10 | 36743 | -0.935 | -0.0463 | No | ||
| 41 | UGCG | ENSG00000148154.10 | 37430 | -1.024 | -0.0404 | No | ||
| 42 | ESYT3 | ENSG00000158220.14 | 38196 | -1.153 | -0.0336 | No | ||
| 43 | ASAH1 | ENSG00000104763.19 | 38422 | -1.192 | -0.0134 | No | ||
| 44 | ESYT1 | ENSG00000139641.12 | 41393 | -2.061 | -0.0384 | No | ||
| 45 | ARSI | ENSG00000183876.9 | 42731 | -3.679 | 0.0090 | No |