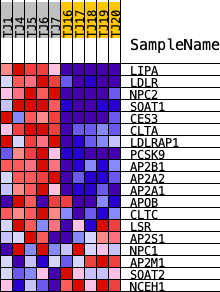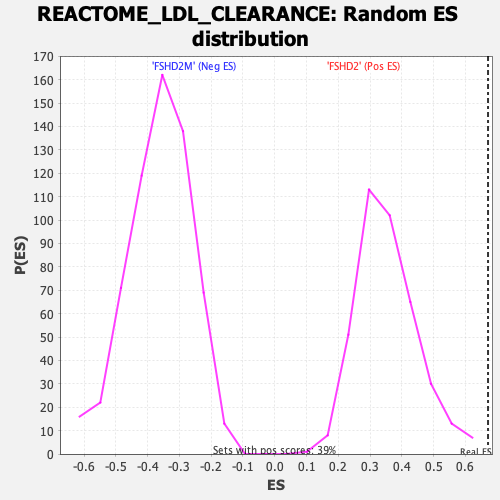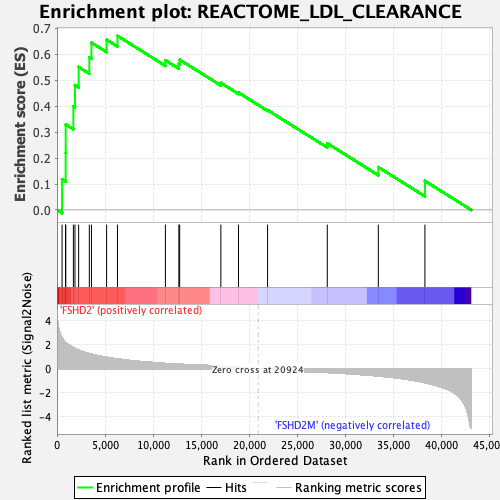
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_LDL_CLEARANCE |
| Enrichment Score (ES) | 0.6713195 |
| Normalized Enrichment Score (NES) | 1.901623 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.01916417 |
| FWER p-Value | 0.311 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LIPA | ENSG00000107798.18 | 528 | 2.571 | 0.1182 | Yes | ||
| 2 | LDLR | ENSG00000130164.13 | 900 | 2.174 | 0.2198 | Yes | ||
| 3 | NPC2 | ENSG00000119655.10 | 904 | 2.166 | 0.3297 | Yes | ||
| 4 | SOAT1 | ENSG00000057252.13 | 1704 | 1.753 | 0.4001 | Yes | ||
| 5 | CES3 | ENSG00000172828.13 | 1871 | 1.680 | 0.4815 | Yes | ||
| 6 | CLTA | ENSG00000122705.17 | 2253 | 1.549 | 0.5512 | Yes | ||
| 7 | LDLRAP1 | ENSG00000157978.11 | 3364 | 1.252 | 0.5890 | Yes | ||
| 8 | PCSK9 | ENSG00000169174.10 | 3586 | 1.208 | 0.6451 | Yes | ||
| 9 | AP2B1 | ENSG00000006125.18 | 5163 | 0.946 | 0.6566 | Yes | ||
| 10 | AP2A2 | ENSG00000183020.13 | 6293 | 0.807 | 0.6713 | Yes | ||
| 11 | AP2A1 | ENSG00000196961.12 | 11282 | 0.428 | 0.5773 | No | ||
| 12 | APOB | ENSG00000084674.14 | 12681 | 0.362 | 0.5632 | No | ||
| 13 | CLTC | ENSG00000141367.12 | 12768 | 0.357 | 0.5794 | No | ||
| 14 | LSR | ENSG00000105699.16 | 17054 | 0.196 | 0.4899 | No | ||
| 15 | AP2S1 | ENSG00000042753.11 | 18891 | 0.104 | 0.4525 | No | ||
| 16 | NPC1 | ENSG00000141458.13 | 21917 | -0.052 | 0.3850 | No | ||
| 17 | AP2M1 | ENSG00000161203.13 | 28129 | -0.319 | 0.2571 | No | ||
| 18 | SOAT2 | ENSG00000167780.12 | 33443 | -0.616 | 0.1651 | No | ||
| 19 | NCEH1 | ENSG00000144959.10 | 38293 | -1.171 | 0.1120 | No |