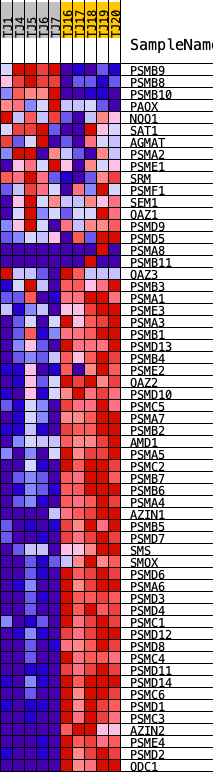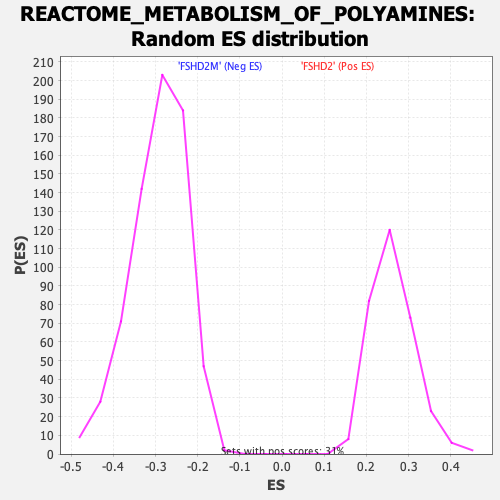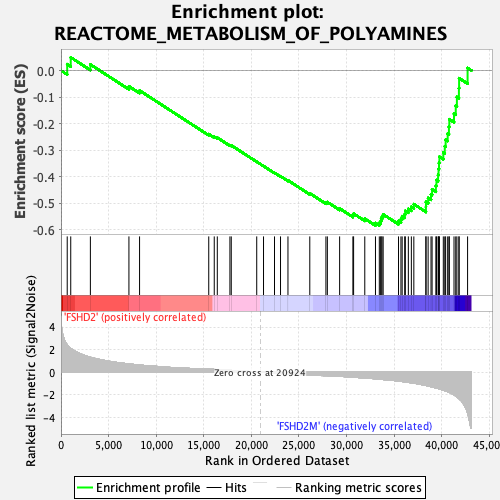
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_METABOLISM_OF_POLYAMINES |
| Enrichment Score (ES) | -0.5836973 |
| Normalized Enrichment Score (NES) | -1.996026 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0011378825 |
| FWER p-Value | 0.042 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMB9 | ENSG00000240065.8 | 653 | 2.409 | 0.0240 | No | ||
| 2 | PSMB8 | ENSG00000204264.11 | 1023 | 2.095 | 0.0495 | No | ||
| 3 | PSMB10 | ENSG00000205220.12 | 3086 | 1.313 | 0.0230 | No | ||
| 4 | PAOX | ENSG00000148832.16 | 7135 | 0.724 | -0.0593 | No | ||
| 5 | NQO1 | ENSG00000181019.13 | 8253 | 0.622 | -0.0751 | No | ||
| 6 | SAT1 | ENSG00000130066.16 | 15520 | 0.274 | -0.2394 | No | ||
| 7 | AGMAT | ENSG00000116771.6 | 16098 | 0.243 | -0.2488 | No | ||
| 8 | PSMA2 | ENSG00000106588.11 | 16418 | 0.224 | -0.2526 | No | ||
| 9 | PSME1 | ENSG00000092010.15 | 17744 | 0.161 | -0.2807 | No | ||
| 10 | SRM | ENSG00000116649.10 | 17885 | 0.152 | -0.2815 | No | ||
| 11 | PSMF1 | ENSG00000125818.18 | 20560 | 0.019 | -0.3433 | No | ||
| 12 | SEM1 | ENSG00000127922.9 | 21274 | -0.020 | -0.3595 | No | ||
| 13 | OAZ1 | ENSG00000104904.12 | 22428 | -0.081 | -0.3850 | No | ||
| 14 | PSMD9 | ENSG00000110801.14 | 23060 | -0.115 | -0.3978 | No | ||
| 15 | PSMD5 | ENSG00000095261.14 | 23841 | -0.161 | -0.4133 | No | ||
| 16 | PSMA8 | ENSG00000154611.14 | 26134 | -0.263 | -0.4622 | No | ||
| 17 | PSMB11 | ENSG00000222028.5 | 27828 | -0.302 | -0.4966 | No | ||
| 18 | OAZ3 | ENSG00000143450.16 | 27980 | -0.307 | -0.4952 | No | ||
| 19 | PSMB3 | ENSG00000277791.5 | 29269 | -0.361 | -0.5192 | No | ||
| 20 | PSMA1 | ENSG00000129084.17 | 30662 | -0.448 | -0.5443 | No | ||
| 21 | PSME3 | ENSG00000131467.11 | 30737 | -0.449 | -0.5387 | No | ||
| 22 | PSMA3 | ENSG00000100567.13 | 31911 | -0.509 | -0.5576 | No | ||
| 23 | PSMB1 | ENSG00000008018.9 | 33034 | -0.586 | -0.5742 | Yes | ||
| 24 | PSMD13 | ENSG00000185627.18 | 33437 | -0.616 | -0.5735 | Yes | ||
| 25 | PSMB4 | ENSG00000159377.11 | 33581 | -0.627 | -0.5666 | Yes | ||
| 26 | PSME2 | ENSG00000100911.16 | 33632 | -0.632 | -0.5575 | Yes | ||
| 27 | OAZ2 | ENSG00000180304.14 | 33726 | -0.639 | -0.5493 | Yes | ||
| 28 | PSMD10 | ENSG00000101843.19 | 33841 | -0.648 | -0.5414 | Yes | ||
| 29 | PSMC5 | ENSG00000087191.13 | 35455 | -0.783 | -0.5661 | Yes | ||
| 30 | PSMA7 | ENSG00000101182.15 | 35709 | -0.810 | -0.5588 | Yes | ||
| 31 | PSMB2 | ENSG00000126067.12 | 35837 | -0.822 | -0.5484 | Yes | ||
| 32 | AMD1 | ENSG00000123505.17 | 36105 | -0.856 | -0.5407 | Yes | ||
| 33 | PSMA5 | ENSG00000143106.13 | 36170 | -0.865 | -0.5281 | Yes | ||
| 34 | PSMC2 | ENSG00000161057.12 | 36487 | -0.899 | -0.5208 | Yes | ||
| 35 | PSMB7 | ENSG00000136930.13 | 36803 | -0.941 | -0.5128 | Yes | ||
| 36 | PSMB6 | ENSG00000142507.10 | 37064 | -0.976 | -0.5030 | Yes | ||
| 37 | PSMA4 | ENSG00000041357.16 | 38333 | -1.177 | -0.5133 | Yes | ||
| 38 | AZIN1 | ENSG00000155096.14 | 38338 | -1.177 | -0.4942 | Yes | ||
| 39 | PSMB5 | ENSG00000100804.19 | 38565 | -1.217 | -0.4797 | Yes | ||
| 40 | PSMD7 | ENSG00000103035.11 | 38868 | -1.282 | -0.4659 | Yes | ||
| 41 | SMS | ENSG00000102172.16 | 38982 | -1.305 | -0.4473 | Yes | ||
| 42 | SMOX | ENSG00000088826.18 | 39379 | -1.393 | -0.4338 | Yes | ||
| 43 | PSMD6 | ENSG00000163636.10 | 39440 | -1.407 | -0.4123 | Yes | ||
| 44 | PSMA6 | ENSG00000100902.11 | 39610 | -1.444 | -0.3928 | Yes | ||
| 45 | PSMD3 | ENSG00000108344.15 | 39660 | -1.456 | -0.3702 | Yes | ||
| 46 | PSMD4 | ENSG00000159352.15 | 39708 | -1.468 | -0.3474 | Yes | ||
| 47 | PSMC1 | ENSG00000100764.14 | 39751 | -1.479 | -0.3244 | Yes | ||
| 48 | PSMD12 | ENSG00000197170.10 | 40160 | -1.590 | -0.3080 | Yes | ||
| 49 | PSMD8 | ENSG00000099341.11 | 40314 | -1.631 | -0.2850 | Yes | ||
| 50 | PSMC4 | ENSG00000013275.8 | 40405 | -1.659 | -0.2601 | Yes | ||
| 51 | PSMD11 | ENSG00000108671.11 | 40619 | -1.724 | -0.2370 | Yes | ||
| 52 | PSMD14 | ENSG00000115233.12 | 40751 | -1.767 | -0.2113 | Yes | ||
| 53 | PSMC6 | ENSG00000100519.12 | 40791 | -1.783 | -0.1833 | Yes | ||
| 54 | PSMD1 | ENSG00000173692.13 | 41284 | -1.994 | -0.1623 | Yes | ||
| 55 | PSMC3 | ENSG00000165916.8 | 41467 | -2.112 | -0.1321 | Yes | ||
| 56 | AZIN2 | ENSG00000142920.17 | 41587 | -2.188 | -0.0993 | Yes | ||
| 57 | PSME4 | ENSG00000068878.15 | 41799 | -2.328 | -0.0664 | Yes | ||
| 58 | PSMD2 | ENSG00000175166.17 | 41820 | -2.343 | -0.0287 | Yes | ||
| 59 | ODC1 | ENSG00000115758.13 | 42710 | -3.622 | 0.0095 | Yes |