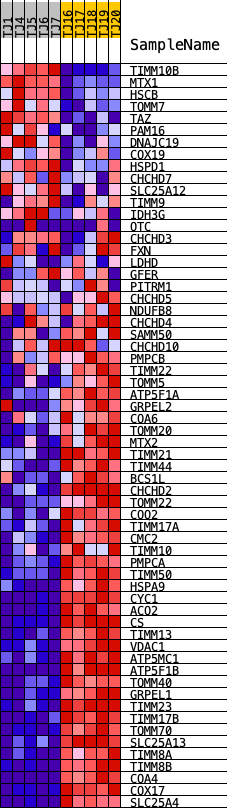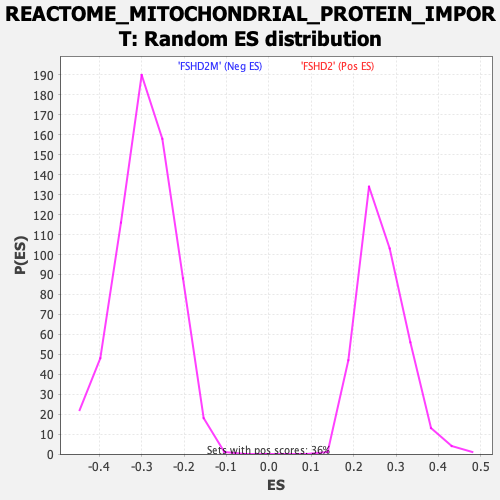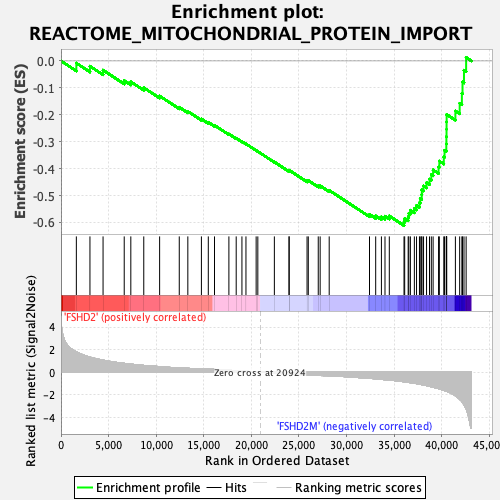
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_MITOCHONDRIAL_PROTEIN_IMPORT |
| Enrichment Score (ES) | -0.6118051 |
| Normalized Enrichment Score (NES) | -2.104097 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.149819E-4 |
| FWER p-Value | 0.006 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TIMM10B | ENSG00000132286.12 | 1612 | 1.795 | -0.0081 | No | ||
| 2 | MTX1 | ENSG00000173171.14 | 3041 | 1.323 | -0.0195 | No | ||
| 3 | HSCB | ENSG00000100209.10 | 4418 | 1.059 | -0.0342 | No | ||
| 4 | TOMM7 | ENSG00000196683.10 | 6641 | 0.774 | -0.0731 | No | ||
| 5 | TAZ | ENSG00000102125.16 | 7334 | 0.707 | -0.0776 | No | ||
| 6 | PAM16 | ENSG00000217930.8 | 8694 | 0.591 | -0.0995 | No | ||
| 7 | DNAJC19 | ENSG00000205981.8 | 10360 | 0.487 | -0.1302 | No | ||
| 8 | COX19 | ENSG00000240230.6 | 12423 | 0.368 | -0.1720 | No | ||
| 9 | HSPD1 | ENSG00000144381.17 | 13323 | 0.338 | -0.1874 | No | ||
| 10 | CHCHD7 | ENSG00000170791.18 | 14749 | 0.295 | -0.2157 | No | ||
| 11 | SLC25A12 | ENSG00000115840.14 | 15473 | 0.276 | -0.2279 | No | ||
| 12 | TIMM9 | ENSG00000100575.14 | 16125 | 0.241 | -0.2391 | No | ||
| 13 | IDH3G | ENSG00000067829.19 | 17636 | 0.167 | -0.2714 | No | ||
| 14 | OTC | ENSG00000036473.8 | 18404 | 0.129 | -0.2871 | No | ||
| 15 | CHCHD3 | ENSG00000106554.13 | 19006 | 0.098 | -0.2995 | No | ||
| 16 | FXN | ENSG00000165060.14 | 19429 | 0.075 | -0.3081 | No | ||
| 17 | LDHD | ENSG00000166816.14 | 20499 | 0.023 | -0.3325 | No | ||
| 18 | GFER | ENSG00000127554.13 | 20676 | 0.014 | -0.3364 | No | ||
| 19 | PITRM1 | ENSG00000107959.16 | 22413 | -0.080 | -0.3754 | No | ||
| 20 | CHCHD5 | ENSG00000125611.15 | 23958 | -0.169 | -0.4085 | No | ||
| 21 | NDUFB8 | ENSG00000166136.16 | 23962 | -0.169 | -0.4058 | No | ||
| 22 | CHCHD4 | ENSG00000163528.13 | 25846 | -0.250 | -0.4454 | No | ||
| 23 | SAMM50 | ENSG00000100347.15 | 25977 | -0.259 | -0.4442 | No | ||
| 24 | CHCHD10 | ENSG00000250479.8 | 27023 | -0.275 | -0.4640 | No | ||
| 25 | PMPCB | ENSG00000105819.14 | 27223 | -0.289 | -0.4639 | No | ||
| 26 | TIMM22 | ENSG00000177370.5 | 28173 | -0.321 | -0.4806 | No | ||
| 27 | TOMM5 | ENSG00000175768.13 | 32407 | -0.540 | -0.5701 | No | ||
| 28 | ATP5F1A | ENSG00000152234.16 | 33057 | -0.587 | -0.5756 | No | ||
| 29 | GRPEL2 | ENSG00000164284.15 | 33658 | -0.635 | -0.5791 | No | ||
| 30 | COA6 | ENSG00000168275.15 | 34034 | -0.663 | -0.5770 | No | ||
| 31 | TOMM20 | ENSG00000173726.11 | 34487 | -0.701 | -0.5760 | No | ||
| 32 | MTX2 | ENSG00000128654.14 | 36031 | -0.847 | -0.5979 | Yes | ||
| 33 | TIMM21 | ENSG00000075336.12 | 36103 | -0.856 | -0.5856 | Yes | ||
| 34 | TIMM44 | ENSG00000104980.8 | 36469 | -0.897 | -0.5794 | Yes | ||
| 35 | BCS1L | ENSG00000074582.14 | 36537 | -0.904 | -0.5661 | Yes | ||
| 36 | CHCHD2 | ENSG00000106153.13 | 36702 | -0.930 | -0.5547 | Yes | ||
| 37 | TOMM22 | ENSG00000100216.6 | 37119 | -0.982 | -0.5483 | Yes | ||
| 38 | COQ2 | ENSG00000173085.14 | 37348 | -1.014 | -0.5370 | Yes | ||
| 39 | TIMM17A | ENSG00000134375.11 | 37654 | -1.060 | -0.5267 | Yes | ||
| 40 | CMC2 | ENSG00000103121.8 | 37732 | -1.073 | -0.5109 | Yes | ||
| 41 | TIMM10 | ENSG00000134809.9 | 37894 | -1.100 | -0.4967 | Yes | ||
| 42 | PMPCA | ENSG00000165688.12 | 37905 | -1.101 | -0.4789 | Yes | ||
| 43 | TIMM50 | ENSG00000105197.11 | 38083 | -1.132 | -0.4644 | Yes | ||
| 44 | HSPA9 | ENSG00000113013.15 | 38400 | -1.187 | -0.4523 | Yes | ||
| 45 | CYC1 | ENSG00000179091.5 | 38710 | -1.245 | -0.4391 | Yes | ||
| 46 | ACO2 | ENSG00000100412.16 | 38900 | -1.290 | -0.4224 | Yes | ||
| 47 | CS | ENSG00000062485.19 | 39089 | -1.328 | -0.4050 | Yes | ||
| 48 | TIMM13 | ENSG00000099800.8 | 39661 | -1.456 | -0.3944 | Yes | ||
| 49 | VDAC1 | ENSG00000213585.11 | 39742 | -1.476 | -0.3721 | Yes | ||
| 50 | ATP5MC1 | ENSG00000159199.14 | 40212 | -1.603 | -0.3568 | Yes | ||
| 51 | ATP5F1B | ENSG00000110955.9 | 40290 | -1.624 | -0.3320 | Yes | ||
| 52 | TOMM40 | ENSG00000130204.13 | 40475 | -1.681 | -0.3087 | Yes | ||
| 53 | GRPEL1 | ENSG00000109519.13 | 40485 | -1.684 | -0.2814 | Yes | ||
| 54 | TIMM23 | ENSG00000265354.4 | 40498 | -1.687 | -0.2540 | Yes | ||
| 55 | TIMM17B | ENSG00000126768.12 | 40502 | -1.688 | -0.2264 | Yes | ||
| 56 | TOMM70 | ENSG00000154174.8 | 40512 | -1.693 | -0.1989 | Yes | ||
| 57 | SLC25A13 | ENSG00000004864.13 | 41429 | -2.086 | -0.1861 | Yes | ||
| 58 | TIMM8A | ENSG00000126953.8 | 41879 | -2.386 | -0.1574 | Yes | ||
| 59 | TIMM8B | ENSG00000150779.12 | 42111 | -2.587 | -0.1204 | Yes | ||
| 60 | COA4 | ENSG00000181924.7 | 42178 | -2.671 | -0.0782 | Yes | ||
| 61 | COX17 | ENSG00000138495.6 | 42332 | -2.886 | -0.0345 | Yes | ||
| 62 | SLC25A4 | ENSG00000151729.11 | 42557 | -3.224 | 0.0131 | Yes |