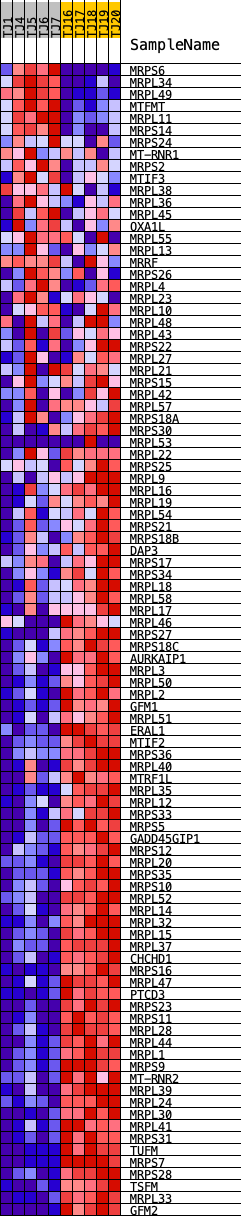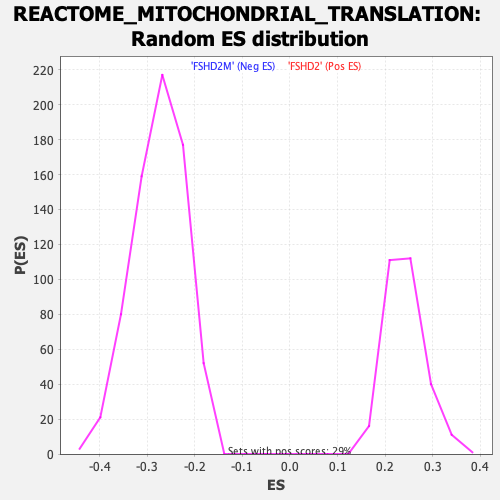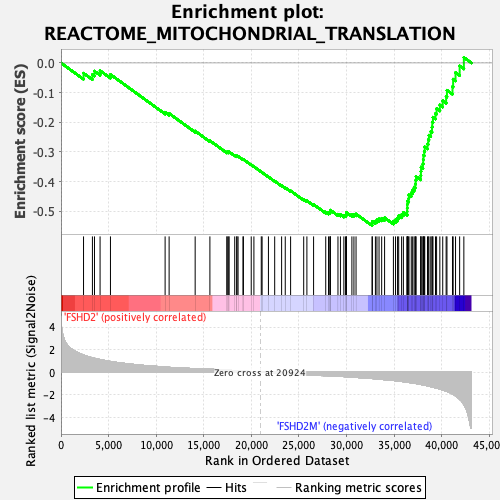
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_MITOCHONDRIAL_TRANSLATION |
| Enrichment Score (ES) | -0.5479773 |
| Normalized Enrichment Score (NES) | -1.9925709 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0012085217 |
| FWER p-Value | 0.047 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MRPS6 | ENSG00000243927.6 | 2367 | 1.515 | -0.0345 | No | ||
| 2 | MRPL34 | ENSG00000130312.6 | 3300 | 1.269 | -0.0390 | No | ||
| 3 | MRPL49 | ENSG00000149792.9 | 3516 | 1.222 | -0.0275 | No | ||
| 4 | MTFMT | ENSG00000103707.10 | 4105 | 1.111 | -0.0262 | No | ||
| 5 | MRPL11 | ENSG00000174547.13 | 5195 | 0.940 | -0.0388 | No | ||
| 6 | MRPS14 | ENSG00000120333.4 | 10936 | 0.449 | -0.1661 | No | ||
| 7 | MRPS24 | ENSG00000062582.14 | 11357 | 0.423 | -0.1701 | No | ||
| 8 | MT-RNR1 | ENSG00000211459.2 | 14093 | 0.311 | -0.2295 | No | ||
| 9 | MRPS2 | ENSG00000122140.11 | 15641 | 0.267 | -0.2618 | No | ||
| 10 | MTIF3 | ENSG00000122033.14 | 17419 | 0.176 | -0.3008 | No | ||
| 11 | MRPL38 | ENSG00000204316.12 | 17462 | 0.174 | -0.2994 | No | ||
| 12 | MRPL36 | ENSG00000171421.13 | 17624 | 0.168 | -0.3009 | No | ||
| 13 | MRPL45 | ENSG00000278845.5 | 17648 | 0.166 | -0.2991 | No | ||
| 14 | OXA1L | ENSG00000155463.14 | 18249 | 0.134 | -0.3113 | No | ||
| 15 | MRPL55 | ENSG00000162910.19 | 18456 | 0.127 | -0.3144 | No | ||
| 16 | MRPL13 | ENSG00000172172.8 | 18482 | 0.125 | -0.3132 | No | ||
| 17 | MRRF | ENSG00000148187.18 | 18600 | 0.119 | -0.3144 | No | ||
| 18 | MRPS26 | ENSG00000125901.6 | 19115 | 0.094 | -0.3250 | No | ||
| 19 | MRPL4 | ENSG00000105364.14 | 19163 | 0.091 | -0.3249 | No | ||
| 20 | MRPL23 | ENSG00000214026.11 | 19983 | 0.048 | -0.3433 | No | ||
| 21 | MRPL10 | ENSG00000159111.13 | 20264 | 0.034 | -0.3493 | No | ||
| 22 | MRPL48 | ENSG00000175581.14 | 21037 | -0.006 | -0.3672 | No | ||
| 23 | MRPL43 | ENSG00000055950.16 | 21124 | -0.012 | -0.3690 | No | ||
| 24 | MRPS22 | ENSG00000175110.11 | 21790 | -0.046 | -0.3839 | No | ||
| 25 | MRPL27 | ENSG00000108826.16 | 22447 | -0.082 | -0.3980 | No | ||
| 26 | MRPL21 | ENSG00000197345.13 | 23175 | -0.123 | -0.4132 | No | ||
| 27 | MRPS15 | ENSG00000116898.12 | 23557 | -0.144 | -0.4201 | No | ||
| 28 | MRPL42 | ENSG00000198015.13 | 24119 | -0.177 | -0.4308 | No | ||
| 29 | MRPL57 | ENSG00000173141.5 | 25494 | -0.227 | -0.4597 | No | ||
| 30 | MRPS18A | ENSG00000096080.11 | 25830 | -0.249 | -0.4641 | No | ||
| 31 | MRPS30 | ENSG00000112996.11 | 26533 | -0.269 | -0.4768 | No | ||
| 32 | MRPL53 | ENSG00000204822.7 | 27809 | -0.302 | -0.5023 | No | ||
| 33 | MRPL22 | ENSG00000082515.18 | 28103 | -0.317 | -0.5048 | No | ||
| 34 | MRPS25 | ENSG00000131368.8 | 28243 | -0.326 | -0.5037 | No | ||
| 35 | MRPL9 | ENSG00000143436.11 | 28288 | -0.329 | -0.5002 | No | ||
| 36 | MRPL16 | ENSG00000166902.5 | 28296 | -0.329 | -0.4959 | No | ||
| 37 | MRPL19 | ENSG00000115364.14 | 29094 | -0.353 | -0.5097 | No | ||
| 38 | MRPL54 | ENSG00000183617.5 | 29353 | -0.366 | -0.5107 | No | ||
| 39 | MRPS21 | ENSG00000266472.5 | 29713 | -0.387 | -0.5139 | No | ||
| 40 | MRPS18B | ENSG00000204568.12 | 29892 | -0.398 | -0.5126 | No | ||
| 41 | DAP3 | ENSG00000132676.16 | 29977 | -0.405 | -0.5091 | No | ||
| 42 | MRPS17 | ENSG00000239789.6 | 29979 | -0.405 | -0.5036 | No | ||
| 43 | MRPS34 | ENSG00000074071.14 | 30557 | -0.438 | -0.5111 | No | ||
| 44 | MRPL18 | ENSG00000112110.10 | 30771 | -0.452 | -0.5100 | No | ||
| 45 | MRPL58 | ENSG00000167862.10 | 30993 | -0.458 | -0.5089 | No | ||
| 46 | MRPL17 | ENSG00000158042.9 | 32675 | -0.558 | -0.5404 | Yes | ||
| 47 | MRPL46 | ENSG00000259494.2 | 32709 | -0.561 | -0.5336 | Yes | ||
| 48 | MRPS27 | ENSG00000113048.16 | 33027 | -0.585 | -0.5331 | Yes | ||
| 49 | MRPS18C | ENSG00000163319.11 | 33187 | -0.597 | -0.5287 | Yes | ||
| 50 | AURKAIP1 | ENSG00000175756.13 | 33409 | -0.613 | -0.5255 | Yes | ||
| 51 | MRPL3 | ENSG00000114686.9 | 33685 | -0.637 | -0.5233 | Yes | ||
| 52 | MRPL50 | ENSG00000136897.8 | 33992 | -0.661 | -0.5215 | Yes | ||
| 53 | MRPL2 | ENSG00000112651.12 | 34918 | -0.735 | -0.5331 | Yes | ||
| 54 | GFM1 | ENSG00000168827.14 | 35142 | -0.754 | -0.5280 | Yes | ||
| 55 | MRPL51 | ENSG00000111639.8 | 35355 | -0.774 | -0.5225 | Yes | ||
| 56 | ERAL1 | ENSG00000132591.12 | 35473 | -0.785 | -0.5146 | Yes | ||
| 57 | MTIF2 | ENSG00000085760.15 | 35779 | -0.816 | -0.5107 | Yes | ||
| 58 | MRPS36 | ENSG00000134056.12 | 35966 | -0.839 | -0.5037 | Yes | ||
| 59 | MRPL40 | ENSG00000185608.8 | 36363 | -0.884 | -0.5009 | Yes | ||
| 60 | MTRF1L | ENSG00000112031.16 | 36368 | -0.885 | -0.4891 | Yes | ||
| 61 | MRPL35 | ENSG00000132313.15 | 36377 | -0.885 | -0.4773 | Yes | ||
| 62 | MRPL12 | ENSG00000262814.8 | 36384 | -0.886 | -0.4654 | Yes | ||
| 63 | MRPS33 | ENSG00000090263.16 | 36529 | -0.904 | -0.4566 | Yes | ||
| 64 | MRPS5 | ENSG00000144029.12 | 36539 | -0.904 | -0.4445 | Yes | ||
| 65 | GADD45GIP1 | ENSG00000179271.3 | 36793 | -0.940 | -0.4377 | Yes | ||
| 66 | MRPS12 | ENSG00000128626.11 | 36940 | -0.958 | -0.4282 | Yes | ||
| 67 | MRPL20 | ENSG00000242485.6 | 37096 | -0.979 | -0.4185 | Yes | ||
| 68 | MRPS35 | ENSG00000061794.13 | 37216 | -0.994 | -0.4079 | Yes | ||
| 69 | MRPS10 | ENSG00000048544.6 | 37256 | -1.000 | -0.3952 | Yes | ||
| 70 | MRPL52 | ENSG00000172590.18 | 37298 | -1.007 | -0.3826 | Yes | ||
| 71 | MRPL14 | ENSG00000180992.7 | 37789 | -1.082 | -0.3793 | Yes | ||
| 72 | MRPL32 | ENSG00000106591.4 | 37806 | -1.084 | -0.3651 | Yes | ||
| 73 | MRPL15 | ENSG00000137547.9 | 37824 | -1.087 | -0.3508 | Yes | ||
| 74 | MRPL37 | ENSG00000116221.15 | 38018 | -1.120 | -0.3401 | Yes | ||
| 75 | CHCHD1 | ENSG00000172586.8 | 38051 | -1.125 | -0.3256 | Yes | ||
| 76 | MRPS16 | ENSG00000182180.14 | 38073 | -1.129 | -0.3109 | Yes | ||
| 77 | MRPL47 | ENSG00000136522.14 | 38164 | -1.147 | -0.2974 | Yes | ||
| 78 | PTCD3 | ENSG00000132300.19 | 38208 | -1.154 | -0.2828 | Yes | ||
| 79 | MRPS23 | ENSG00000181610.13 | 38502 | -1.206 | -0.2733 | Yes | ||
| 80 | MRPS11 | ENSG00000181991.16 | 38580 | -1.220 | -0.2586 | Yes | ||
| 81 | MRPL28 | ENSG00000086504.17 | 38649 | -1.233 | -0.2436 | Yes | ||
| 82 | MRPL44 | ENSG00000135900.4 | 38843 | -1.277 | -0.2308 | Yes | ||
| 83 | MRPL1 | ENSG00000169288.18 | 38971 | -1.303 | -0.2161 | Yes | ||
| 84 | MRPS9 | ENSG00000135972.9 | 39002 | -1.311 | -0.1991 | Yes | ||
| 85 | MT-RNR2 | ENSG00000210082.2 | 39074 | -1.325 | -0.1828 | Yes | ||
| 86 | MRPL39 | ENSG00000154719.13 | 39339 | -1.385 | -0.1702 | Yes | ||
| 87 | MRPL24 | ENSG00000143314.12 | 39454 | -1.410 | -0.1538 | Yes | ||
| 88 | MRPL30 | ENSG00000185414.20 | 39796 | -1.492 | -0.1416 | Yes | ||
| 89 | MRPL41 | ENSG00000182154.8 | 40099 | -1.570 | -0.1274 | Yes | ||
| 90 | MRPS31 | ENSG00000102738.8 | 40457 | -1.673 | -0.1130 | Yes | ||
| 91 | TUFM | ENSG00000178952.11 | 40543 | -1.700 | -0.0920 | Yes | ||
| 92 | MRPS7 | ENSG00000125445.11 | 41124 | -1.920 | -0.0796 | Yes | ||
| 93 | MRPS28 | ENSG00000147586.10 | 41192 | -1.955 | -0.0547 | Yes | ||
| 94 | TSFM | ENSG00000123297.19 | 41445 | -2.098 | -0.0322 | Yes | ||
| 95 | MRPL33 | ENSG00000243147.8 | 41876 | -2.384 | -0.0099 | Yes | ||
| 96 | GFM2 | ENSG00000164347.18 | 42318 | -2.870 | 0.0186 | Yes |