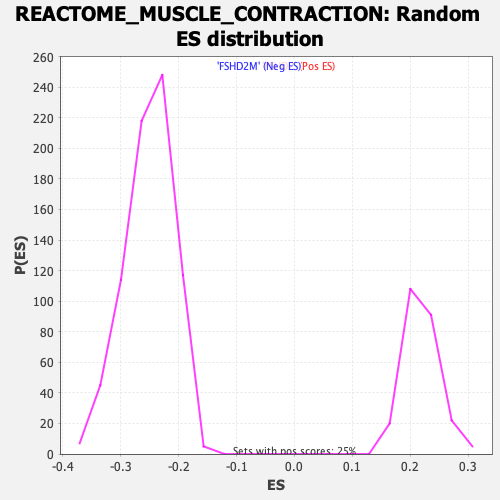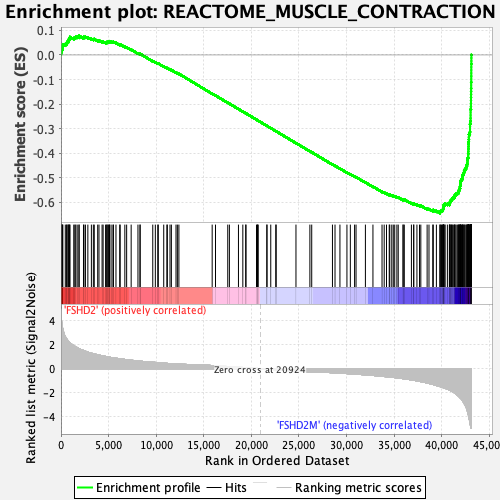
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | -0.6434019 |
| Normalized Enrichment Score (NES) | -2.5713985 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CLIC2 | ENSG00000155962.13 | 4 | 4.773 | 0.0119 | No | ||
| 2 | KCNIP3 | ENSG00000115041.13 | 68 | 3.944 | 0.0203 | No | ||
| 3 | CACNA2D4 | ENSG00000151062.15 | 124 | 3.591 | 0.0280 | No | ||
| 4 | TBX5 | ENSG00000089225.19 | 139 | 3.525 | 0.0365 | No | ||
| 5 | CACNA1C | ENSG00000151067.22 | 193 | 3.311 | 0.0435 | No | ||
| 6 | LMOD1 | ENSG00000163431.13 | 451 | 2.677 | 0.0442 | No | ||
| 7 | ASPH | ENSG00000198363.18 | 590 | 2.483 | 0.0472 | No | ||
| 8 | ATP2B4 | ENSG00000058668.14 | 627 | 2.432 | 0.0525 | No | ||
| 9 | KCNK2 | ENSG00000082482.13 | 754 | 2.305 | 0.0553 | No | ||
| 10 | KCND1 | ENSG00000102057.10 | 763 | 2.298 | 0.0609 | No | ||
| 11 | VIM | ENSG00000026025.16 | 852 | 2.214 | 0.0644 | No | ||
| 12 | SORBS3 | ENSG00000120896.13 | 884 | 2.183 | 0.0691 | No | ||
| 13 | CACNB3 | ENSG00000167535.8 | 947 | 2.135 | 0.0730 | No | ||
| 14 | KCND3 | ENSG00000171385.9 | 1341 | 1.914 | 0.0686 | No | ||
| 15 | NPR2 | ENSG00000159899.14 | 1451 | 1.865 | 0.0708 | No | ||
| 16 | KCNK15 | ENSG00000124249.7 | 1548 | 1.821 | 0.0731 | No | ||
| 17 | TMOD3 | ENSG00000138594.14 | 1698 | 1.756 | 0.0740 | No | ||
| 18 | ORAI2 | ENSG00000160991.16 | 1886 | 1.673 | 0.0738 | No | ||
| 19 | SCN1A | ENSG00000144285.21 | 1902 | 1.668 | 0.0777 | No | ||
| 20 | KCNIP4 | ENSG00000185774.16 | 2362 | 1.517 | 0.0708 | No | ||
| 21 | CACNA1D | ENSG00000157388.19 | 2414 | 1.501 | 0.0733 | No | ||
| 22 | SCN4B | ENSG00000177098.8 | 2566 | 1.451 | 0.0734 | No | ||
| 23 | KCNK12 | ENSG00000184261.4 | 2821 | 1.373 | 0.0710 | No | ||
| 24 | MYL7 | ENSG00000106631.8 | 3191 | 1.291 | 0.0656 | No | ||
| 25 | GUCY1A1 | ENSG00000164116.16 | 3397 | 1.246 | 0.0639 | No | ||
| 26 | KCNE3 | ENSG00000175538.10 | 3493 | 1.226 | 0.0648 | No | ||
| 27 | CACNA2D2 | ENSG00000007402.11 | 3855 | 1.154 | 0.0593 | No | ||
| 28 | SCN2A | ENSG00000136531.16 | 3989 | 1.130 | 0.0590 | No | ||
| 29 | ITGB5 | ENSG00000082781.12 | 4315 | 1.074 | 0.0541 | No | ||
| 30 | NPR1 | ENSG00000169418.10 | 4423 | 1.058 | 0.0543 | No | ||
| 31 | GUCY1B1 | ENSG00000061918.13 | 4682 | 1.017 | 0.0508 | No | ||
| 32 | CACNA2D3 | ENSG00000157445.15 | 4802 | 0.997 | 0.0505 | No | ||
| 33 | FGF12 | ENSG00000114279.13 | 4831 | 0.993 | 0.0524 | No | ||
| 34 | FGF14 | ENSG00000102466.15 | 4922 | 0.980 | 0.0527 | No | ||
| 35 | CACNG7 | ENSG00000105605.7 | 5004 | 0.967 | 0.0532 | No | ||
| 36 | SCN7A | ENSG00000136546.16 | 5073 | 0.957 | 0.0541 | No | ||
| 37 | KCND2 | ENSG00000184408.10 | 5100 | 0.954 | 0.0558 | No | ||
| 38 | CACNB2 | ENSG00000165995.20 | 5260 | 0.933 | 0.0545 | No | ||
| 39 | TMOD2 | ENSG00000128872.10 | 5454 | 0.906 | 0.0522 | No | ||
| 40 | GUCY1A2 | ENSG00000152402.10 | 5511 | 0.899 | 0.0532 | No | ||
| 41 | TRPC1 | ENSG00000144935.15 | 5774 | 0.867 | 0.0492 | No | ||
| 42 | KCNK4 | ENSG00000182450.13 | 6171 | 0.820 | 0.0421 | No | ||
| 43 | RANGRF | ENSG00000108961.14 | 6214 | 0.815 | 0.0431 | No | ||
| 44 | SRI | ENSG00000075142.13 | 6706 | 0.767 | 0.0336 | No | ||
| 45 | CACNG2 | ENSG00000166862.6 | 6896 | 0.747 | 0.0311 | No | ||
| 46 | CACNA1F | ENSG00000102001.12 | 7370 | 0.703 | 0.0218 | No | ||
| 47 | CACNB4 | ENSG00000182389.19 | 8084 | 0.640 | 0.0068 | No | ||
| 48 | FXYD7 | ENSG00000221946.7 | 8301 | 0.620 | 0.0033 | No | ||
| 49 | SCN2B | ENSG00000149575.6 | 8311 | 0.619 | 0.0047 | No | ||
| 50 | KCNK6 | ENSG00000099337.5 | 9639 | 0.527 | -0.0249 | No | ||
| 51 | ANXA2 | ENSG00000182718.16 | 9909 | 0.516 | -0.0299 | No | ||
| 52 | KCNE2 | ENSG00000159197.3 | 10168 | 0.499 | -0.0347 | No | ||
| 53 | KCNK1 | ENSG00000135750.15 | 10219 | 0.496 | -0.0346 | No | ||
| 54 | KCNIP1 | ENSG00000182132.13 | 10791 | 0.460 | -0.0468 | No | ||
| 55 | TPM4 | ENSG00000167460.17 | 11125 | 0.437 | -0.0534 | No | ||
| 56 | ITGA1 | ENSG00000213949.10 | 11169 | 0.434 | -0.0533 | No | ||
| 57 | KAT2B | ENSG00000114166.8 | 11450 | 0.417 | -0.0588 | No | ||
| 58 | KCNK9 | ENSG00000169427.8 | 11595 | 0.408 | -0.0612 | No | ||
| 59 | SCN1B | ENSG00000105711.12 | 12081 | 0.383 | -0.0715 | No | ||
| 60 | AHCYL1 | ENSG00000168710.18 | 12224 | 0.376 | -0.0739 | No | ||
| 61 | ANXA1 | ENSG00000135046.14 | 12384 | 0.370 | -0.0766 | No | ||
| 62 | ORAI1 | ENSG00000276045.3 | 15880 | 0.255 | -0.1574 | No | ||
| 63 | ATP1B2 | ENSG00000129244.9 | 16230 | 0.234 | -0.1650 | No | ||
| 64 | ABCC9 | ENSG00000069431.11 | 17517 | 0.171 | -0.1945 | No | ||
| 65 | SCN11A | ENSG00000168356.12 | 17688 | 0.164 | -0.1981 | No | ||
| 66 | ITPR3 | ENSG00000096433.11 | 18641 | 0.116 | -0.2200 | No | ||
| 67 | SCN8A | ENSG00000196876.17 | 19100 | 0.095 | -0.2304 | No | ||
| 68 | KCNE4 | ENSG00000152049.6 | 19402 | 0.077 | -0.2372 | No | ||
| 69 | MYLK | ENSG00000065534.18 | 19434 | 0.074 | -0.2378 | No | ||
| 70 | KCNK13 | ENSG00000152315.5 | 20537 | 0.020 | -0.2634 | No | ||
| 71 | KCNJ14 | ENSG00000182324.6 | 20632 | 0.015 | -0.2655 | No | ||
| 72 | ATP1A3 | ENSG00000105409.19 | 20648 | 0.014 | -0.2659 | No | ||
| 73 | MYH11 | ENSG00000133392.18 | 20718 | 0.012 | -0.2674 | No | ||
| 74 | CORIN | ENSG00000145244.12 | 21624 | -0.038 | -0.2884 | No | ||
| 75 | CACNG8 | ENSG00000142408.4 | 21638 | -0.039 | -0.2886 | No | ||
| 76 | ATP2A3 | ENSG00000074370.18 | 22034 | -0.059 | -0.2977 | No | ||
| 77 | FKBP1B | ENSG00000119782.14 | 22563 | -0.088 | -0.3098 | No | ||
| 78 | GATA4 | ENSG00000136574.17 | 22602 | -0.091 | -0.3104 | No | ||
| 79 | KCNK10 | ENSG00000100433.15 | 24682 | -0.210 | -0.3583 | No | ||
| 80 | CACNG3 | ENSG00000006116.3 | 26144 | -0.263 | -0.3917 | No | ||
| 81 | KCNK16 | ENSG00000095981.10 | 26337 | -0.263 | -0.3955 | No | ||
| 82 | TLN1 | ENSG00000137076.21 | 28515 | -0.337 | -0.4454 | No | ||
| 83 | RYR2 | ENSG00000198626.17 | 28784 | -0.351 | -0.4508 | No | ||
| 84 | NPPC | ENSG00000163273.4 | 29301 | -0.362 | -0.4619 | No | ||
| 85 | CAMK2D | ENSG00000145349.17 | 30036 | -0.410 | -0.4780 | No | ||
| 86 | SCN9A | ENSG00000169432.16 | 30394 | -0.431 | -0.4852 | No | ||
| 87 | PAK1 | ENSG00000149269.9 | 30840 | -0.456 | -0.4945 | No | ||
| 88 | FXYD2 | ENSG00000137731.14 | 30992 | -0.458 | -0.4968 | No | ||
| 89 | FGF13 | ENSG00000129682.16 | 31976 | -0.511 | -0.5185 | No | ||
| 90 | KCNJ2 | ENSG00000123700.4 | 32768 | -0.566 | -0.5355 | No | ||
| 91 | SLC8A2 | ENSG00000118160.14 | 33723 | -0.639 | -0.5561 | No | ||
| 92 | CALD1 | ENSG00000122786.20 | 33950 | -0.658 | -0.5597 | No | ||
| 93 | CAMK2G | ENSG00000148660.20 | 34193 | -0.675 | -0.5637 | No | ||
| 94 | PAK2 | ENSG00000180370.10 | 34479 | -0.701 | -0.5685 | No | ||
| 95 | KCNK5 | ENSG00000164626.9 | 34538 | -0.705 | -0.5681 | No | ||
| 96 | STIM1 | ENSG00000167323.11 | 34747 | -0.720 | -0.5712 | No | ||
| 97 | FXYD3 | ENSG00000089356.18 | 34904 | -0.733 | -0.5730 | No | ||
| 98 | MYL6 | ENSG00000092841.18 | 35056 | -0.747 | -0.5746 | No | ||
| 99 | NOS1 | ENSG00000089250.19 | 35284 | -0.766 | -0.5780 | No | ||
| 100 | MYBPC1 | ENSG00000196091.14 | 35425 | -0.780 | -0.5793 | No | ||
| 101 | KCNK17 | ENSG00000124780.14 | 35928 | -0.835 | -0.5889 | No | ||
| 102 | KCNK3 | ENSG00000171303.7 | 35941 | -0.836 | -0.5871 | No | ||
| 103 | KCNIP2 | ENSG00000120049.19 | 36062 | -0.851 | -0.5878 | No | ||
| 104 | MYL5 | ENSG00000215375.6 | 36809 | -0.942 | -0.6028 | No | ||
| 105 | ATP1B3 | ENSG00000069849.11 | 37044 | -0.973 | -0.6058 | No | ||
| 106 | KCNQ1 | ENSG00000053918.17 | 37057 | -0.975 | -0.6037 | No | ||
| 107 | PRKACA | ENSG00000072062.14 | 37389 | -1.018 | -0.6088 | No | ||
| 108 | VCL | ENSG00000035403.17 | 37662 | -1.061 | -0.6125 | No | ||
| 109 | MYL12B | ENSG00000118680.13 | 37797 | -1.083 | -0.6129 | No | ||
| 110 | ATP2B3 | ENSG00000067842.17 | 38460 | -1.199 | -0.6253 | No | ||
| 111 | MYBPC2 | ENSG00000086967.10 | 38655 | -1.234 | -0.6268 | No | ||
| 112 | TRDN | ENSG00000186439.14 | 39054 | -1.322 | -0.6327 | No | ||
| 113 | ACTN3 | ENSG00000248746.6 | 39105 | -1.332 | -0.6306 | No | ||
| 114 | HIPK1 | ENSG00000163349.22 | 39433 | -1.406 | -0.6347 | No | ||
| 115 | MYL3 | ENSG00000160808.10 | 39809 | -1.495 | -0.6397 | Yes | ||
| 116 | FXYD4 | ENSG00000150201.15 | 39814 | -1.497 | -0.6360 | Yes | ||
| 117 | MYL1 | ENSG00000168530.16 | 39896 | -1.521 | -0.6341 | Yes | ||
| 118 | NKX2-5 | ENSG00000183072.10 | 39974 | -1.542 | -0.6320 | Yes | ||
| 119 | KCNJ4 | ENSG00000168135.4 | 40091 | -1.569 | -0.6308 | Yes | ||
| 120 | CAMK2B | ENSG00000058404.20 | 40117 | -1.578 | -0.6274 | Yes | ||
| 121 | FXYD1 | ENSG00000266964.5 | 40154 | -1.588 | -0.6243 | Yes | ||
| 122 | ATP2B2 | ENSG00000157087.20 | 40162 | -1.590 | -0.6205 | Yes | ||
| 123 | ATP2B1 | ENSG00000070961.15 | 40174 | -1.593 | -0.6167 | Yes | ||
| 124 | MYL9 | ENSG00000101335.10 | 40187 | -1.596 | -0.6130 | Yes | ||
| 125 | CACNG4 | ENSG00000075461.6 | 40209 | -1.603 | -0.6095 | Yes | ||
| 126 | KCNE5 | ENSG00000176076.7 | 40321 | -1.633 | -0.6080 | Yes | ||
| 127 | SCN3A | ENSG00000153253.18 | 40335 | -1.638 | -0.6042 | Yes | ||
| 128 | ANXA6 | ENSG00000197043.14 | 40587 | -1.711 | -0.6058 | Yes | ||
| 129 | MYL12A | ENSG00000101608.12 | 40797 | -1.785 | -0.6062 | Yes | ||
| 130 | WWTR1 | ENSG00000018408.14 | 40806 | -1.787 | -0.6019 | Yes | ||
| 131 | KCNE1 | ENSG00000180509.12 | 40863 | -1.808 | -0.5987 | Yes | ||
| 132 | PLN | ENSG00000198523.6 | 40873 | -1.812 | -0.5944 | Yes | ||
| 133 | TNNI2 | ENSG00000130598.16 | 40957 | -1.846 | -0.5917 | Yes | ||
| 134 | ITPR1 | ENSG00000150995.19 | 41025 | -1.877 | -0.5885 | Yes | ||
| 135 | MYL10 | ENSG00000106436.7 | 41072 | -1.896 | -0.5849 | Yes | ||
| 136 | SCN10A | ENSG00000185313.8 | 41147 | -1.932 | -0.5817 | Yes | ||
| 137 | TNNI3 | ENSG00000129991.13 | 41255 | -1.980 | -0.5793 | Yes | ||
| 138 | ATP1A1 | ENSG00000163399.16 | 41345 | -2.032 | -0.5763 | Yes | ||
| 139 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.5716 | Yes | ||
| 140 | SCN3B | ENSG00000166257.9 | 41392 | -2.060 | -0.5671 | Yes | ||
| 141 | TPM3 | ENSG00000143549.21 | 41507 | -2.135 | -0.5644 | Yes | ||
| 142 | FXYD6 | ENSG00000137726.17 | 41634 | -2.212 | -0.5618 | Yes | ||
| 143 | ACTG2 | ENSG00000163017.14 | 41741 | -2.285 | -0.5585 | Yes | ||
| 144 | ATP2A1 | ENSG00000196296.13 | 41742 | -2.285 | -0.5528 | Yes | ||
| 145 | AKAP9 | ENSG00000127914.16 | 41818 | -2.342 | -0.5487 | Yes | ||
| 146 | ITPR2 | ENSG00000123104.12 | 41853 | -2.368 | -0.5436 | Yes | ||
| 147 | CACNG6 | ENSG00000130433.7 | 41917 | -2.421 | -0.5390 | Yes | ||
| 148 | SLC8A1 | ENSG00000183023.18 | 41918 | -2.421 | -0.5329 | Yes | ||
| 149 | CACNA2D1 | ENSG00000153956.16 | 41945 | -2.451 | -0.5274 | Yes | ||
| 150 | MYH3 | ENSG00000109063.15 | 41957 | -2.460 | -0.5215 | Yes | ||
| 151 | KCNH2 | ENSG00000055118.15 | 41985 | -2.477 | -0.5159 | Yes | ||
| 152 | RYR3 | ENSG00000198838.13 | 41998 | -2.488 | -0.5100 | Yes | ||
| 153 | CASQ1 | ENSG00000143318.13 | 42074 | -2.550 | -0.5053 | Yes | ||
| 154 | KCNJ12 | ENSG00000184185.10 | 42159 | -2.647 | -0.5007 | Yes | ||
| 155 | ATP2A2 | ENSG00000174437.17 | 42170 | -2.663 | -0.4942 | Yes | ||
| 156 | NPPA | ENSG00000175206.10 | 42186 | -2.680 | -0.4879 | Yes | ||
| 157 | ACTA2 | ENSG00000107796.13 | 42259 | -2.777 | -0.4826 | Yes | ||
| 158 | CALM1 | ENSG00000198668.12 | 42323 | -2.877 | -0.4769 | Yes | ||
| 159 | ATP1A4 | ENSG00000132681.16 | 42353 | -2.919 | -0.4702 | Yes | ||
| 160 | TNNC2 | ENSG00000101470.10 | 42424 | -3.025 | -0.4643 | Yes | ||
| 161 | MYH8 | ENSG00000133020.4 | 42537 | -3.185 | -0.4589 | Yes | ||
| 162 | HIPK2 | ENSG00000064393.16 | 42572 | -3.249 | -0.4516 | Yes | ||
| 163 | ATP1B1 | ENSG00000143153.13 | 42642 | -3.447 | -0.4446 | Yes | ||
| 164 | TTN | ENSG00000155657.26 | 42690 | -3.572 | -0.4367 | Yes | ||
| 165 | KCNK7 | ENSG00000173338.12 | 42693 | -3.578 | -0.4278 | Yes | ||
| 166 | ACTA1 | ENSG00000143632.14 | 42721 | -3.653 | -0.4193 | Yes | ||
| 167 | TMOD1 | ENSG00000136842.14 | 42776 | -3.811 | -0.4110 | Yes | ||
| 168 | CAMK2A | ENSG00000070808.16 | 42777 | -3.811 | -0.4015 | Yes | ||
| 169 | TPM1 | ENSG00000140416.21 | 42794 | -3.877 | -0.3922 | Yes | ||
| 170 | DMPK | ENSG00000104936.17 | 42797 | -3.886 | -0.3825 | Yes | ||
| 171 | CACNB1 | ENSG00000067191.16 | 42803 | -3.923 | -0.3728 | Yes | ||
| 172 | TNNT3 | ENSG00000130595.19 | 42808 | -3.940 | -0.3630 | Yes | ||
| 173 | CASQ2 | ENSG00000118729.11 | 42823 | -4.006 | -0.3533 | Yes | ||
| 174 | SLN | ENSG00000170290.4 | 42839 | -4.058 | -0.3435 | Yes | ||
| 175 | FGF11 | ENSG00000161958.11 | 42850 | -4.106 | -0.3335 | Yes | ||
| 176 | TPM2 | ENSG00000198467.15 | 42854 | -4.136 | -0.3232 | Yes | ||
| 177 | DMD | ENSG00000198947.15 | 42943 | -4.488 | -0.3140 | Yes | ||
| 178 | ACTN2 | ENSG00000077522.13 | 42966 | -4.553 | -0.3031 | Yes | ||
| 179 | MYLPF | ENSG00000180209.12 | 42967 | -4.555 | -0.2917 | Yes | ||
| 180 | MYL6B | ENSG00000196465.10 | 42969 | -4.580 | -0.2803 | Yes | ||
| 181 | CAV3 | ENSG00000182533.6 | 43009 | -4.750 | -0.2693 | Yes | ||
| 182 | SLC8A3 | ENSG00000100678.18 | 43023 | -4.807 | -0.2576 | Yes | ||
| 183 | MYL2 | ENSG00000111245.15 | 43025 | -4.822 | -0.2455 | Yes | ||
| 184 | TCAP | ENSG00000173991.5 | 43026 | -4.827 | -0.2334 | Yes | ||
| 185 | KCNJ11 | ENSG00000187486.5 | 43027 | -4.829 | -0.2214 | Yes | ||
| 186 | SORBS1 | ENSG00000095637.22 | 43061 | -4.908 | -0.2098 | Yes | ||
| 187 | SCN5A | ENSG00000183873.15 | 43064 | -4.912 | -0.1976 | Yes | ||
| 188 | ATP1A2 | ENSG00000018625.15 | 43067 | -4.924 | -0.1853 | Yes | ||
| 189 | MYL4 | ENSG00000198336.9 | 43070 | -4.932 | -0.1730 | Yes | ||
| 190 | RYR1 | ENSG00000196218.12 | 43074 | -4.942 | -0.1607 | Yes | ||
| 191 | CACNG1 | ENSG00000108878.5 | 43077 | -4.944 | -0.1484 | Yes | ||
| 192 | DYSF | ENSG00000135636.14 | 43081 | -4.948 | -0.1361 | Yes | ||
| 193 | DES | ENSG00000175084.12 | 43086 | -4.954 | -0.1238 | Yes | ||
| 194 | TNNT1 | ENSG00000105048.17 | 43087 | -4.955 | -0.1114 | Yes | ||
| 195 | SCN4A | ENSG00000007314.12 | 43094 | -4.963 | -0.0991 | Yes | ||
| 196 | TRIM72 | ENSG00000177238.14 | 43095 | -4.964 | -0.0867 | Yes | ||
| 197 | NEB | ENSG00000183091.19 | 43097 | -4.966 | -0.0743 | Yes | ||
| 198 | TNNT2 | ENSG00000118194.20 | 43098 | -4.967 | -0.0618 | Yes | ||
| 199 | MYH6 | ENSG00000197616.12 | 43099 | -4.967 | -0.0494 | Yes | ||
| 200 | TNNC1 | ENSG00000114854.8 | 43100 | -4.968 | -0.0370 | Yes | ||
| 201 | CACNA1S | ENSG00000081248.11 | 43107 | -4.973 | -0.0247 | Yes | ||
| 202 | ACTC1 | ENSG00000159251.8 | 43108 | -4.973 | -0.0122 | Yes | ||
| 203 | TNNI1 | ENSG00000159173.19 | 43109 | -4.974 | 0.0002 | Yes |