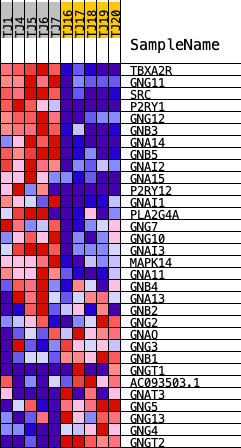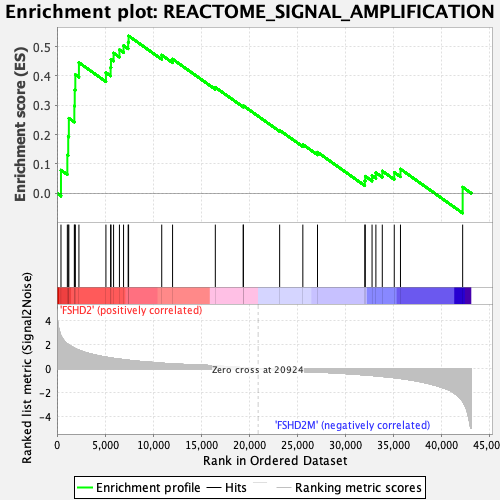
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_SIGNAL_AMPLIFICATION |
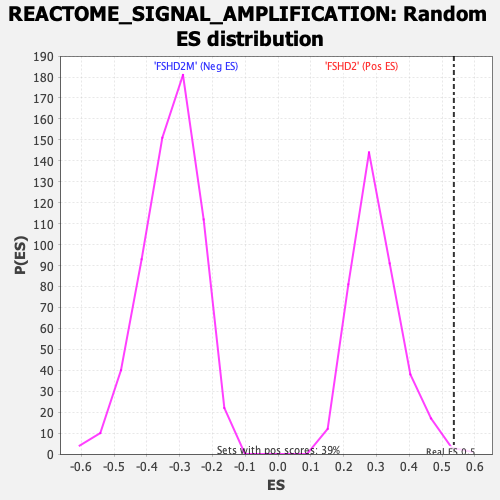
| Enrichment Score (ES) | 0.5361695 |
| Normalized Enrichment Score (NES) | 1.7902458 |
| Nominal p-value | 0.0025839794 |
| FDR q-value | 0.03680809 |
| FWER p-Value | 0.769 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TBXA2R | ENSG00000006638.11 | 414 | 2.734 | 0.0789 | Yes | ||
| 2 | GNG11 | ENSG00000127920.6 | 1076 | 2.064 | 0.1303 | Yes | ||
| 3 | SRC | ENSG00000197122.11 | 1168 | 2.008 | 0.1932 | Yes | ||
| 4 | P2RY1 | ENSG00000169860.7 | 1233 | 1.973 | 0.2555 | Yes | ||
| 5 | GNG12 | ENSG00000172380.6 | 1804 | 1.714 | 0.2978 | Yes | ||
| 6 | GNB3 | ENSG00000111664.10 | 1846 | 1.694 | 0.3516 | Yes | ||
| 7 | GNA14 | ENSG00000156049.7 | 1907 | 1.665 | 0.4041 | Yes | ||
| 8 | GNB5 | ENSG00000069966.18 | 2282 | 1.540 | 0.4452 | Yes | ||
| 9 | GNAI2 | ENSG00000114353.17 | 5091 | 0.956 | 0.4110 | Yes | ||
| 10 | GNA15 | ENSG00000060558.4 | 5581 | 0.890 | 0.4284 | Yes | ||
| 11 | P2RY12 | ENSG00000169313.9 | 5618 | 0.886 | 0.4563 | Yes | ||
| 12 | GNAI1 | ENSG00000127955.17 | 5889 | 0.854 | 0.4776 | Yes | ||
| 13 | PLA2G4A | ENSG00000116711.10 | 6498 | 0.788 | 0.4890 | Yes | ||
| 14 | GNG7 | ENSG00000176533.13 | 6927 | 0.744 | 0.5032 | Yes | ||
| 15 | GNG10 | ENSG00000242616.4 | 7406 | 0.700 | 0.5147 | Yes | ||
| 16 | GNAI3 | ENSG00000065135.11 | 7451 | 0.695 | 0.5362 | Yes | ||
| 17 | MAPK14 | ENSG00000112062.10 | 10900 | 0.451 | 0.4707 | No | ||
| 18 | GNA11 | ENSG00000088256.9 | 12029 | 0.387 | 0.4571 | No | ||
| 19 | GNB4 | ENSG00000114450.10 | 16478 | 0.221 | 0.3610 | No | ||
| 20 | GNA13 | ENSG00000120063.10 | 19384 | 0.078 | 0.2961 | No | ||
| 21 | GNB2 | ENSG00000172354.10 | 19396 | 0.077 | 0.2983 | No | ||
| 22 | GNG2 | ENSG00000186469.8 | 23173 | -0.123 | 0.2147 | No | ||
| 23 | GNAQ | ENSG00000156052.11 | 25587 | -0.232 | 0.1662 | No | ||
| 24 | GNG3 | ENSG00000162188.6 | 27113 | -0.281 | 0.1399 | No | ||
| 25 | GNB1 | ENSG00000078369.18 | 32035 | -0.516 | 0.0424 | No | ||
| 26 | GNGT1 | ENSG00000127928.13 | 32085 | -0.520 | 0.0581 | No | ||
| 27 | AC093503.1 | ENSG00000268423.4 | 32791 | -0.568 | 0.0601 | No | ||
| 28 | GNAT3 | ENSG00000214415.4 | 33190 | -0.597 | 0.0702 | No | ||
| 29 | GNG5 | ENSG00000174021.11 | 33856 | -0.650 | 0.0758 | No | ||
| 30 | GNG13 | ENSG00000127588.5 | 35102 | -0.751 | 0.0711 | No | ||
| 31 | GNG4 | ENSG00000168243.11 | 35749 | -0.813 | 0.0825 | No | ||
| 32 | GNGT2 | ENSG00000167083.7 | 42221 | -2.737 | 0.0208 | No |