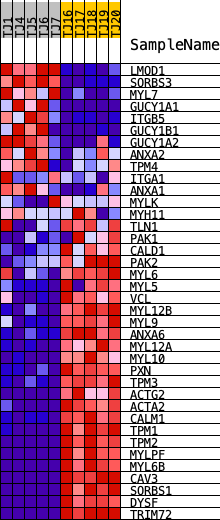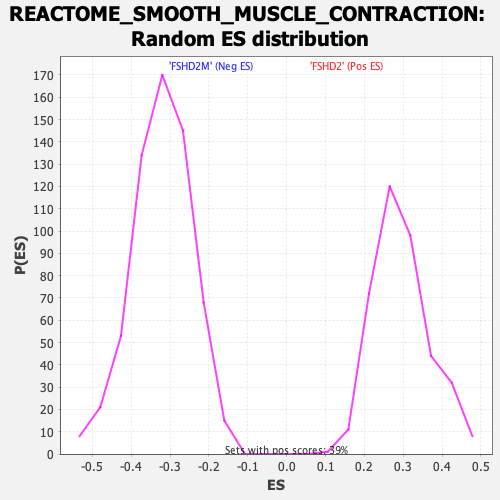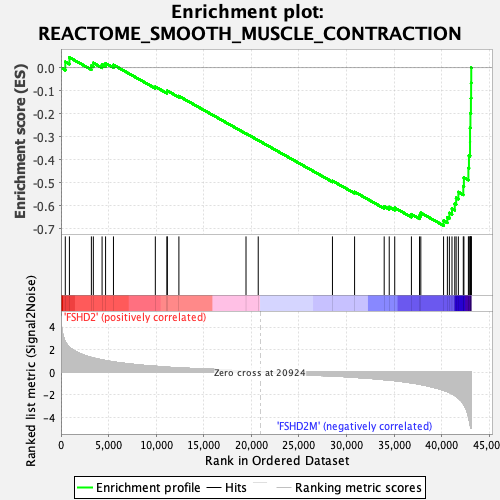
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_SMOOTH_MUSCLE_CONTRACTION |
| Enrichment Score (ES) | -0.6862957 |
| Normalized Enrichment Score (NES) | -2.1387484 |
| Nominal p-value | 0.0 |
| FDR q-value | 4.3756043E-4 |
| FWER p-Value | 0.005 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LMOD1 | ENSG00000163431.13 | 451 | 2.677 | 0.0257 | No | ||
| 2 | SORBS3 | ENSG00000120896.13 | 884 | 2.183 | 0.0452 | No | ||
| 3 | MYL7 | ENSG00000106631.8 | 3191 | 1.291 | 0.0091 | No | ||
| 4 | GUCY1A1 | ENSG00000164116.16 | 3397 | 1.246 | 0.0211 | No | ||
| 5 | ITGB5 | ENSG00000082781.12 | 4315 | 1.074 | 0.0144 | No | ||
| 6 | GUCY1B1 | ENSG00000061918.13 | 4682 | 1.017 | 0.0196 | No | ||
| 7 | GUCY1A2 | ENSG00000152402.10 | 5511 | 0.899 | 0.0125 | No | ||
| 8 | ANXA2 | ENSG00000182718.16 | 9909 | 0.516 | -0.0826 | No | ||
| 9 | TPM4 | ENSG00000167460.17 | 11125 | 0.437 | -0.1049 | No | ||
| 10 | ITGA1 | ENSG00000213949.10 | 11169 | 0.434 | -0.1000 | No | ||
| 11 | ANXA1 | ENSG00000135046.14 | 12384 | 0.370 | -0.1232 | No | ||
| 12 | MYLK | ENSG00000065534.18 | 19434 | 0.074 | -0.2858 | No | ||
| 13 | MYH11 | ENSG00000133392.18 | 20718 | 0.012 | -0.3154 | No | ||
| 14 | TLN1 | ENSG00000137076.21 | 28515 | -0.337 | -0.4918 | No | ||
| 15 | PAK1 | ENSG00000149269.9 | 30840 | -0.456 | -0.5396 | No | ||
| 16 | CALD1 | ENSG00000122786.20 | 33950 | -0.658 | -0.6029 | No | ||
| 17 | PAK2 | ENSG00000180370.10 | 34479 | -0.701 | -0.6057 | No | ||
| 18 | MYL6 | ENSG00000092841.18 | 35056 | -0.747 | -0.6090 | No | ||
| 19 | MYL5 | ENSG00000215375.6 | 36809 | -0.942 | -0.6369 | No | ||
| 20 | VCL | ENSG00000035403.17 | 37662 | -1.061 | -0.6424 | No | ||
| 21 | MYL12B | ENSG00000118680.13 | 37797 | -1.083 | -0.6308 | No | ||
| 22 | MYL9 | ENSG00000101335.10 | 40187 | -1.596 | -0.6647 | Yes | ||
| 23 | ANXA6 | ENSG00000197043.14 | 40587 | -1.711 | -0.6509 | Yes | ||
| 24 | MYL12A | ENSG00000101608.12 | 40797 | -1.785 | -0.6316 | Yes | ||
| 25 | MYL10 | ENSG00000106436.7 | 41072 | -1.896 | -0.6124 | Yes | ||
| 26 | PXN | ENSG00000089159.16 | 41365 | -2.042 | -0.5916 | Yes | ||
| 27 | TPM3 | ENSG00000143549.21 | 41507 | -2.135 | -0.5660 | Yes | ||
| 28 | ACTG2 | ENSG00000163017.14 | 41741 | -2.285 | -0.5406 | Yes | ||
| 29 | ACTA2 | ENSG00000107796.13 | 42259 | -2.777 | -0.5151 | Yes | ||
| 30 | CALM1 | ENSG00000198668.12 | 42323 | -2.877 | -0.4777 | Yes | ||
| 31 | TPM1 | ENSG00000140416.21 | 42794 | -3.877 | -0.4362 | Yes | ||
| 32 | TPM2 | ENSG00000198467.15 | 42854 | -4.136 | -0.3817 | Yes | ||
| 33 | MYLPF | ENSG00000180209.12 | 42967 | -4.555 | -0.3228 | Yes | ||
| 34 | MYL6B | ENSG00000196465.10 | 42969 | -4.580 | -0.2609 | Yes | ||
| 35 | CAV3 | ENSG00000182533.6 | 43009 | -4.750 | -0.1977 | Yes | ||
| 36 | SORBS1 | ENSG00000095637.22 | 43061 | -4.908 | -0.1326 | Yes | ||
| 37 | DYSF | ENSG00000135636.14 | 43081 | -4.948 | -0.0662 | Yes | ||
| 38 | TRIM72 | ENSG00000177238.14 | 43095 | -4.964 | 0.0006 | Yes |