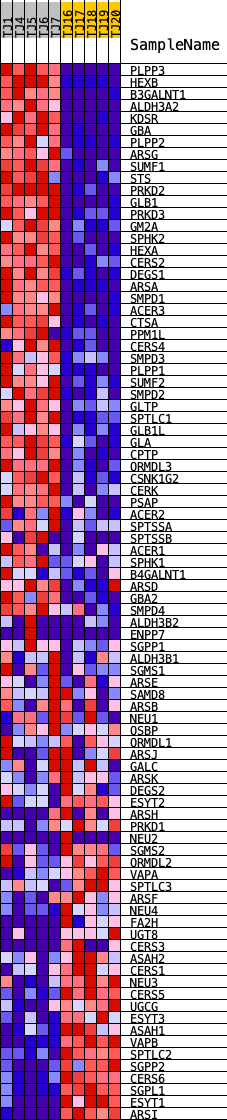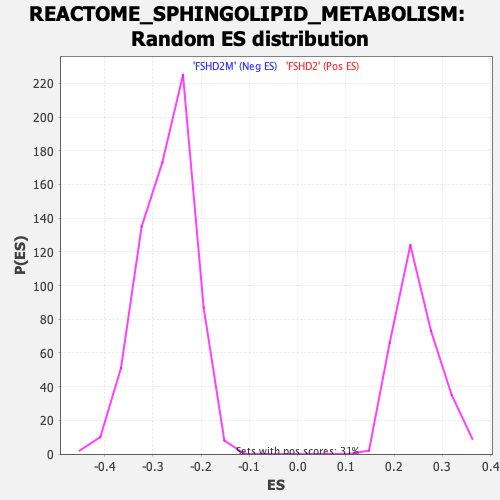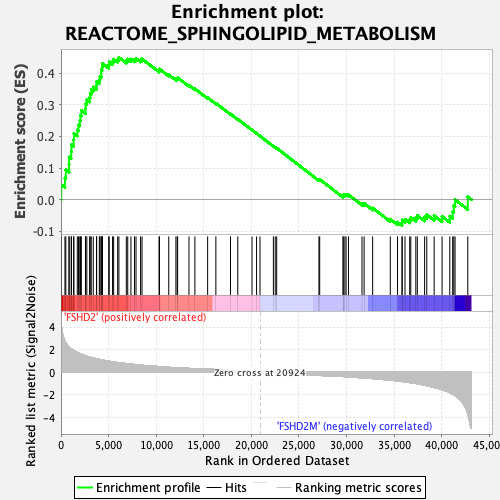
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_SPHINGOLIPID_METABOLISM |
| Enrichment Score (ES) | 0.44926846 |
| Normalized Enrichment Score (NES) | 1.8154521 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.03100423 |
| FWER p-Value | 0.671 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLPP3 | ENSG00000162407.9 | 18 | 4.425 | 0.0477 | Yes | ||
| 2 | HEXB | ENSG00000049860.14 | 415 | 2.734 | 0.0682 | Yes | ||
| 3 | B3GALNT1 | ENSG00000169255.15 | 507 | 2.592 | 0.0943 | Yes | ||
| 4 | ALDH3A2 | ENSG00000072210.19 | 840 | 2.223 | 0.1108 | Yes | ||
| 5 | KDSR | ENSG00000119537.18 | 853 | 2.214 | 0.1346 | Yes | ||
| 6 | GBA | ENSG00000177628.16 | 1072 | 2.067 | 0.1520 | Yes | ||
| 7 | PLPP2 | ENSG00000141934.10 | 1096 | 2.052 | 0.1738 | Yes | ||
| 8 | ARSG | ENSG00000141337.12 | 1326 | 1.924 | 0.1894 | Yes | ||
| 9 | SUMF1 | ENSG00000144455.14 | 1354 | 1.904 | 0.2095 | Yes | ||
| 10 | STS | ENSG00000101846.7 | 1712 | 1.750 | 0.2202 | Yes | ||
| 11 | PRKD2 | ENSG00000105287.12 | 1835 | 1.698 | 0.2359 | Yes | ||
| 12 | GLB1 | ENSG00000170266.16 | 1972 | 1.642 | 0.2505 | Yes | ||
| 13 | PRKD3 | ENSG00000115825.10 | 2052 | 1.612 | 0.2662 | Yes | ||
| 14 | GM2A | ENSG00000196743.9 | 2138 | 1.587 | 0.2815 | Yes | ||
| 15 | SPHK2 | ENSG00000063176.16 | 2570 | 1.450 | 0.2873 | Yes | ||
| 16 | HEXA | ENSG00000213614.9 | 2587 | 1.446 | 0.3026 | Yes | ||
| 17 | CERS2 | ENSG00000143418.19 | 2714 | 1.404 | 0.3150 | Yes | ||
| 18 | DEGS1 | ENSG00000143753.13 | 3004 | 1.332 | 0.3228 | Yes | ||
| 19 | ARSA | ENSG00000100299.18 | 3068 | 1.317 | 0.3356 | Yes | ||
| 20 | SMPD1 | ENSG00000166311.10 | 3178 | 1.292 | 0.3471 | Yes | ||
| 21 | ACER3 | ENSG00000078124.12 | 3389 | 1.247 | 0.3558 | Yes | ||
| 22 | CTSA | ENSG00000064601.18 | 3731 | 1.177 | 0.3607 | Yes | ||
| 23 | PPM1L | ENSG00000163590.14 | 3743 | 1.176 | 0.3732 | Yes | ||
| 24 | CERS4 | ENSG00000090661.12 | 4025 | 1.122 | 0.3789 | Yes | ||
| 25 | SMPD3 | ENSG00000103056.12 | 4087 | 1.114 | 0.3896 | Yes | ||
| 26 | PLPP1 | ENSG00000067113.17 | 4224 | 1.088 | 0.3983 | Yes | ||
| 27 | SUMF2 | ENSG00000129103.18 | 4226 | 1.088 | 0.4101 | Yes | ||
| 28 | SMPD2 | ENSG00000135587.9 | 4336 | 1.071 | 0.4192 | Yes | ||
| 29 | GLTP | ENSG00000139433.10 | 4348 | 1.069 | 0.4306 | Yes | ||
| 30 | SPTLC1 | ENSG00000090054.15 | 5014 | 0.966 | 0.4256 | Yes | ||
| 31 | GLB1L | ENSG00000163521.16 | 5059 | 0.960 | 0.4350 | Yes | ||
| 32 | GLA | ENSG00000102393.10 | 5414 | 0.912 | 0.4367 | Yes | ||
| 33 | CPTP | ENSG00000224051.7 | 5532 | 0.896 | 0.4438 | Yes | ||
| 34 | ORMDL3 | ENSG00000172057.10 | 5953 | 0.847 | 0.4432 | Yes | ||
| 35 | CSNK1G2 | ENSG00000133275.16 | 6082 | 0.831 | 0.4493 | Yes | ||
| 36 | CERK | ENSG00000100422.14 | 6861 | 0.751 | 0.4394 | No | ||
| 37 | PSAP | ENSG00000197746.14 | 7009 | 0.737 | 0.4440 | No | ||
| 38 | ACER2 | ENSG00000177076.6 | 7345 | 0.706 | 0.4439 | No | ||
| 39 | SPTSSA | ENSG00000165389.7 | 7716 | 0.671 | 0.4426 | No | ||
| 40 | SPTSSB | ENSG00000196542.8 | 7880 | 0.657 | 0.4459 | No | ||
| 41 | ACER1 | ENSG00000167769.5 | 8360 | 0.615 | 0.4415 | No | ||
| 42 | SPHK1 | ENSG00000176170.13 | 8512 | 0.605 | 0.4445 | No | ||
| 43 | B4GALNT1 | ENSG00000135454.14 | 10301 | 0.491 | 0.4083 | No | ||
| 44 | ARSD | ENSG00000006756.16 | 10345 | 0.488 | 0.4126 | No | ||
| 45 | GBA2 | ENSG00000070610.14 | 11319 | 0.426 | 0.3947 | No | ||
| 46 | SMPD4 | ENSG00000136699.19 | 12070 | 0.384 | 0.3814 | No | ||
| 47 | ALDH3B2 | ENSG00000132746.14 | 12210 | 0.377 | 0.3823 | No | ||
| 48 | ENPP7 | ENSG00000182156.10 | 12251 | 0.375 | 0.3854 | No | ||
| 49 | SGPP1 | ENSG00000126821.8 | 13443 | 0.331 | 0.3613 | No | ||
| 50 | ALDH3B1 | ENSG00000006534.16 | 14064 | 0.314 | 0.3503 | No | ||
| 51 | SGMS1 | ENSG00000198964.14 | 15404 | 0.280 | 0.3223 | No | ||
| 52 | ARSE | ENSG00000157399.15 | 16268 | 0.232 | 0.3047 | No | ||
| 53 | SAMD8 | ENSG00000156671.15 | 17804 | 0.157 | 0.2708 | No | ||
| 54 | ARSB | ENSG00000113273.17 | 18569 | 0.120 | 0.2543 | No | ||
| 55 | NEU1 | ENSG00000204386.11 | 20064 | 0.045 | 0.2201 | No | ||
| 56 | OSBP | ENSG00000110048.12 | 20544 | 0.019 | 0.2092 | No | ||
| 57 | ORMDL1 | ENSG00000128699.14 | 20901 | 0.002 | 0.2009 | No | ||
| 58 | ARSJ | ENSG00000180801.14 | 22309 | -0.075 | 0.1690 | No | ||
| 59 | GALC | ENSG00000054983.17 | 22503 | -0.084 | 0.1655 | No | ||
| 60 | ARSK | ENSG00000164291.17 | 22655 | -0.093 | 0.1630 | No | ||
| 61 | DEGS2 | ENSG00000168350.8 | 27095 | -0.280 | 0.0629 | No | ||
| 62 | ESYT2 | ENSG00000117868.17 | 27175 | -0.286 | 0.0641 | No | ||
| 63 | ARSH | ENSG00000205667.2 | 29627 | -0.383 | 0.0114 | No | ||
| 64 | PRKD1 | ENSG00000184304.15 | 29646 | -0.384 | 0.0151 | No | ||
| 65 | NEU2 | ENSG00000115488.3 | 29819 | -0.394 | 0.0154 | No | ||
| 66 | SGMS2 | ENSG00000164023.14 | 29947 | -0.403 | 0.0168 | No | ||
| 67 | ORMDL2 | ENSG00000123353.10 | 30197 | -0.421 | 0.0156 | No | ||
| 68 | VAPA | ENSG00000101558.13 | 31618 | -0.495 | -0.0120 | No | ||
| 69 | SPTLC3 | ENSG00000172296.13 | 31841 | -0.504 | -0.0117 | No | ||
| 70 | ARSF | ENSG00000062096.15 | 32737 | -0.564 | -0.0263 | No | ||
| 71 | NEU4 | ENSG00000204099.11 | 34590 | -0.709 | -0.0617 | No | ||
| 72 | FA2H | ENSG00000103089.9 | 35352 | -0.773 | -0.0709 | No | ||
| 73 | UGT8 | ENSG00000174607.11 | 35823 | -0.820 | -0.0729 | No | ||
| 74 | CERS3 | ENSG00000154227.13 | 35840 | -0.823 | -0.0644 | No | ||
| 75 | ASAH2 | ENSG00000188611.15 | 36136 | -0.860 | -0.0619 | No | ||
| 76 | CERS1 | ENSG00000223802.7 | 36629 | -0.919 | -0.0633 | No | ||
| 77 | NEU3 | ENSG00000162139.10 | 36743 | -0.935 | -0.0558 | No | ||
| 78 | CERS5 | ENSG00000139624.13 | 37273 | -1.002 | -0.0572 | No | ||
| 79 | UGCG | ENSG00000148154.10 | 37430 | -1.024 | -0.0496 | No | ||
| 80 | ESYT3 | ENSG00000158220.14 | 38196 | -1.153 | -0.0549 | No | ||
| 81 | ASAH1 | ENSG00000104763.19 | 38422 | -1.192 | -0.0472 | No | ||
| 82 | VAPB | ENSG00000124164.15 | 39198 | -1.352 | -0.0505 | No | ||
| 83 | SPTLC2 | ENSG00000100596.6 | 40030 | -1.556 | -0.0528 | No | ||
| 84 | SGPP2 | ENSG00000163082.10 | 40840 | -1.798 | -0.0521 | No | ||
| 85 | CERS6 | ENSG00000172292.14 | 41144 | -1.929 | -0.0382 | No | ||
| 86 | SGPL1 | ENSG00000166224.17 | 41263 | -1.982 | -0.0193 | No | ||
| 87 | ESYT1 | ENSG00000139641.12 | 41393 | -2.061 | 0.0001 | No | ||
| 88 | ARSI | ENSG00000183876.9 | 42731 | -3.679 | 0.0090 | No |