Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
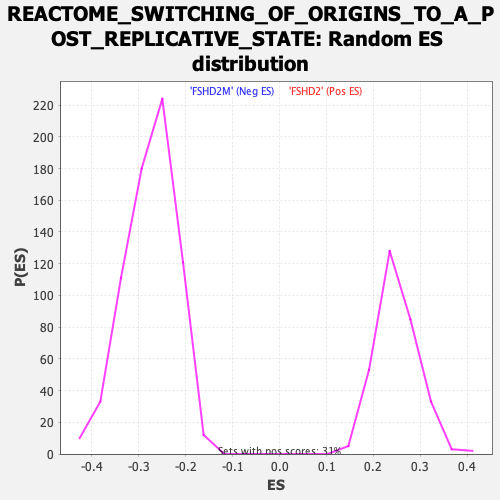
| GeneSet | REACTOME_SWITCHING_OF_ORIGINS_TO_A_POST_REPLICATIVE_STATE |
| Enrichment Score (ES) | -0.5733322 |
| Normalized Enrichment Score (NES) | -2.082754 |
| Nominal p-value | 0.0 |
| FDR q-value | 3.633605E-4 |
| FWER p-Value | 0.008 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMB9 | ENSG00000240065.8 | 653 | 2.409 | 0.0161 | No | ||
| 2 | PSMB8 | ENSG00000204264.11 | 1023 | 2.095 | 0.0346 | No | ||
| 3 | PSMB10 | ENSG00000205220.12 | 3086 | 1.313 | 0.0038 | No | ||
| 4 | CDC26 | ENSG00000176386.9 | 7372 | 0.703 | -0.0867 | No | ||
| 5 | ANAPC16 | ENSG00000166295.9 | 8822 | 0.582 | -0.1128 | No | ||
| 6 | UBA52 | ENSG00000221983.8 | 9987 | 0.512 | -0.1333 | No | ||
| 7 | PSMA2 | ENSG00000106588.11 | 16418 | 0.224 | -0.2798 | No | ||
| 8 | UBB | ENSG00000170315.14 | 17176 | 0.189 | -0.2949 | No | ||
| 9 | PSME1 | ENSG00000092010.15 | 17744 | 0.161 | -0.3060 | No | ||
| 10 | ANAPC4 | ENSG00000053900.10 | 19109 | 0.094 | -0.3365 | No | ||
| 11 | ANAPC5 | ENSG00000089053.13 | 19924 | 0.051 | -0.3548 | No | ||
| 12 | PSMF1 | ENSG00000125818.18 | 20560 | 0.019 | -0.3693 | No | ||
| 13 | MCM7 | ENSG00000166508.17 | 20623 | 0.016 | -0.3705 | No | ||
| 14 | SEM1 | ENSG00000127922.9 | 21274 | -0.020 | -0.3854 | No | ||
| 15 | ANAPC10 | ENSG00000164162.14 | 21601 | -0.037 | -0.3924 | No | ||
| 16 | UBC | ENSG00000150991.15 | 21944 | -0.054 | -0.3997 | No | ||
| 17 | CCNE2 | ENSG00000175305.17 | 21963 | -0.055 | -0.3994 | No | ||
| 18 | CDT1 | ENSG00000167513.9 | 22064 | -0.060 | -0.4009 | No | ||
| 19 | UBE2D1 | ENSG00000072401.15 | 22471 | -0.083 | -0.4093 | No | ||
| 20 | CDC16 | ENSG00000130177.16 | 23003 | -0.112 | -0.4202 | No | ||
| 21 | PSMD9 | ENSG00000110801.14 | 23060 | -0.115 | -0.4200 | No | ||
| 22 | PSMD5 | ENSG00000095261.14 | 23841 | -0.161 | -0.4360 | No | ||
| 23 | RPS27A | ENSG00000143947.13 | 24249 | -0.185 | -0.4431 | No | ||
| 24 | ORC5 | ENSG00000164815.11 | 24869 | -0.222 | -0.4546 | No | ||
| 25 | PSMA8 | ENSG00000154611.14 | 26134 | -0.263 | -0.4806 | No | ||
| 26 | CUL1 | ENSG00000055130.17 | 26496 | -0.266 | -0.4855 | No | ||
| 27 | ORC6 | ENSG00000091651.9 | 27104 | -0.281 | -0.4960 | No | ||
| 28 | PSMB11 | ENSG00000222028.5 | 27828 | -0.302 | -0.5089 | No | ||
| 29 | ANAPC2 | ENSG00000176248.9 | 27964 | -0.306 | -0.5080 | No | ||
| 30 | ORC2 | ENSG00000115942.9 | 28093 | -0.316 | -0.5069 | No | ||
| 31 | MCM6 | ENSG00000076003.5 | 28225 | -0.325 | -0.5057 | No | ||
| 32 | PSMB3 | ENSG00000277791.5 | 29269 | -0.361 | -0.5253 | No | ||
| 33 | ORC3 | ENSG00000135336.14 | 29356 | -0.366 | -0.5225 | No | ||
| 34 | MCM3 | ENSG00000112118.20 | 29614 | -0.382 | -0.5236 | No | ||
| 35 | PSMA1 | ENSG00000129084.17 | 30662 | -0.448 | -0.5421 | No | ||
| 36 | PSME3 | ENSG00000131467.11 | 30737 | -0.449 | -0.5380 | No | ||
| 37 | PSMA3 | ENSG00000100567.13 | 31911 | -0.509 | -0.5587 | No | ||
| 38 | FZR1 | ENSG00000105325.14 | 31978 | -0.512 | -0.5536 | No | ||
| 39 | CDC27 | ENSG00000004897.12 | 32830 | -0.571 | -0.5659 | Yes | ||
| 40 | MCM5 | ENSG00000100297.16 | 32907 | -0.577 | -0.5602 | Yes | ||
| 41 | PSMB1 | ENSG00000008018.9 | 33034 | -0.586 | -0.5556 | Yes | ||
| 42 | RBX1 | ENSG00000100387.9 | 33259 | -0.602 | -0.5530 | Yes | ||
| 43 | ORC4 | ENSG00000115947.13 | 33408 | -0.613 | -0.5484 | Yes | ||
| 44 | PSMD13 | ENSG00000185627.18 | 33437 | -0.616 | -0.5411 | Yes | ||
| 45 | PSMB4 | ENSG00000159377.11 | 33581 | -0.627 | -0.5363 | Yes | ||
| 46 | PSME2 | ENSG00000100911.16 | 33632 | -0.632 | -0.5293 | Yes | ||
| 47 | PSMD10 | ENSG00000101843.19 | 33841 | -0.648 | -0.5257 | Yes | ||
| 48 | CCNA1 | ENSG00000133101.10 | 34815 | -0.726 | -0.5389 | Yes | ||
| 49 | UBE2C | ENSG00000175063.17 | 34985 | -0.740 | -0.5332 | Yes | ||
| 50 | CCNA2 | ENSG00000145386.10 | 35426 | -0.780 | -0.5333 | Yes | ||
| 51 | PSMC5 | ENSG00000087191.13 | 35455 | -0.783 | -0.5238 | Yes | ||
| 52 | ANAPC1 | ENSG00000153107.13 | 35605 | -0.799 | -0.5170 | Yes | ||
| 53 | PSMA7 | ENSG00000101182.15 | 35709 | -0.810 | -0.5089 | Yes | ||
| 54 | PSMB2 | ENSG00000126067.12 | 35837 | -0.822 | -0.5011 | Yes | ||
| 55 | MCM4 | ENSG00000104738.18 | 35969 | -0.840 | -0.4933 | Yes | ||
| 56 | PSMA5 | ENSG00000143106.13 | 36170 | -0.865 | -0.4867 | Yes | ||
| 57 | UBE2E1 | ENSG00000170142.12 | 36422 | -0.891 | -0.4810 | Yes | ||
| 58 | PSMC2 | ENSG00000161057.12 | 36487 | -0.899 | -0.4708 | Yes | ||
| 59 | ORC1 | ENSG00000085840.13 | 36693 | -0.928 | -0.4636 | Yes | ||
| 60 | PSMB7 | ENSG00000136930.13 | 36803 | -0.941 | -0.4539 | Yes | ||
| 61 | SKP1 | ENSG00000113558.18 | 36892 | -0.952 | -0.4436 | Yes | ||
| 62 | CDC23 | ENSG00000094880.11 | 36926 | -0.956 | -0.4320 | Yes | ||
| 63 | MCM2 | ENSG00000073111.14 | 36965 | -0.962 | -0.4204 | Yes | ||
| 64 | CDK2 | ENSG00000123374.11 | 37047 | -0.974 | -0.4096 | Yes | ||
| 65 | PSMB6 | ENSG00000142507.10 | 37064 | -0.976 | -0.3974 | Yes | ||
| 66 | MCM8 | ENSG00000125885.13 | 37350 | -1.014 | -0.3908 | Yes | ||
| 67 | PSMA4 | ENSG00000041357.16 | 38333 | -1.177 | -0.3984 | Yes | ||
| 68 | PSMB5 | ENSG00000100804.19 | 38565 | -1.217 | -0.3880 | Yes | ||
| 69 | PSMD7 | ENSG00000103035.11 | 38868 | -1.282 | -0.3784 | Yes | ||
| 70 | ANAPC7 | ENSG00000196510.12 | 38905 | -1.290 | -0.3625 | Yes | ||
| 71 | ANAPC11 | ENSG00000141552.17 | 39180 | -1.349 | -0.3514 | Yes | ||
| 72 | PSMD6 | ENSG00000163636.10 | 39440 | -1.407 | -0.3391 | Yes | ||
| 73 | PSMA6 | ENSG00000100902.11 | 39610 | -1.444 | -0.3243 | Yes | ||
| 74 | PSMD3 | ENSG00000108344.15 | 39660 | -1.456 | -0.3066 | Yes | ||
| 75 | PSMD4 | ENSG00000159352.15 | 39708 | -1.468 | -0.2887 | Yes | ||
| 76 | ANAPC15 | ENSG00000110200.8 | 39750 | -1.479 | -0.2704 | Yes | ||
| 77 | PSMC1 | ENSG00000100764.14 | 39751 | -1.479 | -0.2513 | Yes | ||
| 78 | PSMD12 | ENSG00000197170.10 | 40160 | -1.590 | -0.2401 | Yes | ||
| 79 | PSMD8 | ENSG00000099341.11 | 40314 | -1.631 | -0.2225 | Yes | ||
| 80 | PSMC4 | ENSG00000013275.8 | 40405 | -1.659 | -0.2031 | Yes | ||
| 81 | PSMD11 | ENSG00000108671.11 | 40619 | -1.724 | -0.1857 | Yes | ||
| 82 | PSMD14 | ENSG00000115233.12 | 40751 | -1.767 | -0.1658 | Yes | ||
| 83 | PSMC6 | ENSG00000100519.12 | 40791 | -1.783 | -0.1436 | Yes | ||
| 84 | CCNE1 | ENSG00000105173.14 | 41239 | -1.974 | -0.1284 | Yes | ||
| 85 | PSMD1 | ENSG00000173692.13 | 41284 | -1.994 | -0.1036 | Yes | ||
| 86 | PSMC3 | ENSG00000165916.8 | 41467 | -2.112 | -0.0804 | Yes | ||
| 87 | CDC6 | ENSG00000094804.12 | 41614 | -2.200 | -0.0553 | Yes | ||
| 88 | SKP2 | ENSG00000145604.16 | 41748 | -2.288 | -0.0287 | Yes | ||
| 89 | PSME4 | ENSG00000068878.15 | 41799 | -2.328 | 0.0003 | Yes | ||
| 90 | PSMD2 | ENSG00000175166.17 | 41820 | -2.343 | 0.0302 | Yes |