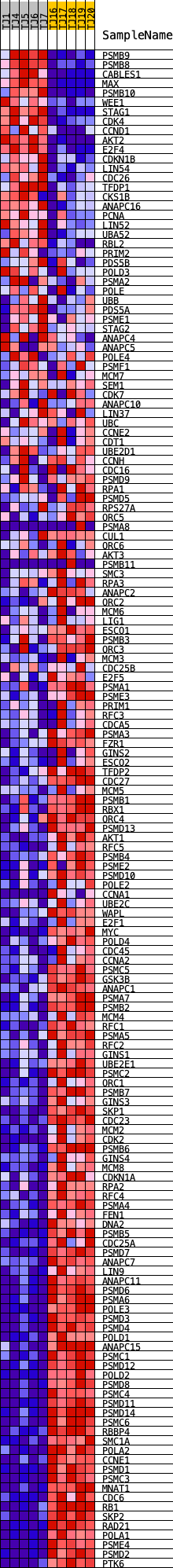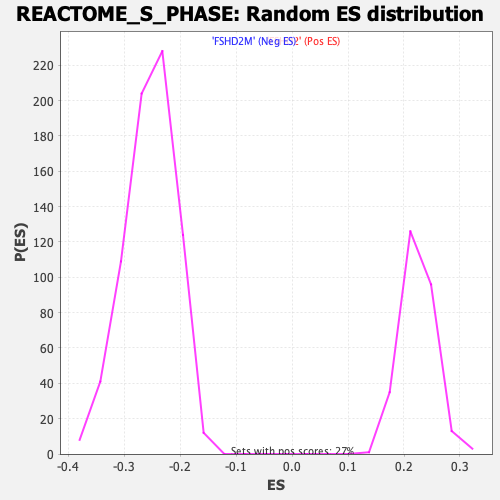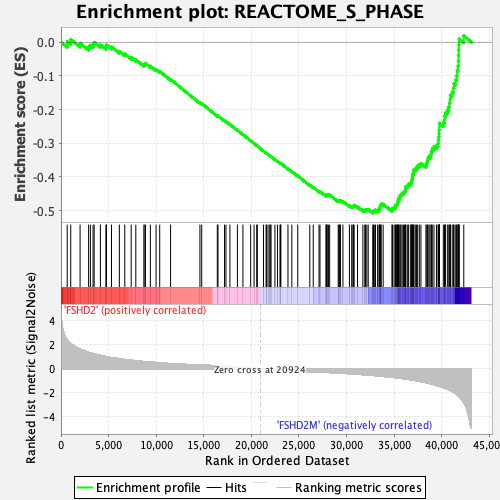
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2M |
| GeneSet | REACTOME_S_PHASE |
| Enrichment Score (ES) | -0.5076968 |
| Normalized Enrichment Score (NES) | -2.001254 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0012074279 |
| FWER p-Value | 0.04 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMB9 | ENSG00000240065.8 | 653 | 2.409 | 0.0020 | No | ||
| 2 | PSMB8 | ENSG00000204264.11 | 1023 | 2.095 | 0.0085 | No | ||
| 3 | CABLES1 | ENSG00000134508.12 | 2004 | 1.629 | -0.0027 | No | ||
| 4 | MAX | ENSG00000125952.20 | 2901 | 1.356 | -0.0139 | No | ||
| 5 | PSMB10 | ENSG00000205220.12 | 3086 | 1.313 | -0.0087 | No | ||
| 6 | WEE1 | ENSG00000166483.11 | 3367 | 1.252 | -0.0063 | No | ||
| 7 | STAG1 | ENSG00000118007.13 | 3483 | 1.228 | -0.0002 | No | ||
| 8 | CDK4 | ENSG00000135446.17 | 4137 | 1.105 | -0.0075 | No | ||
| 9 | CCND1 | ENSG00000110092.3 | 4717 | 1.011 | -0.0137 | No | ||
| 10 | AKT2 | ENSG00000105221.18 | 4775 | 1.001 | -0.0079 | No | ||
| 11 | E2F4 | ENSG00000205250.9 | 5303 | 0.926 | -0.0135 | No | ||
| 12 | CDKN1B | ENSG00000111276.11 | 6128 | 0.825 | -0.0268 | No | ||
| 13 | LIN54 | ENSG00000189308.11 | 6699 | 0.767 | -0.0346 | No | ||
| 14 | CDC26 | ENSG00000176386.9 | 7372 | 0.703 | -0.0452 | No | ||
| 15 | TFDP1 | ENSG00000198176.13 | 7858 | 0.659 | -0.0517 | No | ||
| 16 | CKS1B | ENSG00000173207.13 | 8704 | 0.590 | -0.0672 | No | ||
| 17 | ANAPC16 | ENSG00000166295.9 | 8822 | 0.582 | -0.0657 | No | ||
| 18 | PCNA | ENSG00000132646.11 | 8869 | 0.578 | -0.0627 | No | ||
| 19 | LIN52 | ENSG00000205659.10 | 9384 | 0.545 | -0.0707 | No | ||
| 20 | UBA52 | ENSG00000221983.8 | 9987 | 0.512 | -0.0811 | No | ||
| 21 | RBL2 | ENSG00000103479.16 | 10368 | 0.487 | -0.0865 | No | ||
| 22 | PRIM2 | ENSG00000146143.19 | 11511 | 0.413 | -0.1101 | No | ||
| 23 | PDS5B | ENSG00000083642.19 | 14594 | 0.304 | -0.1796 | No | ||
| 24 | POLD3 | ENSG00000077514.9 | 14781 | 0.293 | -0.1819 | No | ||
| 25 | PSMA2 | ENSG00000106588.11 | 16418 | 0.224 | -0.2184 | No | ||
| 26 | POLE | ENSG00000177084.17 | 16492 | 0.220 | -0.2185 | No | ||
| 27 | UBB | ENSG00000170315.14 | 17176 | 0.189 | -0.2330 | No | ||
| 28 | PDS5A | ENSG00000121892.15 | 17329 | 0.182 | -0.2353 | No | ||
| 29 | PSME1 | ENSG00000092010.15 | 17744 | 0.161 | -0.2438 | No | ||
| 30 | STAG2 | ENSG00000101972.18 | 18529 | 0.122 | -0.2611 | No | ||
| 31 | ANAPC4 | ENSG00000053900.10 | 19109 | 0.094 | -0.2739 | No | ||
| 32 | ANAPC5 | ENSG00000089053.13 | 19924 | 0.051 | -0.2925 | No | ||
| 33 | POLE4 | ENSG00000115350.12 | 20281 | 0.033 | -0.3006 | No | ||
| 34 | PSMF1 | ENSG00000125818.18 | 20560 | 0.019 | -0.3069 | No | ||
| 35 | MCM7 | ENSG00000166508.17 | 20623 | 0.016 | -0.3082 | No | ||
| 36 | SEM1 | ENSG00000127922.9 | 21274 | -0.020 | -0.3232 | No | ||
| 37 | CDK7 | ENSG00000134058.12 | 21543 | -0.034 | -0.3292 | No | ||
| 38 | ANAPC10 | ENSG00000164162.14 | 21601 | -0.037 | -0.3303 | No | ||
| 39 | LIN37 | ENSG00000267796.8 | 21753 | -0.045 | -0.3335 | No | ||
| 40 | UBC | ENSG00000150991.15 | 21944 | -0.054 | -0.3375 | No | ||
| 41 | CCNE2 | ENSG00000175305.17 | 21963 | -0.055 | -0.3375 | No | ||
| 42 | CDT1 | ENSG00000167513.9 | 22064 | -0.060 | -0.3394 | No | ||
| 43 | UBE2D1 | ENSG00000072401.15 | 22471 | -0.083 | -0.3483 | No | ||
| 44 | CCNH | ENSG00000134480.15 | 22740 | -0.098 | -0.3538 | No | ||
| 45 | CDC16 | ENSG00000130177.16 | 23003 | -0.112 | -0.3591 | No | ||
| 46 | PSMD9 | ENSG00000110801.14 | 23060 | -0.115 | -0.3596 | No | ||
| 47 | RPA1 | ENSG00000132383.12 | 23094 | -0.118 | -0.3595 | No | ||
| 48 | PSMD5 | ENSG00000095261.14 | 23841 | -0.161 | -0.3757 | No | ||
| 49 | RPS27A | ENSG00000143947.13 | 24249 | -0.185 | -0.3839 | No | ||
| 50 | ORC5 | ENSG00000164815.11 | 24869 | -0.222 | -0.3967 | No | ||
| 51 | PSMA8 | ENSG00000154611.14 | 26134 | -0.263 | -0.4243 | No | ||
| 52 | CUL1 | ENSG00000055130.17 | 26496 | -0.266 | -0.4308 | No | ||
| 53 | ORC6 | ENSG00000091651.9 | 27104 | -0.281 | -0.4429 | No | ||
| 54 | AKT3 | ENSG00000117020.18 | 27205 | -0.288 | -0.4431 | No | ||
| 55 | PSMB11 | ENSG00000222028.5 | 27828 | -0.302 | -0.4555 | No | ||
| 56 | SMC3 | ENSG00000108055.9 | 27903 | -0.302 | -0.4550 | No | ||
| 57 | RPA3 | ENSG00000106399.11 | 27907 | -0.302 | -0.4529 | No | ||
| 58 | ANAPC2 | ENSG00000176248.9 | 27964 | -0.306 | -0.4520 | No | ||
| 59 | ORC2 | ENSG00000115942.9 | 28093 | -0.316 | -0.4528 | No | ||
| 60 | MCM6 | ENSG00000076003.5 | 28225 | -0.325 | -0.4535 | No | ||
| 61 | LIG1 | ENSG00000105486.14 | 29118 | -0.354 | -0.4717 | No | ||
| 62 | ESCO1 | ENSG00000141446.11 | 29190 | -0.359 | -0.4708 | No | ||
| 63 | PSMB3 | ENSG00000277791.5 | 29269 | -0.361 | -0.4700 | No | ||
| 64 | ORC3 | ENSG00000135336.14 | 29356 | -0.366 | -0.4694 | No | ||
| 65 | MCM3 | ENSG00000112118.20 | 29614 | -0.382 | -0.4726 | No | ||
| 66 | CDC25B | ENSG00000101224.17 | 30297 | -0.429 | -0.4854 | No | ||
| 67 | E2F5 | ENSG00000133740.11 | 30537 | -0.437 | -0.4879 | No | ||
| 68 | PSMA1 | ENSG00000129084.17 | 30662 | -0.448 | -0.4876 | No | ||
| 69 | PSME3 | ENSG00000131467.11 | 30737 | -0.449 | -0.4861 | No | ||
| 70 | PRIM1 | ENSG00000198056.15 | 30787 | -0.453 | -0.4840 | No | ||
| 71 | RFC3 | ENSG00000133119.13 | 31151 | -0.472 | -0.4890 | No | ||
| 72 | CDCA5 | ENSG00000146670.10 | 31708 | -0.498 | -0.4984 | No | ||
| 73 | PSMA3 | ENSG00000100567.13 | 31911 | -0.509 | -0.4995 | No | ||
| 74 | FZR1 | ENSG00000105325.14 | 31978 | -0.512 | -0.4974 | No | ||
| 75 | GINS2 | ENSG00000131153.9 | 32126 | -0.521 | -0.4970 | No | ||
| 76 | ESCO2 | ENSG00000171320.15 | 32271 | -0.532 | -0.4966 | No | ||
| 77 | TFDP2 | ENSG00000114126.17 | 32749 | -0.564 | -0.5037 | Yes | ||
| 78 | CDC27 | ENSG00000004897.12 | 32830 | -0.571 | -0.5014 | Yes | ||
| 79 | MCM5 | ENSG00000100297.16 | 32907 | -0.577 | -0.4991 | Yes | ||
| 80 | PSMB1 | ENSG00000008018.9 | 33034 | -0.586 | -0.4978 | Yes | ||
| 81 | RBX1 | ENSG00000100387.9 | 33259 | -0.602 | -0.4987 | Yes | ||
| 82 | ORC4 | ENSG00000115947.13 | 33408 | -0.613 | -0.4978 | Yes | ||
| 83 | PSMD13 | ENSG00000185627.18 | 33437 | -0.616 | -0.4940 | Yes | ||
| 84 | AKT1 | ENSG00000142208.16 | 33520 | -0.623 | -0.4915 | Yes | ||
| 85 | RFC5 | ENSG00000111445.14 | 33522 | -0.623 | -0.4870 | Yes | ||
| 86 | PSMB4 | ENSG00000159377.11 | 33581 | -0.627 | -0.4839 | Yes | ||
| 87 | PSME2 | ENSG00000100911.16 | 33632 | -0.632 | -0.4805 | Yes | ||
| 88 | PSMD10 | ENSG00000101843.19 | 33841 | -0.648 | -0.4807 | Yes | ||
| 89 | POLE2 | ENSG00000100479.13 | 34764 | -0.721 | -0.4970 | Yes | ||
| 90 | CCNA1 | ENSG00000133101.10 | 34815 | -0.726 | -0.4930 | Yes | ||
| 91 | UBE2C | ENSG00000175063.17 | 34985 | -0.740 | -0.4916 | Yes | ||
| 92 | WAPL | ENSG00000062650.19 | 35121 | -0.752 | -0.4894 | Yes | ||
| 93 | E2F1 | ENSG00000101412.13 | 35122 | -0.753 | -0.4840 | Yes | ||
| 94 | MYC | ENSG00000136997.19 | 35251 | -0.763 | -0.4815 | Yes | ||
| 95 | POLD4 | ENSG00000175482.9 | 35331 | -0.771 | -0.4778 | Yes | ||
| 96 | CDC45 | ENSG00000093009.10 | 35359 | -0.774 | -0.4729 | Yes | ||
| 97 | CCNA2 | ENSG00000145386.10 | 35426 | -0.780 | -0.4689 | Yes | ||
| 98 | PSMC5 | ENSG00000087191.13 | 35455 | -0.783 | -0.4639 | Yes | ||
| 99 | GSK3B | ENSG00000082701.15 | 35603 | -0.798 | -0.4616 | Yes | ||
| 100 | ANAPC1 | ENSG00000153107.13 | 35605 | -0.799 | -0.4560 | Yes | ||
| 101 | PSMA7 | ENSG00000101182.15 | 35709 | -0.810 | -0.4526 | Yes | ||
| 102 | PSMB2 | ENSG00000126067.12 | 35837 | -0.822 | -0.4496 | Yes | ||
| 103 | MCM4 | ENSG00000104738.18 | 35969 | -0.840 | -0.4467 | Yes | ||
| 104 | RFC1 | ENSG00000035928.16 | 36076 | -0.852 | -0.4430 | Yes | ||
| 105 | PSMA5 | ENSG00000143106.13 | 36170 | -0.865 | -0.4390 | Yes | ||
| 106 | RFC2 | ENSG00000049541.11 | 36206 | -0.868 | -0.4336 | Yes | ||
| 107 | GINS1 | ENSG00000101003.10 | 36223 | -0.870 | -0.4278 | Yes | ||
| 108 | UBE2E1 | ENSG00000170142.12 | 36422 | -0.891 | -0.4260 | Yes | ||
| 109 | PSMC2 | ENSG00000161057.12 | 36487 | -0.899 | -0.4210 | Yes | ||
| 110 | ORC1 | ENSG00000085840.13 | 36693 | -0.928 | -0.4192 | Yes | ||
| 111 | PSMB7 | ENSG00000136930.13 | 36803 | -0.941 | -0.4150 | Yes | ||
| 112 | GINS3 | ENSG00000181938.13 | 36855 | -0.948 | -0.4094 | Yes | ||
| 113 | SKP1 | ENSG00000113558.18 | 36892 | -0.952 | -0.4034 | Yes | ||
| 114 | CDC23 | ENSG00000094880.11 | 36926 | -0.956 | -0.3973 | Yes | ||
| 115 | MCM2 | ENSG00000073111.14 | 36965 | -0.962 | -0.3913 | Yes | ||
| 116 | CDK2 | ENSG00000123374.11 | 37047 | -0.974 | -0.3862 | Yes | ||
| 117 | PSMB6 | ENSG00000142507.10 | 37064 | -0.976 | -0.3796 | Yes | ||
| 118 | GINS4 | ENSG00000147536.12 | 37217 | -0.994 | -0.3760 | Yes | ||
| 119 | MCM8 | ENSG00000125885.13 | 37350 | -1.014 | -0.3718 | Yes | ||
| 120 | CDKN1A | ENSG00000124762.13 | 37465 | -1.029 | -0.3671 | Yes | ||
| 121 | RPA2 | ENSG00000117748.10 | 37640 | -1.058 | -0.3636 | Yes | ||
| 122 | RFC4 | ENSG00000163918.10 | 37809 | -1.085 | -0.3598 | Yes | ||
| 123 | PSMA4 | ENSG00000041357.16 | 38333 | -1.177 | -0.3635 | Yes | ||
| 124 | FEN1 | ENSG00000168496.4 | 38431 | -1.194 | -0.3572 | Yes | ||
| 125 | DNA2 | ENSG00000138346.15 | 38510 | -1.209 | -0.3504 | Yes | ||
| 126 | PSMB5 | ENSG00000100804.19 | 38565 | -1.217 | -0.3429 | Yes | ||
| 127 | CDC25A | ENSG00000164045.12 | 38736 | -1.251 | -0.3379 | Yes | ||
| 128 | PSMD7 | ENSG00000103035.11 | 38868 | -1.282 | -0.3318 | Yes | ||
| 129 | ANAPC7 | ENSG00000196510.12 | 38905 | -1.290 | -0.3234 | Yes | ||
| 130 | LIN9 | ENSG00000183814.15 | 39000 | -1.310 | -0.3162 | Yes | ||
| 131 | ANAPC11 | ENSG00000141552.17 | 39180 | -1.349 | -0.3107 | Yes | ||
| 132 | PSMD6 | ENSG00000163636.10 | 39440 | -1.407 | -0.3067 | Yes | ||
| 133 | PSMA6 | ENSG00000100902.11 | 39610 | -1.444 | -0.3003 | Yes | ||
| 134 | POLE3 | ENSG00000148229.13 | 39649 | -1.454 | -0.2907 | Yes | ||
| 135 | PSMD3 | ENSG00000108344.15 | 39660 | -1.456 | -0.2806 | Yes | ||
| 136 | PSMD4 | ENSG00000159352.15 | 39708 | -1.468 | -0.2711 | Yes | ||
| 137 | POLD1 | ENSG00000062822.14 | 39718 | -1.471 | -0.2608 | Yes | ||
| 138 | ANAPC15 | ENSG00000110200.8 | 39750 | -1.479 | -0.2510 | Yes | ||
| 139 | PSMC1 | ENSG00000100764.14 | 39751 | -1.479 | -0.2404 | Yes | ||
| 140 | PSMD12 | ENSG00000197170.10 | 40160 | -1.590 | -0.2385 | Yes | ||
| 141 | POLD2 | ENSG00000106628.10 | 40288 | -1.623 | -0.2298 | Yes | ||
| 142 | PSMD8 | ENSG00000099341.11 | 40314 | -1.631 | -0.2187 | Yes | ||
| 143 | PSMC4 | ENSG00000013275.8 | 40405 | -1.659 | -0.2089 | Yes | ||
| 144 | PSMD11 | ENSG00000108671.11 | 40619 | -1.724 | -0.2016 | Yes | ||
| 145 | PSMD14 | ENSG00000115233.12 | 40751 | -1.767 | -0.1920 | Yes | ||
| 146 | PSMC6 | ENSG00000100519.12 | 40791 | -1.783 | -0.1801 | Yes | ||
| 147 | RBBP4 | ENSG00000162521.19 | 40876 | -1.813 | -0.1691 | Yes | ||
| 148 | SMC1A | ENSG00000072501.17 | 40902 | -1.825 | -0.1566 | Yes | ||
| 149 | POLA2 | ENSG00000014138.9 | 41096 | -1.906 | -0.1475 | Yes | ||
| 150 | CCNE1 | ENSG00000105173.14 | 41239 | -1.974 | -0.1366 | Yes | ||
| 151 | PSMD1 | ENSG00000173692.13 | 41284 | -1.994 | -0.1234 | Yes | ||
| 152 | PSMC3 | ENSG00000165916.8 | 41467 | -2.112 | -0.1125 | Yes | ||
| 153 | MNAT1 | ENSG00000020426.11 | 41558 | -2.168 | -0.0991 | Yes | ||
| 154 | CDC6 | ENSG00000094804.12 | 41614 | -2.200 | -0.0846 | Yes | ||
| 155 | RB1 | ENSG00000139687.15 | 41706 | -2.257 | -0.0706 | Yes | ||
| 156 | SKP2 | ENSG00000145604.16 | 41748 | -2.288 | -0.0551 | Yes | ||
| 157 | RAD21 | ENSG00000164754.14 | 41762 | -2.297 | -0.0390 | Yes | ||
| 158 | POLA1 | ENSG00000101868.11 | 41785 | -2.314 | -0.0229 | Yes | ||
| 159 | PSME4 | ENSG00000068878.15 | 41799 | -2.328 | -0.0066 | Yes | ||
| 160 | PSMD2 | ENSG00000175166.17 | 41820 | -2.343 | 0.0097 | Yes | ||
| 161 | PTK6 | ENSG00000101213.7 | 42311 | -2.863 | 0.0188 | Yes |