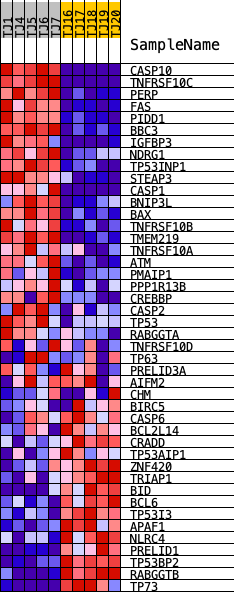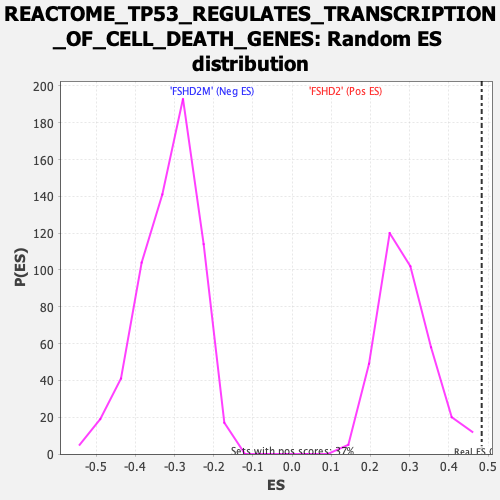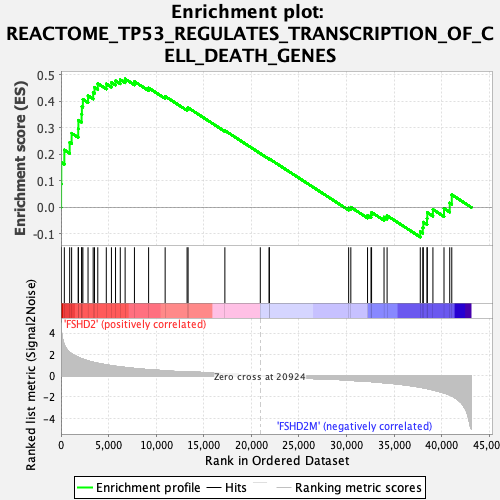
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Phenotype | Phenotype.FSHD2.FSHD2M.cls#FSHD2_versus_FSHD2M |
| Upregulated in class | FSHD2 |
| GeneSet | REACTOME_TP53_REGULATES_TRANSCRIPTION_OF_CELL_DEATH_GENES |
| Enrichment Score (ES) | 0.48391217 |
| Normalized Enrichment Score (NES) | 1.6764537 |
| Nominal p-value | 0.0027322404 |
| FDR q-value | 0.07939785 |
| FWER p-Value | 0.982 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CASP10 | ENSG00000003400.14 | 10 | 4.582 | 0.0876 | Yes | ||
| 2 | TNFRSF10C | ENSG00000173535.14 | 31 | 4.283 | 0.1692 | Yes | ||
| 3 | PERP | ENSG00000112378.12 | 350 | 2.857 | 0.2165 | Yes | ||
| 4 | FAS | ENSG00000026103.22 | 911 | 2.164 | 0.2450 | Yes | ||
| 5 | PIDD1 | ENSG00000177595.18 | 1116 | 2.042 | 0.2793 | Yes | ||
| 6 | BBC3 | ENSG00000105327.17 | 1808 | 1.712 | 0.2961 | Yes | ||
| 7 | IGFBP3 | ENSG00000146674.15 | 1828 | 1.702 | 0.3283 | Yes | ||
| 8 | NDRG1 | ENSG00000104419.14 | 2162 | 1.579 | 0.3508 | Yes | ||
| 9 | TP53INP1 | ENSG00000164938.14 | 2194 | 1.568 | 0.3801 | Yes | ||
| 10 | STEAP3 | ENSG00000115107.20 | 2299 | 1.536 | 0.4071 | Yes | ||
| 11 | CASP1 | ENSG00000137752.24 | 2840 | 1.369 | 0.4208 | Yes | ||
| 12 | BNIP3L | ENSG00000104765.16 | 3379 | 1.249 | 0.4322 | Yes | ||
| 13 | BAX | ENSG00000087088.20 | 3520 | 1.222 | 0.4524 | Yes | ||
| 14 | TNFRSF10B | ENSG00000120889.13 | 3864 | 1.152 | 0.4665 | Yes | ||
| 15 | TMEM219 | ENSG00000149932.17 | 4758 | 1.005 | 0.4651 | Yes | ||
| 16 | TNFRSF10A | ENSG00000104689.10 | 5298 | 0.927 | 0.4703 | Yes | ||
| 17 | ATM | ENSG00000149311.18 | 5728 | 0.872 | 0.4770 | Yes | ||
| 18 | PMAIP1 | ENSG00000141682.11 | 6227 | 0.814 | 0.4811 | Yes | ||
| 19 | PPP1R13B | ENSG00000088808.18 | 6736 | 0.763 | 0.4839 | Yes | ||
| 20 | CREBBP | ENSG00000005339.14 | 7714 | 0.671 | 0.4741 | No | ||
| 21 | CASP2 | ENSG00000106144.20 | 9203 | 0.555 | 0.4502 | No | ||
| 22 | TP53 | ENSG00000141510.17 | 10940 | 0.449 | 0.4185 | No | ||
| 23 | RABGGTA | ENSG00000100949.14 | 13244 | 0.343 | 0.3716 | No | ||
| 24 | TNFRSF10D | ENSG00000173530.6 | 13350 | 0.336 | 0.3756 | No | ||
| 25 | TP63 | ENSG00000073282.13 | 17214 | 0.187 | 0.2895 | No | ||
| 26 | PRELID3A | ENSG00000141391.14 | 20936 | -0.001 | 0.2031 | No | ||
| 27 | AIFM2 | ENSG00000042286.15 | 21859 | -0.049 | 0.1827 | No | ||
| 28 | CHM | ENSG00000188419.14 | 21888 | -0.051 | 0.1830 | No | ||
| 29 | BIRC5 | ENSG00000089685.15 | 30216 | -0.422 | -0.0022 | No | ||
| 30 | CASP6 | ENSG00000138794.10 | 30445 | -0.435 | 0.0008 | No | ||
| 31 | BCL2L14 | ENSG00000121380.12 | 32201 | -0.527 | -0.0299 | No | ||
| 32 | CRADD | ENSG00000169372.13 | 32566 | -0.548 | -0.0278 | No | ||
| 33 | TP53AIP1 | ENSG00000120471.15 | 32624 | -0.554 | -0.0185 | No | ||
| 34 | ZNF420 | ENSG00000197050.11 | 33933 | -0.657 | -0.0363 | No | ||
| 35 | TRIAP1 | ENSG00000170855.4 | 34255 | -0.679 | -0.0307 | No | ||
| 36 | BID | ENSG00000015475.18 | 37741 | -1.075 | -0.0911 | No | ||
| 37 | BCL6 | ENSG00000113916.18 | 38004 | -1.118 | -0.0757 | No | ||
| 38 | TP53I3 | ENSG00000115129.14 | 38060 | -1.126 | -0.0554 | No | ||
| 39 | APAF1 | ENSG00000120868.13 | 38448 | -1.197 | -0.0415 | No | ||
| 40 | NLRC4 | ENSG00000091106.19 | 38479 | -1.201 | -0.0192 | No | ||
| 41 | PRELID1 | ENSG00000169230.10 | 39069 | -1.324 | -0.0075 | No | ||
| 42 | TP53BP2 | ENSG00000143514.17 | 40233 | -1.609 | -0.0036 | No | ||
| 43 | RABGGTB | ENSG00000137955.16 | 40835 | -1.796 | 0.0168 | No | ||
| 44 | TP73 | ENSG00000078900.15 | 41049 | -1.889 | 0.0481 | No |