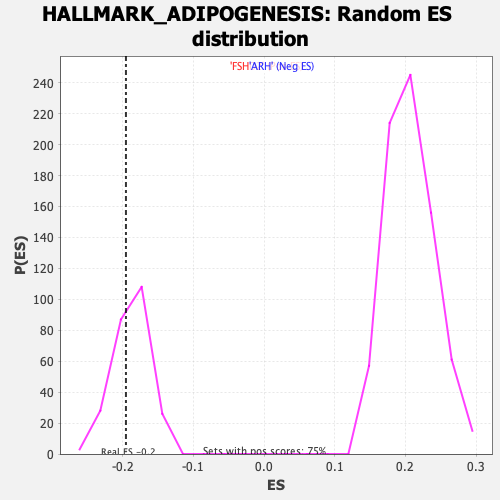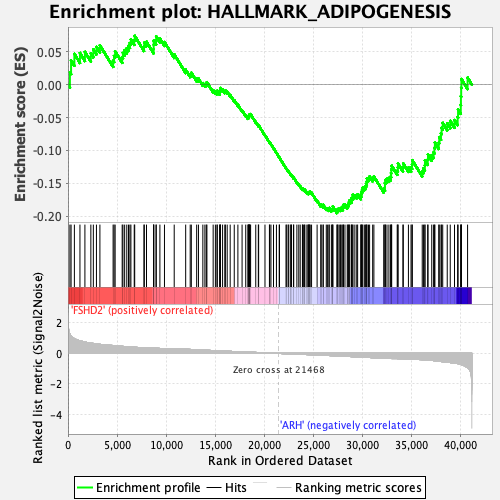
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | ARH |
| GeneSet | HALLMARK_ADIPOGENESIS |
| Enrichment Score (ES) | -0.19521315 |
| Normalized Enrichment Score (NES) | -1.0383682 |
| Nominal p-value | 0.32936507 |
| FDR q-value | 0.41460314 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LPL | ENSG00000175445.17 | 194 | 1.247 | 0.0187 | No | ||
| 2 | FABP4 | ENSG00000170323.9 | 313 | 1.123 | 0.0368 | No | ||
| 3 | TKT | ENSG00000163931.16 | 650 | 0.958 | 0.0466 | No | ||
| 4 | ACADL | ENSG00000115361.8 | 1218 | 0.813 | 0.0480 | No | ||
| 5 | EPHX2 | ENSG00000120915.14 | 1723 | 0.744 | 0.0497 | No | ||
| 6 | IFNGR1 | ENSG00000027697.14 | 2329 | 0.669 | 0.0474 | No | ||
| 7 | BCKDHA | ENSG00000248098.12 | 2582 | 0.645 | 0.0534 | No | ||
| 8 | LTC4S | ENSG00000213316.10 | 2899 | 0.614 | 0.0572 | No | ||
| 9 | MARC2 | ENSG00000117791.16 | 3252 | 0.587 | 0.0596 | No | ||
| 10 | DHCR7 | ENSG00000172893.15 | 4598 | 0.514 | 0.0364 | No | ||
| 11 | ABCA1 | ENSG00000165029.16 | 4697 | 0.511 | 0.0436 | No | ||
| 12 | MRAP | ENSG00000170262.12 | 4804 | 0.504 | 0.0505 | No | ||
| 13 | CD302 | ENSG00000241399.7 | 5523 | 0.458 | 0.0415 | No | ||
| 14 | DGAT1 | ENSG00000185000.12 | 5599 | 0.453 | 0.0482 | No | ||
| 15 | PIM3 | ENSG00000198355.5 | 5774 | 0.444 | 0.0523 | No | ||
| 16 | CCNG2 | ENSG00000138764.15 | 5991 | 0.434 | 0.0551 | No | ||
| 17 | SPARCL1 | ENSG00000152583.12 | 6166 | 0.425 | 0.0589 | No | ||
| 18 | SQOR | ENSG00000137767.14 | 6309 | 0.417 | 0.0632 | No | ||
| 19 | FAH | ENSG00000103876.12 | 6411 | 0.412 | 0.0685 | No | ||
| 20 | FZD4 | ENSG00000174804.4 | 6751 | 0.398 | 0.0677 | No | ||
| 21 | POR | ENSG00000127948.16 | 6799 | 0.396 | 0.0740 | No | ||
| 22 | ADIG | ENSG00000182035.12 | 7743 | 0.363 | 0.0577 | No | ||
| 23 | ADIPOQ | ENSG00000181092.10 | 7776 | 0.363 | 0.0638 | No | ||
| 24 | PFKFB3 | ENSG00000170525.21 | 8002 | 0.361 | 0.0650 | No | ||
| 25 | PEMT | ENSG00000133027.18 | 8729 | 0.346 | 0.0538 | No | ||
| 26 | STAT5A | ENSG00000126561.16 | 8732 | 0.346 | 0.0603 | No | ||
| 27 | CAT | ENSG00000121691.7 | 8736 | 0.346 | 0.0667 | No | ||
| 28 | LPCAT3 | ENSG00000111684.11 | 8979 | 0.336 | 0.0671 | No | ||
| 29 | ELMOD3 | ENSG00000115459.18 | 8991 | 0.336 | 0.0731 | No | ||
| 30 | SLC5A6 | ENSG00000138074.15 | 9367 | 0.320 | 0.0700 | No | ||
| 31 | MIGA2 | ENSG00000148343.18 | 9833 | 0.318 | 0.0646 | No | ||
| 32 | APOE | ENSG00000130203.10 | 10826 | 0.298 | 0.0460 | No | ||
| 33 | PFKL | ENSG00000141959.17 | 11995 | 0.271 | 0.0225 | No | ||
| 34 | ADCY6 | ENSG00000174233.11 | 12456 | 0.256 | 0.0161 | No | ||
| 35 | GPHN | ENSG00000171723.15 | 12570 | 0.252 | 0.0181 | No | ||
| 36 | C3 | ENSG00000125730.17 | 13115 | 0.235 | 0.0092 | No | ||
| 37 | CIDEA | ENSG00000176194.18 | 13297 | 0.229 | 0.0091 | No | ||
| 38 | ABCB8 | ENSG00000197150.12 | 13745 | 0.215 | 0.0022 | No | ||
| 39 | UCK1 | ENSG00000130717.13 | 13954 | 0.207 | 0.0010 | No | ||
| 40 | PTGER3 | ENSG00000050628.20 | 14098 | 0.203 | 0.0013 | No | ||
| 41 | SNCG | ENSG00000173267.14 | 14159 | 0.200 | 0.0036 | No | ||
| 42 | COQ9 | ENSG00000088682.14 | 14796 | 0.181 | -0.0086 | No | ||
| 43 | BAZ2A | ENSG00000076108.11 | 15023 | 0.173 | -0.0108 | No | ||
| 44 | CMBL | ENSG00000164237.9 | 15188 | 0.168 | -0.0117 | No | ||
| 45 | GPX4 | ENSG00000167468.17 | 15215 | 0.168 | -0.0092 | No | ||
| 46 | SCARB1 | ENSG00000073060.16 | 15471 | 0.160 | -0.0124 | No | ||
| 47 | BCL6 | ENSG00000113916.18 | 15472 | 0.160 | -0.0094 | No | ||
| 48 | CD151 | ENSG00000177697.19 | 15524 | 0.158 | -0.0077 | No | ||
| 49 | ALDH2 | ENSG00000111275.13 | 15545 | 0.158 | -0.0052 | No | ||
| 50 | SOWAHC | ENSG00000198142.5 | 15757 | 0.152 | -0.0075 | No | ||
| 51 | DNAJB9 | ENSG00000128590.5 | 15986 | 0.146 | -0.0103 | No | ||
| 52 | DHRS7 | ENSG00000100612.14 | 16041 | 0.144 | -0.0090 | No | ||
| 53 | RAB34 | ENSG00000109113.20 | 16227 | 0.139 | -0.0109 | No | ||
| 54 | SLC27A1 | ENSG00000130304.17 | 16531 | 0.130 | -0.0158 | No | ||
| 55 | JAGN1 | ENSG00000171135.15 | 16946 | 0.118 | -0.0237 | No | ||
| 56 | ME1 | ENSG00000065833.9 | 17317 | 0.109 | -0.0307 | No | ||
| 57 | ACADM | ENSG00000117054.13 | 17744 | 0.098 | -0.0392 | No | ||
| 58 | SLC19A1 | ENSG00000173638.19 | 18117 | 0.088 | -0.0467 | No | ||
| 59 | OMD | ENSG00000127083.7 | 18338 | 0.083 | -0.0505 | No | ||
| 60 | HSPB8 | ENSG00000152137.6 | 18377 | 0.082 | -0.0499 | No | ||
| 61 | TALDO1 | ENSG00000177156.11 | 18425 | 0.081 | -0.0495 | No | ||
| 62 | ANGPT1 | ENSG00000154188.10 | 18432 | 0.081 | -0.0481 | No | ||
| 63 | GPX3 | ENSG00000211445.12 | 18462 | 0.080 | -0.0473 | No | ||
| 64 | ACOX1 | ENSG00000161533.12 | 18471 | 0.080 | -0.0460 | No | ||
| 65 | NDUFS3 | ENSG00000213619.10 | 18514 | 0.078 | -0.0456 | No | ||
| 66 | IDH3G | ENSG00000067829.19 | 18543 | 0.078 | -0.0448 | No | ||
| 67 | LIPE | ENSG00000079435.10 | 18614 | 0.076 | -0.0451 | No | ||
| 68 | TST | ENSG00000128311.14 | 19138 | 0.061 | -0.0567 | No | ||
| 69 | ECH1 | ENSG00000104823.9 | 19400 | 0.054 | -0.0621 | No | ||
| 70 | DECR1 | ENSG00000104325.7 | 19432 | 0.053 | -0.0619 | No | ||
| 71 | ARAF | ENSG00000078061.13 | 20078 | 0.037 | -0.0769 | No | ||
| 72 | ITGA7 | ENSG00000135424.16 | 20532 | 0.024 | -0.0875 | No | ||
| 73 | SULT1A1 | ENSG00000196502.11 | 20558 | 0.023 | -0.0877 | No | ||
| 74 | PHLDB1 | ENSG00000019144.19 | 20685 | 0.020 | -0.0904 | No | ||
| 75 | UBC | ENSG00000150991.15 | 20948 | 0.013 | -0.0965 | No | ||
| 76 | RREB1 | ENSG00000124782.20 | 21246 | 0.005 | -0.1037 | No | ||
| 77 | ACO2 | ENSG00000100412.16 | 21529 | -0.002 | -0.1105 | No | ||
| 78 | ESRRA | ENSG00000173153.14 | 21558 | -0.003 | -0.1112 | No | ||
| 79 | PPP1R15B | ENSG00000158615.9 | 22221 | -0.019 | -0.1270 | No | ||
| 80 | AGPAT3 | ENSG00000160216.19 | 22294 | -0.021 | -0.1283 | No | ||
| 81 | MAP4K3 | ENSG00000011566.15 | 22445 | -0.025 | -0.1315 | No | ||
| 82 | MCCC1 | ENSG00000078070.13 | 22504 | -0.027 | -0.1324 | No | ||
| 83 | SAMM50 | ENSG00000100347.15 | 22709 | -0.031 | -0.1368 | No | ||
| 84 | NDUFB7 | ENSG00000099795.7 | 22725 | -0.031 | -0.1366 | No | ||
| 85 | NABP1 | ENSG00000173559.13 | 22786 | -0.033 | -0.1375 | No | ||
| 86 | IDH1 | ENSG00000138413.13 | 22996 | -0.039 | -0.1418 | No | ||
| 87 | AK2 | ENSG00000004455.17 | 22999 | -0.039 | -0.1411 | No | ||
| 88 | APLP2 | ENSG00000084234.17 | 23330 | -0.048 | -0.1483 | No | ||
| 89 | ACADS | ENSG00000122971.9 | 23524 | -0.053 | -0.1520 | No | ||
| 90 | GPAT4 | ENSG00000158669.11 | 23669 | -0.057 | -0.1545 | No | ||
| 91 | GBE1 | ENSG00000114480.13 | 23894 | -0.063 | -0.1587 | No | ||
| 92 | SLC25A1 | ENSG00000100075.10 | 23913 | -0.064 | -0.1580 | No | ||
| 93 | GADD45A | ENSG00000116717.13 | 23989 | -0.064 | -0.1586 | No | ||
| 94 | HADH | ENSG00000138796.16 | 24127 | -0.067 | -0.1607 | No | ||
| 95 | CPT2 | ENSG00000157184.7 | 24128 | -0.068 | -0.1594 | No | ||
| 96 | SLC25A10 | ENSG00000183048.12 | 24351 | -0.075 | -0.1635 | No | ||
| 97 | CYC1 | ENSG00000179091.5 | 24456 | -0.076 | -0.1646 | No | ||
| 98 | ALDOA | ENSG00000149925.20 | 24530 | -0.078 | -0.1649 | No | ||
| 99 | HIBCH | ENSG00000198130.16 | 24556 | -0.079 | -0.1640 | No | ||
| 100 | PREB | ENSG00000138073.14 | 24574 | -0.079 | -0.1630 | No | ||
| 101 | REEP6 | ENSG00000115255.11 | 24644 | -0.081 | -0.1631 | No | ||
| 102 | PEX14 | ENSG00000142655.13 | 24680 | -0.082 | -0.1624 | No | ||
| 103 | LAMA4 | ENSG00000112769.20 | 24827 | -0.086 | -0.1644 | No | ||
| 104 | ITSN1 | ENSG00000205726.15 | 25392 | -0.102 | -0.1762 | No | ||
| 105 | RETSAT | ENSG00000042445.14 | 25766 | -0.113 | -0.1832 | No | ||
| 106 | COX6A1 | ENSG00000111775.3 | 25808 | -0.114 | -0.1821 | No | ||
| 107 | PPARG | ENSG00000132170.21 | 26008 | -0.121 | -0.1847 | No | ||
| 108 | UQCRC1 | ENSG00000010256.11 | 26010 | -0.121 | -0.1824 | No | ||
| 109 | PTCD3 | ENSG00000132300.19 | 26326 | -0.129 | -0.1877 | No | ||
| 110 | AIFM1 | ENSG00000156709.14 | 26439 | -0.133 | -0.1879 | No | ||
| 111 | ENPP2 | ENSG00000136960.12 | 26598 | -0.138 | -0.1892 | No | ||
| 112 | ACAA2 | ENSG00000167315.18 | 26626 | -0.139 | -0.1873 | No | ||
| 113 | DRAM2 | ENSG00000156171.14 | 26845 | -0.145 | -0.1899 | No | ||
| 114 | ATL2 | ENSG00000119787.14 | 26930 | -0.147 | -0.1892 | No | ||
| 115 | DHRS7B | ENSG00000109016.17 | 26990 | -0.149 | -0.1878 | No | ||
| 116 | UQCR11 | ENSG00000127540.12 | 26991 | -0.149 | -0.1850 | No | ||
| 117 | COL15A1 | ENSG00000204291.11 | 27409 | -0.161 | -0.1922 | Yes | ||
| 118 | COX7B | ENSG00000131174.7 | 27456 | -0.163 | -0.1903 | Yes | ||
| 119 | SDHB | ENSG00000117118.9 | 27519 | -0.165 | -0.1887 | Yes | ||
| 120 | ECHS1 | ENSG00000127884.5 | 27650 | -0.169 | -0.1887 | Yes | ||
| 121 | COX8A | ENSG00000176340.4 | 27783 | -0.173 | -0.1887 | Yes | ||
| 122 | PHYH | ENSG00000107537.14 | 27848 | -0.175 | -0.1869 | Yes | ||
| 123 | SLC1A5 | ENSG00000105281.12 | 28017 | -0.181 | -0.1876 | Yes | ||
| 124 | ACLY | ENSG00000131473.17 | 28035 | -0.182 | -0.1847 | Yes | ||
| 125 | RMDN3 | ENSG00000137824.16 | 28083 | -0.183 | -0.1824 | Yes | ||
| 126 | ATP5PO | ENSG00000241837.7 | 28230 | -0.187 | -0.1824 | Yes | ||
| 127 | ETFB | ENSG00000105379.9 | 28464 | -0.195 | -0.1844 | Yes | ||
| 128 | DBT | ENSG00000137992.14 | 28531 | -0.197 | -0.1823 | Yes | ||
| 129 | SUCLG1 | ENSG00000163541.12 | 28603 | -0.199 | -0.1803 | Yes | ||
| 130 | NMT1 | ENSG00000136448.12 | 28656 | -0.201 | -0.1778 | Yes | ||
| 131 | MDH2 | ENSG00000146701.12 | 28720 | -0.203 | -0.1756 | Yes | ||
| 132 | CMPK1 | ENSG00000162368.13 | 28889 | -0.208 | -0.1758 | Yes | ||
| 133 | ANGPTL4 | ENSG00000167772.12 | 28905 | -0.208 | -0.1722 | Yes | ||
| 134 | ELOVL6 | ENSG00000170522.9 | 29029 | -0.213 | -0.1712 | Yes | ||
| 135 | CRAT | ENSG00000095321.17 | 29033 | -0.213 | -0.1673 | Yes | ||
| 136 | UQCR10 | ENSG00000184076.13 | 29240 | -0.219 | -0.1683 | Yes | ||
| 137 | UQCRQ | ENSG00000164405.11 | 29432 | -0.226 | -0.1687 | Yes | ||
| 138 | G3BP2 | ENSG00000138757.14 | 29523 | -0.228 | -0.1666 | Yes | ||
| 139 | ITIH5 | ENSG00000123243.15 | 29842 | -0.239 | -0.1699 | Yes | ||
| 140 | LIFR | ENSG00000113594.10 | 29881 | -0.241 | -0.1663 | Yes | ||
| 141 | TANK | ENSG00000136560.14 | 29913 | -0.241 | -0.1625 | Yes | ||
| 142 | COQ5 | ENSG00000110871.15 | 29985 | -0.243 | -0.1597 | Yes | ||
| 143 | ADIPOR2 | ENSG00000006831.10 | 30043 | -0.245 | -0.1565 | Yes | ||
| 144 | PLIN2 | ENSG00000147872.10 | 30207 | -0.250 | -0.1558 | Yes | ||
| 145 | RTN3 | ENSG00000133318.14 | 30339 | -0.255 | -0.1542 | Yes | ||
| 146 | IDH3A | ENSG00000166411.14 | 30386 | -0.256 | -0.1505 | Yes | ||
| 147 | STOM | ENSG00000148175.13 | 30456 | -0.259 | -0.1473 | Yes | ||
| 148 | PGM1 | ENSG00000079739.17 | 30468 | -0.259 | -0.1428 | Yes | ||
| 149 | PPM1B | ENSG00000138032.21 | 30632 | -0.263 | -0.1418 | Yes | ||
| 150 | CDKN2C | ENSG00000123080.12 | 30743 | -0.267 | -0.1395 | Yes | ||
| 151 | DNAJC15 | ENSG00000120675.6 | 31055 | -0.279 | -0.1418 | Yes | ||
| 152 | NDUFAB1 | ENSG00000004779.10 | 31179 | -0.284 | -0.1395 | Yes | ||
| 153 | NDUFA5 | ENSG00000128609.15 | 32180 | -0.303 | -0.1582 | Yes | ||
| 154 | SOD1 | ENSG00000142168.14 | 32307 | -0.310 | -0.1555 | Yes | ||
| 155 | YWHAG | ENSG00000170027.7 | 32308 | -0.310 | -0.1497 | Yes | ||
| 156 | SORBS1 | ENSG00000095637.22 | 32350 | -0.311 | -0.1448 | Yes | ||
| 157 | ESYT1 | ENSG00000139641.12 | 32515 | -0.317 | -0.1429 | Yes | ||
| 158 | SDHC | ENSG00000143252.14 | 32711 | -0.324 | -0.1416 | Yes | ||
| 159 | GRPEL1 | ENSG00000109519.13 | 32886 | -0.331 | -0.1396 | Yes | ||
| 160 | CS | ENSG00000062485.19 | 32933 | -0.333 | -0.1345 | Yes | ||
| 161 | GHITM | ENSG00000165678.21 | 32954 | -0.334 | -0.1287 | Yes | ||
| 162 | CD36 | ENSG00000135218.19 | 32994 | -0.336 | -0.1233 | Yes | ||
| 163 | IMMT | ENSG00000132305.20 | 33557 | -0.338 | -0.1307 | Yes | ||
| 164 | UBQLN1 | ENSG00000135018.14 | 33620 | -0.341 | -0.1258 | Yes | ||
| 165 | DDT | ENSG00000099977.14 | 33641 | -0.342 | -0.1199 | Yes | ||
| 166 | CHCHD10 | ENSG00000250479.8 | 34140 | -0.347 | -0.1256 | Yes | ||
| 167 | MRPL15 | ENSG00000137547.9 | 34174 | -0.348 | -0.1198 | Yes | ||
| 168 | VEGFB | ENSG00000173511.9 | 34699 | -0.356 | -0.1260 | Yes | ||
| 169 | SCP2 | ENSG00000116171.18 | 34965 | -0.364 | -0.1256 | Yes | ||
| 170 | MYLK | ENSG00000065534.18 | 35071 | -0.368 | -0.1213 | Yes | ||
| 171 | RIOK3 | ENSG00000101782.15 | 35090 | -0.368 | -0.1148 | Yes | ||
| 172 | CHUK | ENSG00000213341.11 | 36107 | -0.409 | -0.1319 | Yes | ||
| 173 | REEP5 | ENSG00000129625.13 | 36230 | -0.412 | -0.1272 | Yes | ||
| 174 | GPD2 | ENSG00000115159.16 | 36374 | -0.420 | -0.1228 | Yes | ||
| 175 | COL4A1 | ENSG00000187498.16 | 36388 | -0.420 | -0.1152 | Yes | ||
| 176 | GPAM | ENSG00000119927.14 | 36667 | -0.434 | -0.1139 | Yes | ||
| 177 | QDPR | ENSG00000151552.12 | 36694 | -0.435 | -0.1063 | Yes | ||
| 178 | BCL2L13 | ENSG00000099968.18 | 37078 | -0.457 | -0.1071 | Yes | ||
| 179 | DLAT | ENSG00000150768.15 | 37266 | -0.468 | -0.1029 | Yes | ||
| 180 | SSPN | ENSG00000123096.11 | 37348 | -0.474 | -0.0960 | Yes | ||
| 181 | UCP2 | ENSG00000175567.10 | 37400 | -0.478 | -0.0883 | Yes | ||
| 182 | DLD | ENSG00000091140.14 | 37792 | -0.512 | -0.0882 | Yes | ||
| 183 | TOB1 | ENSG00000141232.5 | 37842 | -0.516 | -0.0797 | Yes | ||
| 184 | COQ3 | ENSG00000132423.12 | 38018 | -0.529 | -0.0741 | Yes | ||
| 185 | PDCD4 | ENSG00000150593.18 | 38083 | -0.533 | -0.0657 | Yes | ||
| 186 | MGLL | ENSG00000074416.14 | 38196 | -0.542 | -0.0582 | Yes | ||
| 187 | MGST3 | ENSG00000143198.13 | 38656 | -0.567 | -0.0588 | Yes | ||
| 188 | RNF11 | ENSG00000123091.5 | 38963 | -0.596 | -0.0551 | Yes | ||
| 189 | LEP | ENSG00000174697.5 | 39396 | -0.626 | -0.0539 | Yes | ||
| 190 | ATP1B3 | ENSG00000069849.11 | 39716 | -0.663 | -0.0492 | Yes | ||
| 191 | MTCH2 | ENSG00000109919.10 | 39761 | -0.668 | -0.0378 | Yes | ||
| 192 | NKIRAS1 | ENSG00000197885.10 | 40036 | -0.720 | -0.0310 | Yes | ||
| 193 | ARL4A | ENSG00000122644.13 | 40051 | -0.722 | -0.0178 | Yes | ||
| 194 | PRDX3 | ENSG00000165672.7 | 40078 | -0.727 | -0.0048 | Yes | ||
| 195 | CAVIN1 | ENSG00000177469.13 | 40101 | -0.731 | 0.0084 | Yes | ||
| 196 | CAVIN2 | ENSG00000168497.5 | 40730 | -0.930 | 0.0105 | Yes |