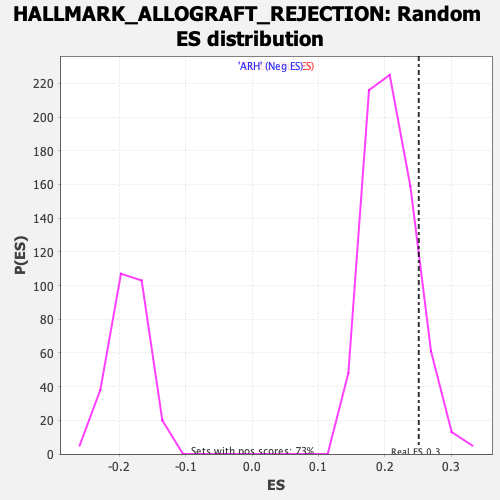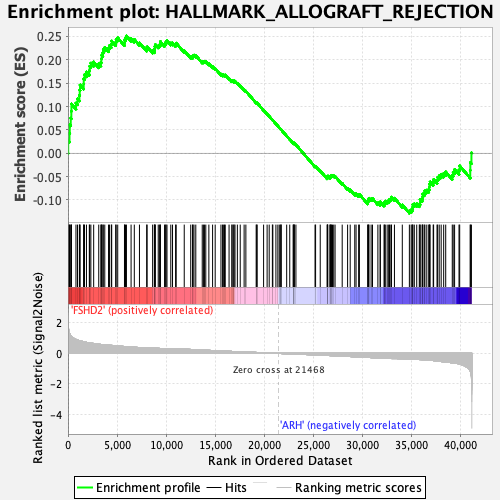
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Normalized.Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Phenotype | Phenotype.FSHD2.ARH.cls#FSHD2_versus_ARH |
| Upregulated in class | FSHD2 |
| GeneSet | HALLMARK_ALLOGRAFT_REJECTION |
| Enrichment Score (ES) | 0.25127137 |
| Normalized Enrichment Score (NES) | 1.2034577 |
| Nominal p-value | 0.13480055 |
| FDR q-value | 0.1992838 |
| FWER p-Value | 0.996 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | NLRP3 | ENSG00000162711.17 | 20 | 1.857 | 0.0267 | Yes | ||
| 2 | TGFB2 | ENSG00000092969.12 | 152 | 1.317 | 0.0428 | Yes | ||
| 3 | IL13 | ENSG00000169194.9 | 180 | 1.275 | 0.0608 | Yes | ||
| 4 | KLRD1 | ENSG00000134539.17 | 278 | 1.149 | 0.0752 | Yes | ||
| 5 | TPD52 | ENSG00000076554.15 | 340 | 1.096 | 0.0898 | Yes | ||
| 6 | TIMP1 | ENSG00000102265.12 | 352 | 1.090 | 0.1055 | Yes | ||
| 7 | IRF7 | ENSG00000185507.21 | 812 | 0.910 | 0.1076 | Yes | ||
| 8 | ITGAL | ENSG00000005844.18 | 971 | 0.866 | 0.1164 | Yes | ||
| 9 | IL16 | ENSG00000172349.17 | 1160 | 0.825 | 0.1239 | Yes | ||
| 10 | WAS | ENSG00000015285.10 | 1194 | 0.819 | 0.1351 | Yes | ||
| 11 | IRF4 | ENSG00000137265.15 | 1229 | 0.811 | 0.1461 | Yes | ||
| 12 | TLR2 | ENSG00000137462.8 | 1599 | 0.759 | 0.1482 | Yes | ||
| 13 | LIF | ENSG00000128342.5 | 1603 | 0.758 | 0.1593 | Yes | ||
| 14 | IGSF6 | ENSG00000140749.9 | 1683 | 0.748 | 0.1683 | Yes | ||
| 15 | CCL7 | ENSG00000108688.11 | 1881 | 0.720 | 0.1740 | Yes | ||
| 16 | ELANE | ENSG00000197561.7 | 2173 | 0.686 | 0.1770 | Yes | ||
| 17 | IL2RB | ENSG00000100385.14 | 2208 | 0.683 | 0.1861 | Yes | ||
| 18 | IFNGR1 | ENSG00000027697.14 | 2329 | 0.669 | 0.1930 | Yes | ||
| 19 | CD7 | ENSG00000173762.8 | 2617 | 0.642 | 0.1954 | Yes | ||
| 20 | SPI1 | ENSG00000066336.11 | 3134 | 0.596 | 0.1915 | Yes | ||
| 21 | PRF1 | ENSG00000180644.8 | 3351 | 0.580 | 0.1947 | Yes | ||
| 22 | LTB | ENSG00000227507.2 | 3397 | 0.576 | 0.2021 | Yes | ||
| 23 | IL1B | ENSG00000125538.12 | 3430 | 0.572 | 0.2097 | Yes | ||
| 24 | CCL4 | ENSG00000275302.2 | 3550 | 0.564 | 0.2150 | Yes | ||
| 25 | CD40LG | ENSG00000102245.7 | 3595 | 0.563 | 0.2222 | Yes | ||
| 26 | CFP | ENSG00000126759.13 | 3748 | 0.554 | 0.2266 | Yes | ||
| 27 | CD1D | ENSG00000158473.6 | 4133 | 0.535 | 0.2250 | Yes | ||
| 28 | BCL3 | ENSG00000069399.15 | 4224 | 0.529 | 0.2306 | Yes | ||
| 29 | LCP2 | ENSG00000043462.12 | 4413 | 0.519 | 0.2336 | Yes | ||
| 30 | PRKCG | ENSG00000126583.11 | 4454 | 0.519 | 0.2402 | Yes | ||
| 31 | CCND2 | ENSG00000118971.8 | 4862 | 0.500 | 0.2376 | Yes | ||
| 32 | HCLS1 | ENSG00000180353.11 | 4907 | 0.497 | 0.2438 | Yes | ||
| 33 | IL12B | ENSG00000113302.4 | 5068 | 0.486 | 0.2470 | Yes | ||
| 34 | IL18RAP | ENSG00000115607.9 | 5756 | 0.445 | 0.2368 | Yes | ||
| 35 | CD3G | ENSG00000160654.10 | 5767 | 0.445 | 0.2430 | Yes | ||
| 36 | ACHE | ENSG00000087085.15 | 5861 | 0.441 | 0.2472 | Yes | ||
| 37 | INHBB | ENSG00000163083.6 | 5957 | 0.435 | 0.2513 | Yes | ||
| 38 | TLR1 | ENSG00000174125.8 | 6430 | 0.410 | 0.2458 | No | ||
| 39 | CD96 | ENSG00000153283.13 | 6758 | 0.398 | 0.2436 | No | ||
| 40 | ZAP70 | ENSG00000115085.13 | 7283 | 0.376 | 0.2363 | No | ||
| 41 | HLA-A | ENSG00000206503.13 | 8022 | 0.360 | 0.2236 | No | ||
| 42 | HLA-G | ENSG00000204632.11 | 8061 | 0.359 | 0.2279 | No | ||
| 43 | CCR5 | ENSG00000160791.13 | 8627 | 0.348 | 0.2192 | No | ||
| 44 | CCR1 | ENSG00000163823.4 | 8802 | 0.344 | 0.2200 | No | ||
| 45 | CCL19 | ENSG00000172724.12 | 8838 | 0.342 | 0.2242 | No | ||
| 46 | RPL3L | ENSG00000140986.8 | 8854 | 0.341 | 0.2288 | No | ||
| 47 | IL4R | ENSG00000077238.14 | 8901 | 0.339 | 0.2326 | No | ||
| 48 | GBP2 | ENSG00000162645.13 | 9197 | 0.327 | 0.2302 | No | ||
| 49 | TAP1 | ENSG00000168394.11 | 9314 | 0.323 | 0.2321 | No | ||
| 50 | MBL2 | ENSG00000165471.6 | 9409 | 0.319 | 0.2345 | No | ||
| 51 | CD2 | ENSG00000116824.5 | 9412 | 0.319 | 0.2391 | No | ||
| 52 | CCL5 | ENSG00000271503.6 | 9843 | 0.317 | 0.2333 | No | ||
| 53 | ICAM1 | ENSG00000090339.9 | 9924 | 0.314 | 0.2359 | No | ||
| 54 | CD74 | ENSG00000019582.15 | 9998 | 0.311 | 0.2387 | No | ||
| 55 | F2R | ENSG00000181104.7 | 10089 | 0.309 | 0.2410 | No | ||
| 56 | CD3E | ENSG00000198851.9 | 10478 | 0.305 | 0.2360 | No | ||
| 57 | ICOSLG | ENSG00000160223.17 | 10654 | 0.304 | 0.2362 | No | ||
| 58 | SIT1 | ENSG00000137078.9 | 10969 | 0.292 | 0.2328 | No | ||
| 59 | HLA-E | ENSG00000204592.9 | 11025 | 0.290 | 0.2357 | No | ||
| 60 | LCK | ENSG00000182866.17 | 11848 | 0.276 | 0.2197 | No | ||
| 61 | IL11 | ENSG00000095752.7 | 12510 | 0.254 | 0.2073 | No | ||
| 62 | CTSS | ENSG00000163131.11 | 12694 | 0.248 | 0.2065 | No | ||
| 63 | PF4 | ENSG00000163737.4 | 12734 | 0.247 | 0.2091 | No | ||
| 64 | F2 | ENSG00000180210.14 | 12841 | 0.243 | 0.2101 | No | ||
| 65 | STAB1 | ENSG00000010327.10 | 13026 | 0.237 | 0.2091 | No | ||
| 66 | TLR6 | ENSG00000174130.12 | 13682 | 0.216 | 0.1962 | No | ||
| 67 | CD40 | ENSG00000101017.14 | 13803 | 0.213 | 0.1964 | No | ||
| 68 | MAP4K1 | ENSG00000104814.13 | 13908 | 0.209 | 0.1969 | No | ||
| 69 | GPR65 | ENSG00000140030.6 | 14045 | 0.204 | 0.1966 | No | ||
| 70 | CCL11 | ENSG00000172156.4 | 14358 | 0.196 | 0.1919 | No | ||
| 71 | CCL2 | ENSG00000108691.9 | 14738 | 0.183 | 0.1853 | No | ||
| 72 | NOS2 | ENSG00000007171.18 | 15003 | 0.173 | 0.1814 | No | ||
| 73 | TRAF2 | ENSG00000127191.18 | 15591 | 0.157 | 0.1694 | No | ||
| 74 | CD247 | ENSG00000198821.10 | 15755 | 0.152 | 0.1676 | No | ||
| 75 | AARS | ENSG00000090861.15 | 15834 | 0.149 | 0.1679 | No | ||
| 76 | IL2RA | ENSG00000134460.17 | 15953 | 0.146 | 0.1671 | No | ||
| 77 | HLA-DRA | ENSG00000204287.14 | 16028 | 0.144 | 0.1674 | No | ||
| 78 | CXCL9 | ENSG00000138755.6 | 16417 | 0.133 | 0.1599 | No | ||
| 79 | C2 | ENSG00000166278.15 | 16704 | 0.125 | 0.1548 | No | ||
| 80 | IL7 | ENSG00000104432.14 | 16729 | 0.124 | 0.1560 | No | ||
| 81 | LY75 | ENSG00000054219.11 | 16900 | 0.120 | 0.1536 | No | ||
| 82 | IFNGR2 | ENSG00000159128.14 | 16912 | 0.119 | 0.1551 | No | ||
| 83 | RPS3A | ENSG00000145425.10 | 17021 | 0.117 | 0.1542 | No | ||
| 84 | IL4 | ENSG00000113520.11 | 17263 | 0.110 | 0.1499 | No | ||
| 85 | THY1 | ENSG00000154096.13 | 17556 | 0.102 | 0.1443 | No | ||
| 86 | CSF1 | ENSG00000184371.14 | 17952 | 0.092 | 0.1360 | No | ||
| 87 | LYN | ENSG00000254087.8 | 18130 | 0.087 | 0.1329 | No | ||
| 88 | TLR3 | ENSG00000164342.12 | 19180 | 0.060 | 0.1082 | No | ||
| 89 | JAK2 | ENSG00000096968.13 | 19235 | 0.058 | 0.1078 | No | ||
| 90 | ST8SIA4 | ENSG00000113532.13 | 19279 | 0.057 | 0.1075 | No | ||
| 91 | PSMB10 | ENSG00000205220.12 | 19937 | 0.040 | 0.0921 | No | ||
| 92 | NPM1 | ENSG00000181163.13 | 20296 | 0.031 | 0.0838 | No | ||
| 93 | EREG | ENSG00000124882.4 | 20508 | 0.024 | 0.0790 | No | ||
| 94 | PTPN6 | ENSG00000111679.17 | 20832 | 0.017 | 0.0714 | No | ||
| 95 | IFNAR2 | ENSG00000159110.19 | 20879 | 0.015 | 0.0705 | No | ||
| 96 | RIPK2 | ENSG00000104312.8 | 21192 | 0.007 | 0.0630 | No | ||
| 97 | RPS9 | ENSG00000170889.14 | 21392 | 0.001 | 0.0581 | No | ||
| 98 | IKBKB | ENSG00000104365.15 | 21539 | -0.002 | 0.0546 | No | ||
| 99 | AKT1 | ENSG00000142208.16 | 21695 | -0.006 | 0.0509 | No | ||
| 100 | TNF | ENSG00000232810.4 | 21705 | -0.006 | 0.0508 | No | ||
| 101 | IL27RA | ENSG00000104998.4 | 21719 | -0.007 | 0.0506 | No | ||
| 102 | IL12RB1 | ENSG00000096996.16 | 22290 | -0.020 | 0.0370 | No | ||
| 103 | CXCL13 | ENSG00000156234.7 | 22603 | -0.029 | 0.0298 | No | ||
| 104 | B2M | ENSG00000166710.20 | 22947 | -0.038 | 0.0219 | No | ||
| 105 | HDAC9 | ENSG00000048052.21 | 23030 | -0.040 | 0.0205 | No | ||
| 106 | GALNT1 | ENSG00000141429.13 | 23031 | -0.040 | 0.0211 | No | ||
| 107 | RPS19 | ENSG00000105372.8 | 23032 | -0.040 | 0.0217 | No | ||
| 108 | CD8A | ENSG00000153563.15 | 23057 | -0.041 | 0.0217 | No | ||
| 109 | APBB1 | ENSG00000166313.19 | 23232 | -0.046 | 0.0181 | No | ||
| 110 | MMP9 | ENSG00000100985.7 | 25181 | -0.096 | -0.0280 | No | ||
| 111 | ACVR2A | ENSG00000121989.15 | 25237 | -0.097 | -0.0279 | No | ||
| 112 | PRKCB | ENSG00000166501.14 | 25710 | -0.111 | -0.0378 | No | ||
| 113 | TAPBP | ENSG00000231925.11 | 26413 | -0.133 | -0.0530 | No | ||
| 114 | BCAT1 | ENSG00000060982.15 | 26443 | -0.133 | -0.0517 | No | ||
| 115 | MTIF2 | ENSG00000085760.15 | 26449 | -0.134 | -0.0499 | No | ||
| 116 | MRPL3 | ENSG00000114686.9 | 26484 | -0.135 | -0.0488 | No | ||
| 117 | SOCS1 | ENSG00000185338.5 | 26705 | -0.141 | -0.0521 | No | ||
| 118 | CCL13 | ENSG00000181374.8 | 26725 | -0.141 | -0.0505 | No | ||
| 119 | CD8B | ENSG00000172116.23 | 26744 | -0.142 | -0.0488 | No | ||
| 120 | IL18 | ENSG00000150782.12 | 26822 | -0.144 | -0.0486 | No | ||
| 121 | TGFB1 | ENSG00000105329.10 | 26855 | -0.145 | -0.0473 | No | ||
| 122 | HLA-DOB | ENSG00000241106.8 | 26961 | -0.148 | -0.0477 | No | ||
| 123 | CD79A | ENSG00000105369.9 | 27085 | -0.152 | -0.0484 | No | ||
| 124 | EIF3D | ENSG00000100353.18 | 27222 | -0.156 | -0.0495 | No | ||
| 125 | ITGB2 | ENSG00000160255.18 | 27949 | -0.179 | -0.0646 | No | ||
| 126 | CRTAM | ENSG00000109943.9 | 28500 | -0.197 | -0.0751 | No | ||
| 127 | WARS | ENSG00000140105.18 | 28765 | -0.204 | -0.0786 | No | ||
| 128 | HIF1A | ENSG00000100644.17 | 29230 | -0.218 | -0.0867 | No | ||
| 129 | MAP3K7 | ENSG00000135341.18 | 29362 | -0.223 | -0.0866 | No | ||
| 130 | IL6 | ENSG00000136244.12 | 29624 | -0.231 | -0.0896 | No | ||
| 131 | CSK | ENSG00000103653.16 | 29702 | -0.235 | -0.0881 | No | ||
| 132 | EIF3J | ENSG00000104131.13 | 30550 | -0.261 | -0.1049 | No | ||
| 133 | RPL9 | ENSG00000163682.16 | 30577 | -0.262 | -0.1017 | No | ||
| 134 | STAT1 | ENSG00000115415.18 | 30631 | -0.263 | -0.0992 | No | ||
| 135 | UBE2D1 | ENSG00000072401.15 | 30679 | -0.265 | -0.0964 | No | ||
| 136 | GLMN | ENSG00000174842.17 | 30897 | -0.273 | -0.0977 | No | ||
| 137 | EIF5A | ENSG00000132507.18 | 31039 | -0.279 | -0.0971 | No | ||
| 138 | CCL22 | ENSG00000102962.5 | 31587 | -0.285 | -0.1062 | No | ||
| 139 | CD47 | ENSG00000196776.16 | 31817 | -0.290 | -0.1076 | No | ||
| 140 | ETS1 | ENSG00000134954.14 | 31835 | -0.291 | -0.1037 | No | ||
| 141 | ELF4 | ENSG00000102034.16 | 32222 | -0.306 | -0.1087 | No | ||
| 142 | EIF4G3 | ENSG00000075151.20 | 32249 | -0.307 | -0.1048 | No | ||
| 143 | CD80 | ENSG00000121594.12 | 32342 | -0.311 | -0.1025 | No | ||
| 144 | CCND3 | ENSG00000112576.12 | 32532 | -0.317 | -0.1025 | No | ||
| 145 | EIF3A | ENSG00000107581.13 | 32686 | -0.323 | -0.1015 | No | ||
| 146 | CAPG | ENSG00000042493.16 | 32769 | -0.326 | -0.0987 | No | ||
| 147 | EGFR | ENSG00000146648.18 | 32910 | -0.332 | -0.0973 | No | ||
| 148 | HLA-DOA | ENSG00000204252.14 | 32977 | -0.335 | -0.0940 | No | ||
| 149 | CD3D | ENSG00000167286.9 | 33274 | -0.336 | -0.0963 | No | ||
| 150 | FAS | ENSG00000026103.22 | 34075 | -0.344 | -0.1107 | No | ||
| 151 | STAT4 | ENSG00000138378.19 | 34796 | -0.361 | -0.1230 | No | ||
| 152 | FGR | ENSG00000000938.13 | 34978 | -0.364 | -0.1221 | No | ||
| 153 | IL12A | ENSG00000168811.7 | 35073 | -0.368 | -0.1190 | No | ||
| 154 | CDKN2A | ENSG00000147889.17 | 35138 | -0.371 | -0.1152 | No | ||
| 155 | SOCS5 | ENSG00000171150.9 | 35144 | -0.371 | -0.1099 | No | ||
| 156 | TAP2 | ENSG00000204267.15 | 35299 | -0.378 | -0.1081 | No | ||
| 157 | DARS | ENSG00000115866.11 | 35546 | -0.391 | -0.1084 | No | ||
| 158 | NCK1 | ENSG00000158092.7 | 35806 | -0.402 | -0.1088 | No | ||
| 159 | NCR1 | ENSG00000189430.13 | 35885 | -0.402 | -0.1048 | No | ||
| 160 | CD86 | ENSG00000114013.16 | 35887 | -0.402 | -0.0989 | No | ||
| 161 | UBE2N | ENSG00000177889.10 | 36090 | -0.408 | -0.0979 | No | ||
| 162 | FCGR2B | ENSG00000072694.20 | 36148 | -0.411 | -0.0933 | No | ||
| 163 | IL2RG | ENSG00000147168.12 | 36157 | -0.412 | -0.0874 | No | ||
| 164 | RPL39 | ENSG00000198918.8 | 36314 | -0.417 | -0.0851 | No | ||
| 165 | ABI1 | ENSG00000136754.17 | 36378 | -0.420 | -0.0805 | No | ||
| 166 | CXCR3 | ENSG00000186810.8 | 36572 | -0.430 | -0.0789 | No | ||
| 167 | ABCE1 | ENSG00000164163.11 | 36773 | -0.438 | -0.0774 | No | ||
| 168 | INHBA | ENSG00000122641.11 | 36833 | -0.442 | -0.0724 | No | ||
| 169 | DEGS1 | ENSG00000143753.13 | 36839 | -0.442 | -0.0660 | No | ||
| 170 | RARS | ENSG00000113643.9 | 36912 | -0.447 | -0.0612 | No | ||
| 171 | HLA-DMA | ENSG00000204257.15 | 37225 | -0.465 | -0.0620 | No | ||
| 172 | DYRK3 | ENSG00000143479.17 | 37267 | -0.468 | -0.0562 | No | ||
| 173 | SRGN | ENSG00000122862.5 | 37616 | -0.495 | -0.0574 | No | ||
| 174 | HLA-DMB | ENSG00000242574.9 | 37679 | -0.501 | -0.0516 | No | ||
| 175 | FLNA | ENSG00000196924.17 | 37839 | -0.515 | -0.0480 | No | ||
| 176 | NME1 | ENSG00000239672.7 | 38038 | -0.530 | -0.0450 | No | ||
| 177 | GCNT1 | ENSG00000187210.14 | 38293 | -0.551 | -0.0432 | No | ||
| 178 | GZMB | ENSG00000100453.13 | 38501 | -0.558 | -0.0400 | No | ||
| 179 | CD28 | ENSG00000178562.18 | 39168 | -0.615 | -0.0473 | No | ||
| 180 | BCL10 | ENSG00000142867.14 | 39261 | -0.619 | -0.0405 | No | ||
| 181 | ITK | ENSG00000113263.12 | 39406 | -0.627 | -0.0348 | No | ||
| 182 | BRCA1 | ENSG00000012048.22 | 39857 | -0.687 | -0.0357 | No | ||
| 183 | CD4 | ENSG00000010610.10 | 39926 | -0.700 | -0.0271 | No | ||
| 184 | FYB1 | ENSG00000082074.18 | 41002 | -1.149 | -0.0365 | No | ||
| 185 | IL15 | ENSG00000164136.17 | 41012 | -1.156 | -0.0199 | No | ||
| 186 | PTPRC | ENSG00000081237.20 | 41122 | -1.603 | 0.0010 | No |